Abstract
The sodium channel NaV1.6 is widely expressed in neurons of the central and peripheral nervous systems, which plays a critical role in regulating neuronal excitability. Dysfunction of NaV1.6 has been linked to epileptic encephalopathy, intellectual disability and movement disorders. Here we present cryo-EM structures of human NaV1.6/β1/β2 alone and complexed with a guanidinium neurotoxin 4,9-anhydro-tetrodotoxin (4,9-ah-TTX), revealing molecular mechanism of NaV1.6 inhibition by the blocker. The apo-form structure reveals two potential Na+ binding sites within the selectivity filter, suggesting a possible mechanism for Na+ selectivity and conductance. In the 4,9-ah-TTX bound structure, 4,9-ah-TTX binds to a pocket similar to the tetrodotoxin (TTX) binding site, which occupies the Na+ binding sites and completely blocks the channel. Molecular dynamics simulation results show that subtle conformational differences in the selectivity filter affect the affinity of TTX analogues. Taken together, our results provide important insights into NaV1.6 structure, ion conductance, and inhibition.
Similar content being viewed by others
Introduction
Voltage-gated sodium (NaV) channels mediate the generation and propagation of action potentials in excitable cells1,2. In humans, nine NaV channel subtypes (NaV1.1–1.9) had been identified, which are involved in a broad range of physiological processes due to their tissue-specific distributions in various excitable tissues3,4. Subtype NaV1.6, encoded by the gene SCN8A, is ubiquitously expressed in neurons of both the central nervous system (CNS) and the peripheral nervous system (PNS), especially enriched in the distal end of axon initial segment (AIS) and in the node of Ranvier of myelinated excitatory neurons. The NaV1.6 channel is believed to play a primary role in the initiation and propagation of action potentials in those neurons by lowering the threshold voltage5,6,7,8,9,10,11. Emerging evidence suggests that NaV1.6 is also expressed in some inhibitory interneurons and plays a role in establishing synaptic inhibition in the thalamic networks12,13,14. Compared with other NaV channel subtypes, NaV1.6 possesses unique biophysical properties including activation at more hyperpolarized voltage, higher levels of persistent current and resurgent current, and higher frequency of repetitive neuronal firing in neurons such as cerebellar Purkinje cells15,16,17,18,19,20,21,22,23. These features make NaV1.6 a critical and favorable mediator in regulating neuronal excitability in those neurons. Meanwhile, dozens of mutations in NaV1.6 have been linked to human diseases, most of which exhibit gain-of-function phenotypes, increase neuronal excitability, and cause different types of epileptic encephalopathy24,25,26,27,28; whereas loss-of-function mutations are often associated with later onset seizures, intellectual disability, isolated cognitive impairment and movement disorders29,30,31. Thus, NaV1.6 is an important drug target; effective and subtype-selective therapeutics are eagerly awaited for the treatment of NaV1.6-related epilepsy and other neurological diseases.
Eukaryotic NaV channels are composed of a pore-forming α subunit and auxiliary β subunits32. The four-domain α subunit exerts voltage sensing, gate opening, ion permeation, and inactivation4,33. Meanwhile, one or two β subunits bind to the α subunit to regulate NaV channel kinetics and trafficking. Among the four types of β subunits34,35,36,37, β1 and β3 subunits non-covalently bind to the α subunit, while β2 and β4 subunits are covalently linked to the α subunit via a disulfide bond32,38. To date, high-resolution cryo-electron microscopy (cryo-EM) structures of seven mammalian NaV channels (NaV1.1–1.5, NaV1.7–1.8) have been reported39,40,41,42,43,44,45. Together with the resting-state46, open-state47, and multiple ligand-bound NaV channel structures48,49,50, these structures revealed the general molecular mechanisms of voltage-sensing, electromechanically coupling, fast inactivation, sodium permeation, and ligand modulation. Among those NaV channel modulators, the guanidinium neurotoxin tetrodotoxin (TTX) has long been used as a useful tool to study NaV channels, which can potently inhibit NaV1.1–1.4 and NaV1.6–1.7 at nanomolar level (TTX-sensitive NaV channels), and less potently inhibit NaV1.5, NaV1.8, and NaV1.9 at a micromolar concentration (TTX-insensitive NaV channels). The detailed binding mode of TTX had been revealed in the NaV channel-TTX complex structures44,51. Furthermore, two guanidinium neurotoxin derivatives, ST-2262 and ST-2530, were reported as potent and selective inhibitors for NaV1.7, indicating that TTX analogs could potentially be developed as selective therapeutics52,53. Interestingly, 4,9-anhydro-tetrodotoxin (4,9-ah-TTX), a metabolite of TTX, has been reported to selectively block NaV1.6 with a blocking efficacy of 40- to 160-fold higher than other TTX-sensitive NaV channels54. However, the structure of NaV1.6 and how 4,9-ah-TTX blocks NaV1.6 remain elusive.
