Abstract
Sustained ryanodine receptor (RyR) Ca2+ leak is associated with pathological conditions such as heart failure or skeletal muscle weakness. We report that a single session of sprint interval training (SIT), but not of moderate intensity continuous training (MICT), triggers RyR1 protein oxidation and nitrosylation leading to calstabin1 dissociation in healthy human muscle and in in vitro SIT models (simulated SIT or S-SIT). This is accompanied by decreased sarcoplasmic reticulum Ca2+ content, increased levels of mitochondrial oxidative phosphorylation proteins, supercomplex formation and enhanced NADH-linked mitochondrial respiratory capacity. Mechanistically, (S-)SIT increases mitochondrial Ca2+ uptake in mouse myotubes and muscle fibres, and decreases pyruvate dehydrogenase phosphorylation in human muscle and mouse myotubes. Countering Ca2+ leak or preventing mitochondrial Ca2+ uptake blunts S-SIT-induced adaptations, a result supported by proteomic analyses. Here we show that triggering acute transient Ca2+ leak through RyR1 in healthy muscle may contribute to the multiple health promoting benefits of exercise.
Similar content being viewed by others
Introduction
Ryanodine receptors (RyRs) encode intracellular Ca2+ release channels located on the endo/sarcoplasmic reticulum (ER/SR)1. In mammals, there are three isoforms. RyR1 and RyR2 are involved in skeletal and cardiac muscle excitation–contraction coupling, respectively2, and are also found in non-muscle tissues3, while RyR3, originally identified in the brain4, is expressed in many tissues5. RyRs are >2 million dalton homotetramers and are the largest known ion channels. RyR1 and RyR2 are stabilized by two forms of FKBP-encoded proteins: calstabin (calcium channel stabilizing binding protein, referring to its role in calcium homeostasis) 1 and calstabin2 (FKBP12 and FKBP12.6, respectively)6. The function of RyRs is further modulated by kinases7, phosphatases8, phosphodiesterases7,9 and calmodulin7. RyR oxidation, phosphorylation and nitrosylation can lead to dissociation of calstabin from the channel, leading to Ca2+ leaking from the ER/SR7. Sustained leak can excessively elevate cytosolic and mitochondrial Ca2+, inducing increased ROS production, mitochondrial dysfunction and cell damage10. This has been linked to pathological conditions such as heart failure a–c Proteomic analysis of protein groups