In this work, we show a fully-functional shorter-form construct of human NaV1.6 suitable for structural studies, and present cryo-EM structures of NaV1.6/β1/β2 apo-form and in complex with 4,9-ah-TTX. Complemented with electrophysiological results and molecular dynamics (MD) simulations, our structures reveal NaV1.6 structural features, sodium conductance, and pore-blockade by 4,9-ah-TTX.
Results
Construct optimization of NaV1.6 for cryo-EM study
To conduct structural studies of NaV1.6, human wide-type NaV1.6 (named NaV1.6WT) was co-expressed with human β1 and β2 subunits in HEK293F cells and was purified similarly to previously reported NaV channels41,44. Although the amino acid sequence of NaV1.6 is highly conserved with other NaV channel subtypes (e.g., 70% identity with NaV1.7); however, the purified NaV1.6WT sample exhibited poor quality and did not permit high-resolution structural analysis (Supplementary Fig. 1a, b). Construct optimization had been proven to be successful in improving the sample quality of NaV1.7 and NaV1.555,56, we therefore carried out construct screening of human NaV1.6 by removing unstructured intracellular loops and C-terminus. We found that deletion of S478-G692 between DI and DII (NaV1.6ΔDI-DII), S1115-L1180 between DII and DIII (NaV1.6ΔDII-DIII), or R1932-C1980 of the C-terminus (NaV1.6ΔCter) showed improved sample homogeneity based on the size-exclusion chromatography (SEC) profiles (Supplementary Fig. 1a). Strikingly, when we combined these modifications and deleted all of the three unstructured regions, it displayed a sharp mono-disperse SEC profile, which is much better than that of NaV1.6WT and any of the single-deletion constructs (Fig. 1a, b, Supplementary Fig. 1a). We next examined the functional characteristics of the triple-deletion construct by whole-cell voltage-clamp recording of NaV1.6-expressing HEK293T cells. The candidate construct exhibits typical voltage-dependent activation and inactivation (Fig. 1c). The resulting V1/2 values of the voltage-dependence of activation and steady-state fast inactivation are −31.3 ± 0.3 mV (n = 15) and −77.3 ± 0.2 mV (n = 15), respectively, which are close to the reported V1/2 values of human wide-type NaV1.657,58. These results confirmed that the triple-deletion construct fulfills similar electrophysiological functions to the NaV1.6WT. The preliminary cryo-EM analysis of this triple-deletion construct showed that the micrograph contains a rich distribution of mono-disperse particles, which gave rise to much better 2D class averages with well-resolved features than the NaV1.6WT (Supplementary Fig. 1b, c). Thus, this triple-deletion construct (named NaV1.6EM) was selected for further structural studies.
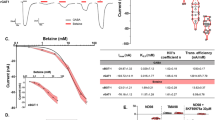
a Topology of the NaV1.6/β1/β2 complex. The α subunit consists of DI (purple), DII (yellow), DIII (pink), and DIV (cyan) connected by intracellular linkers, a mCherry fluorescent protein tag fused at the C-terminus. Scissors indicate the truncated sites. The β1 fused with a GFP tag at the C-terminus and the β2 subunit are highlighted in light blue and gray, respectively. The same color codes for NaV1.6/β1/β2 are applied throughout the manuscript unless specified. b Size exclusion chromatogram profiles of the purified NaV1.6WT (black) and the NaV1.6EM (red). c Electrophysiological characterization of the NaV1.6EM construct. The voltage protocols and representative current traces are shown on the left panels. To characterize the voltage-dependence of activation, NaV1.6EM expressing HEK293T cells were stimulated by a 100 ms test pulse varying from −80 mV to 10 mV in 5 mV increments from a holding potential of −120 mV, with a stimulus frequency of 0.2 Hz. To measure the steady-state fast inactivation, HEK293T cells were stimulated by a test step to −10 mV after a 500 ms prepulse varying from −130 mV to −20 mV in 5 mV increments, from a holding potential of −120 mV and a stimulus frequency of 0.2 Hz. The resulting normalized conductance-voltage (G/V) relationship (squares, n = 15) and steady-state fast inactivation (circles, n = 15) curves are shown on the right panel. Data are presented as mean ± SEM. n biological independent cells. Source data are provided as a Source data file.
The overall structure of human NaV1.6
The purified NaV1.6EM/β1/β2 sample was frozen in vitreous ice for cryo-EM data collection (Supplementary Fig. 2). After processing, the final reconstruction map from the best class of ~41 k particles was refined to an overall resolution of 3.4 Å (Fig. 2a, Supplementary Figs. 3–5). As expected, the resulting NaV1.6EM/β1/β2 structure closely resembles the reported structures of human NaV channels due to high sequence similarity (Fig. 2b). For example, the binding modes of the β subunits are consistent with the structures of human NaV1.7/β1/β2 and NaV1.3/β1/β241,44; the pore-forming α-subunit of NaV1.6EM can be well superimposed with NaV1.7 with a backbone (1107 Cα) root mean square deviation (RMSD) of 1.4 Å (Fig. 2c). However, marked local conformational differences were observed between the two structures, especially in the extracellular loops (ECLs) (Fig. 2c, d). The ECLs are less conserved regions among the nine NaV channel subtypes (Supplementary Fig. 6a), which form the outer mouth of the selectivity filters (SFs) and contribute to the binding of β subunits. Superposition of the Domain I ECLs of NaV1.6EM and NaV1.7 shows that the ECLI of NaV1.6EM lacks the short α2 helix, but instead forms an extended hairpin-like turn (Fig. 2d). Importantly, the ECLI of NaV1.6EM exhibits more N-linked glycosylation modification sites than NaV1.7; N308-linked glycosylation site appears to be unique for NaV1.6 based on the sequence alignment (Supplementary Fig. 6a). Although these structural differences in the ECLs do not affect the binding of β subunits to NaV1.6 (Fig. 2a), the glycosylation and other modifications shape the surface properties of NaV1.6, which play important roles in its trafficking, localization, and pathology59,60. For instance, a unique glycosylation site in the ECLI of NaV1.5 blocks the binding of the β1 subunit to NaV1.543.
a, b The cryo-EM density map (a) and cartoon representation (b) of the NaV1.6EM/β1/β2 complex. c Structural comparison of NaV1.6EM and NaV1.7 (PDB code: 7W9K, colored in gray). The black dashed-line squares indicate the areas shown in panels (d), (e), and (f). d Superimposition of the ECLI between NaV1.6EM and NaV1.7. N-linked glycosylation moieties are shown in sticks. e Comparison of the IFM motif. The IFM motif were depicted side chains in sticks and spheres with half transparency. f Comparison of the intracellular activation gate of NaV1.6EM and NaV1.7 viewed from intracellular side. Key residues from four S6 helices were shown side-chains sticks and spheres with half transparency.
We next compared the fast inactivation gate and intracellular activation gate between NaV1.6EM and NaV1.7, which only display subtle conformational shifts (Fig. 2e, f), indicating that those key structural elements are highly conserved to fulfill their similar biological roles. Consistently, the signature fast inactivation gate, Ile-Phe-Met motif (IFM-motif), binds tightly to its receptor site adjacent to the intracellular activation gate (Fig. 2e), resulting in a non-conductive activation gate constricted by A411, L977, I1464, and I1765 from the four S6 helices, respectively (Fig. 2f). The van der Waals diameter of the activation gate is less than 6 Å, suggesting that the gate is functionally closed (Fig. 3a, b).
a The ion conductance path of NaV1.6EM calculated by HOLE. The diagonal repeats of pore domain only including the S5–S6 and pore-helices were shown for clarity. b Plot of the pore radii of NaV1.6EM. The dashed line indicates pore radius at 2 Å. The key residues constituting the selectivity filter (SF) and the intracellular activation gate (AG) were shown as sticks. c The SF of NaV1.6EM viewed from the extracellular side. Potential Na+ ions were shown as blue balls. Black dashed lines represent polar interactions. d, e Comparison of the Na+ binding sites of NaV1.6EM and the Ca2+ binding sites of CaVAb (PDB code: 4MS2, colored in gray). The diagonal repeats of DI and DIII (d), DII and DIV (e) are shown separately for clarity. The EM densities for putative Na+ and key residues are shown in orange meshes contoured at 4 σ and 5 σ, respectively. A third possible Na+ ion with weaker density contoured at 3 σ was shown as a light blue ball with half transparency. Ca2+ ions are shown as green balls.
Potential Na+ sites in the SF
The ion path of NaV1.6 has two constriction sites, the extracellular SF and intracellular activation gate, respectively (Fig. 3a, b). The sodium selectivity of mammalian NaV channels is determined by the extracellular SF61,62, which is composed of an Asp from DI, Glu from DII, Lys from DIII, and Ala from DIV, known as the DEKA-locus63,64. Based on structural analysis, the acidic residues of the DEKA-locus are believed to act as a high-field strength site, which attracts and coordinates Na+; and the Lys in DIII was proposed as a favorable binding ligand for Na+ which facilitates the ions passing through the SF43,65. In coincidence with other mammalian NaV channels43,44, the SF of NaV1.6EM adopts an asymmetric conformation composed of the DEKA-locus (Fig. 3b, c). No oblivious Na+ binding site had been identified in previous structures of mammalian NaV channels. In contrast, densities for Ca2+ were consistently reported in the structures of bacterial CaVAb channel and mammalian CaV1.1, CaV2.2, and CaV3.1 channels66,67,68,69. Interestingly, two strong blobs of EM densities were observed in the SF of NaV1.6EM (Fig. 3d, e), which are deduced as potential Na+ binding sites because Na+ ions are the only major cations in the solutions throughout the purification processes. The upper site (namely Na1) closely engages E936 of the DEKA-locus and an additional acidic residue E939 (Fig. 3c). The distances of this Na1 to the E936 and E939 are at ~3.5 Å, suggesting that Na+ in Na1 site may still be hydrated. Meanwhile, D370 of the DEKA-locus contributes minorly to this Na+ binding site at a distance of ~7.5 Å (Fig. 3c). This observation is in line with previous studies showing that E936/K1413 of the DEKA-locus are the most prominent residues for Na+ permeation and selectivity, while D370 of the DEKA-locus is not absolutely required63. This potential Na1 site may represent the first step for Na+ conductance, that is, E936 of the DEKA-locus attracts and captures one hydrated Na+ from the extracellular solution with the assistance of E939. The second blob of density is located inside the SF, namely the Na2 site, which is about ~5.3 Å away from the Na1 site (Fig. 3d, e). Interestingly, the Na2 is close to the short side-chain residue A1705 of the DEKA-locus and is coordinated with the strictly conserved E373 at a distance of ~3.3 Å (Fig. 3c, d). We also noticed that D370/E936 of the DEKA-locus contribute negligibly to the Na2 at distances of 5.6–6.6 Å (Fig. 3c). Thus, we hypothesize that the Na2 may represent the second step for sodium conductance, that is, after captured and partially dehydrated in Na1 site, at least partially-dehydrated Na+ can fit into the Na2 site which is going to enter the narrowest asymmetric constriction site of the SF. The possible partial dehydration of Na+ in the Na2 site is reflected by its relatively weaker density compared to the Na1 (Fig. 3d, e). Furthermore, the K1413 points its long side-chain deep into the SF, forming the narrowest part of the SF. It has been proposed that this residue serves as a key coordination ligand in favor of Na+ or Li+ but is unfavorable for other cations43. In line with this hypothesis, Na+ from the Na2 site can quickly pass through the SF and enter the central cavity accelerated by the amino group of the K1413. We found additional elongated density below the K1413 at a distance of ~3.5 Å, which may represent a third Na+ site (namely Na3) (Fig. 3d, e). Consistently, previous MD simulations studies suggested that two Na+ ions spontaneously occupy the symmetric SF of the bacterial NaV channels, and three Na+ sites were proposed in the asymmetric SF of the eukaryotic NaV channel70,71,
Discussion
In this study, we presented cryo-EM structures of human NaV1.6/β1/β2 apo-form and complexed with the NaV1.6 preferred blocker 4,9-ah-TTX. To facilitate the structural studies, we obtained the core construct of NaV1.6EM which displayed improved sample quality. This construct and the structures can be a useful tool for future NaV1.6-related structural and biochemical studies. The apo-form NaV1.6 structure reveals three potential Na+ sites, which are coordinated by the important residues in the SF, suggesting a possible mechanism for Na+ recognition, selection, and conductance. By comparison with the Ca2+ sites in the bacterial and mammalian CaV channels66,67,68,69, the unique asymmetric SF of mammalian NaV channels provides a precise tunnel to separate Na+ from other cations. However, the exact hydration state of those potential Na+ sites cannot be identified here due to the resolution limit. Future high-resolution structure of NaV1.6 would be required to investigate more detailed mechanisms of Na+ conductance. The 4,9-anhydro-TTX bound NaV1.6 structure demonstrated that 4,9-anhydro-TTX and its closely-related analog TTX share a similar binding pocket, which is composed of nearly identical residues above the SFs. However, TTX has greater potency than 4,9-anhydro-TTX in inhibiting NaV1.6 very likely due to TTX can form two additional hydrogen bonds with NaV1.6. Our MD simulations show that 4,9-anhydro-TTX exhibits a more stable binding mode and greater binding energy with NaV1.6 than NaV1.7. Specifically, the increased flexibility of E930 and E927 may cause the loose binding of 4,9-anhydro-TTX to NaV1.7. Those results potentially explain the higher potency of TTX to NaV1.6 than other TTX-sensitive NaV channels and the favorable inhibition of NaV1.6 by 4,9-anhydro-TTX. In addition, an interesting observation needed to be mentioned here is the existence of some differences between the binding poses of 4,9-anhydro-TTX in the NaV1.64,9-ahTTX EM structure and our MD simulation models (Fig. 4f, Supplementary Fig. 8f). The MD study was conducted with the assumption that the NH group of guanidine in 4,9-anhydro-TTX is fully protonated into NH2+. However, since such NH in the EM structure is only ~3 Å from the amine group of Y371, it implies an uncertainty of the protonation state of the guanidine of 4,9-anhydro-TTX. When we performed another MD study using unprotonated 4,9-anhydro-TTX and found that the ligand adopts a similar binding pose as observed in the EM structure. Our findings on the protonation state of 4,9-anhydro-TTX binding with NaV1.6 requires further systemic investigation. Taken together, our results provide important insights into NaV channel structure, Na+ selectivity and conductance, and modulation by TTX and its analog 4,9-anhydro-TTX.
Methods
Whole-cell recordings
HEK293T cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM, Gibco, USA) supplemented with 15% Fetal Bovine Serum (FBS, PAN-Biotech, Germany) at 37 °C and 5% CO2. The P2 viruses of NaV1.6EM and NaV1.6 variants were obtained using Sf9 insect cells and used to infect HEK293T cells for 9 h. The plasmids expressing NaV1.2WT or NaV1.7WT were transfected into HEK293T cells using lipofectamine 2000 (Thermo Fisher Scientific, USA). 12–24 h after transfection or infection, whole-cell recordings were obtained using a HEKA EPC-10 patch-clamp amplifier (HEKA Electronic, Germany) and PatchMaster software (HEKA Electronic, Germany). The extracellular recording solution contained (in mM): 140 NaCl, 3 KCl, 1 CaCl2, 1 MgCl2, 10 Glucose, and 10 HEPES (310 mOsm/L, pH 7.30 with NaOH). The recording pipette intracellular solution contained (in mM): 140 CsF, 10 NaCl, 1 EGTA, and 10 HEPES (300 mOsm/L, pH 7.30 with CsOH). The pipettes were fabricated by a DMZ Universal Electrode puller (Zeitz Instruments, Germany) using borosilicate glass, with a resistance of 1.5–2.5 MΩ. The currents were acquired at a 50 kHz sample rate and series resistance (Rs) compensation was set to 70–90%. All experiments were performed at room temperature.
Data analyses were performed using Origin 2020b (OriginLab, USA), Excel 2016 (Microsoft, USA), and GraphPad Prism 9.1.1 (GraphPad Software, USA). Steady-state fast inactivation (I–V) and conductance-voltage (G–V) relationships were fitted to Boltzmann equations:
where I is the peak current, G is conductance, Vm is the stimulus potential, V1/2 is the half-maximal activation potential, ENa is the equilibrium potential, and k is the slope factor.
To assess the potency of 4,9-anhydro-TTX and TTX on NaV channels, HEK293T cells were held at −120 mV and the inward sodium currents were elicited by a 50-ms step to −10 mV with a low frequency of 1/15 Hz. The concentration-response curves were fitted to a four-parameter Hill equation with constraints of Bottom = 0 and Top = 1:
where Y is the value of IDrug/IControl, Top is the maximum response, Bottom is the minimum response, X is the lg of drug concentration, and IC50 is the drug concentration producing the half-maximum response. The significance of fitted IC50 values compared to the control was analyzed using the extra sum-of-squares F test. The drug inhibition statistics are presented in Supplementary Table 1.
NaV1.6/β1/β2 cloning and expression
The DNA fragments encoding human NaV1.6 (UniProt ID: Q9UQD0), β1 (Uniprot ID: Q07699), and β2 (Uniprot ID: O60939) were amplified from a HEK293 cDNA library. The full-length or truncated NaV1.6, β1, and β2 genes were cloned into the pEG BacMam vector, respectively. For NaV1.6EM, residues of inter-domain linkers 478–692, 1115–1180, and 1932 to the last residue were deleted by PCR to optimize the biochemical properties of the purified protein sample. Specifically, NaV1.6EM was fused before a PreScission Protease recognition site, which is succeeded by a mCherry fluorescent protein and a Twin-Strep II tag at the C terminus. A superfolder green fluorescent protein (sfGFP) and His10 tag were introduced at the C terminus of β1. The sequences of all primers used in this study are provided in Supplementary Table 2. For protein expression, recombinant baculoviruses were generated in Sf9 cells using the Bac-to-Bac baculovirus expression system (Invitrogen, 11496015). HEK293F (Gibco, 11625019) cells were cultured under 5% CO2 at 37 °C and were used for transfection at a density of 2.5 × 106 cells/ml. The NaV1.6EM, β1, and β2 viruses were co-transfected into HEK 293F cells at a ratio of 1% (v/v) supplemented with 1% (v/v) FBS. After 8–12 h, sodium butyrate was added into the culture at a final concentration of 10 mM, and the cell was incubated for another 48 h under 30 °C. Cells were then harvested by centrifugation at 1640 × g for 5 min, and finally stored at −80 °C after freezing in liquid nitrogen.
Purification of human NaV1.6/β1/β2 complex
The NaV1.6/β1/β2 complex was purified following a protocol as was applied in the purification of the NaV1.3/β1/β2 complex41. Cells expressing NaV1.6EM complex were resuspended in buffer A (20 mM HEPES pH 7.5, 150 mM NaCl, 2 mM β-mercaptoethanol (β-ME), aprotinin (2 μg/mL), leupeptin (1.4 μg/mL), pepstatin A (0.5 μg/mL)) using a Dounce homogenizer and centrifuged at 100,000 × g for 1 h. After resuspension in buffer B (buffer A supplemented with 1% (w/v) n-Dodecyl-β-D-maltoside (DDM, Anatrace), 0.15% (w/v) cholesteryl hemisuccinate (CHS, Anatrace), 5 mM MgCl2 and 5 mM ATP), the suspension was agitated at 4 °C for 2 h and the insoluble fraction was removed by centrifugation again at 100,000 × g for 1 h. The supernatant containing solubilized NaV1.6EM was then passed through Streptactin Beads (Smart-Lifesciences, China) via gravity flow at 4 °C to enrich the protein complex. The resin was subsequently washed with buffer C (buffer A supplemented with 0.03% (w/v) glycol-diosgenin (GDN, Anatrace)) for 10 column volumes. The purified NaV1.6EM complex was eluted with buffer D (buffer C plus 5 mM desthiobiotin (Sigma, USA)) and was subsequently concentrated to 1 mL using a 100 kDa cut-off Amicon ultra centrifugal filter (Merck Millipore, Germany). The concentrated protein sample was further purified by size exclusion chromatography (SEC) using a Superose 6 Increase 10/300 GL (GE Healthcare) column pre-equilibrated with the buffer E (20 mM HEPES pH 7.5, 150 mM NaCl, 2 mM β-ME, 0.007% GDN). Finally, the fractions containing homogeneous-distributed protein particles were collected and concentrated to ~4 mg/mL for cryo-EM sample preparation.
Cryo-EM sample preparation and data acquisition
For the preparation of cryo-EM grids, 300-mesh Cu R1.2/1.3 grids (Quantifoil Micro Tools, Germany) were glow-discharged under H2-O2 condition for 60 s. A droplet of 2.5 μL of purified NaV1.6EM complex was applied to the grid followed by blotting for 4–5 s at 4 °C under 100% humidity using a Vitrobot Mark IV (Thermo Fisher Scientific, USA). In the case of the preparation of NaV1.6EM complex with 4,9-anhydro-TTX, 50 μM 4,9-anhydro-TTX (Tocris, UK) was added to the sample before vitrification. Cryo-EM data were collected on a 300-kV Titan Krios transmission electron microscope (Thermo Fisher Scientific, USA) equipped with a Gatan K2 Summit Direct Electron Detector (Gatan, USA) located behind the GIF quantum energy filter (20 e-V). SerialEM81 was used to collect movie stacks at a magnification of ×130,000 (1.04 Å pixel size) with a nominal defocus range from –1.2 to –2.2 μm. A total dose of 50–60 e−/Å2 was acquired for each movie stack under a dose rate of ~9.2 e-/(Å2s) and dose-fractionated into 32 frames. A total of 3,985 and 2929 movie stacks were collected for the apo- and 4,9-anhydro-TTX-bound NaV1.6 complex, respectively.
Data processing
For the data processing of apo and 4,9-anhydro-TTX-bound NaV1.6 complex, a similar procedure was performed and a detailed diagram was presented in Supplementary Figs. 3 and 4. All the data were processed in RELION3.082 or cryoSPARC83. Movies were motion-corrected and dose-weighted using MotionCor2. Contrast transfer function (CTF) estimation was performed with GCTF84. Particles were picked using the AutoPick tool in RELION with templates and extracted into 256 × 256-pixel boxes. Several rounds of 2D and 3D classifications were performed to remove junk particles, followed by 3D autorefine, Bayesian polish, and CTF refinement to improve the map quality. The final EM density maps were generated by the non-uniform (NU) refinement in cryoSPARC and reported at 3.4 Å and 3.3 Å, respectively, according to the golden standard Fourier shell correlation (GSFSC) criterion.
Model building
The sequence of human NaV1.6 and NaV1.7 were aligned using Jalview85, and a homology model of NaV1.6 was generated using the molecular replacement tool in PHENIX86. The atomic models of β1 and β2 subunits were extracted from the structure of NaV1.7 (PDB ID: 6J8I). All of the models were fitted into the cryo-EM map as rigid bodies using the UCSF Chimera87. Restraints for 4,9-anhydro-TTX were derived by eLBOW in PHENIX and examined in Coot88. All residues were manually checked and adjusted to fit the map in Coot and were subsequently subjected to rounds of real-space refinement in PHENIX. Model validation was performed using the comprehensive validation (cryo-EM) in PHENIX. The cryo-EM data collection, refinement, and model validation statistics are presented in Supplementary Table 3.
All figures were prepared with UCSF ChimeraX89 or PyMOL (Schrödinger, USA)90.
Molecular dynamics simulations
The structures and force fields for protein, DMPC lipids, and ligands were prepared using the CHARMM-GUI website. The Amber ff14SB force field was used for both protein and lipids with the TIP3P model for water molecules91. The GAFF2 force field parameters were used for the ligands92. The simulated systems were solvated in water with 150 mM NaCl. The energy minimization was performed using the steepest descent method, followed by six equilibrium steps. During the 2 ns equilibrium steps, the protein backbone atoms were restrained to their initial positions using a harmonic potential with a force constant of 1 kcal mol−1 Å−2 and the restraints were subsequently removed. Berendsen’s coupling scheme was used for both temperature and pressure93. Water molecules and all bond lengths to hydrogen atoms were constrained using LINCS94. Finally, six independent production runs were performed for 100 ns. The overall temperature of the system was kept constant, coupling independently for protein, lipids, and solvents at 303.15 K with a Nose-Hoover thermostat95. A constant pressure of 1 bar was maintained using a Parrinello–Rahman barostat in a semi-isotropic coupling type for x/y, and z directions, respectively96. The temperature and pressure time constants of the coupling were 1 and 5 ps, and the compressibility was 4.5 × 10−5 bar−1 for pressure. The integration of the equations of motion was performed by using a leapfrog algorithm with a time step of 2 fs. Periodic boundary conditions were implemented in all systems. A cutoff of 0.9 nm was implemented for the Lennard–Jones and the direct space part of the Ewald sum for Coulombic interactions. The Fourier space part of the Ewald splitting was computed by using the particle-mesh-Ewald method97, with a grid length of 0.12 nm on the side and a cubic spline interpolation.
The binding affinities were calculated by MM/GBSA method98,99,100,101. The MM part consists of the bonded (bond, angle, and dihedral), electrostatic, and van der Waals interactions. The solvation free energies were obtained by using the generalized Born model (GB part), and the non-polar term is obtained from a linear relation to the solvent-accessible surface area (SA part). For each independent trajectory, the first 20 ns trajectory was discarded and 800 frames from 20–100 ns were used for MM/GBSA calculations. The final binding affinity for each ligand-protein complex was obtained by taking the average of the six independent trajectories. Regarding the clustering analysis, structure alignment was first performed for each two of the structures in the trajectory by using Least Squares algorithm which aligns two sets of structure by rotating and translating one of the structures so that the RMSD between matching atoms of the two structures is minimal. Then the clustering analysis was performed by using GROMOS102 with a RMSD cut-off of 1.5 Å to determine the structurally similar clusters. All the simulations were performed using the GROMACS 2021 suite of programs103.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.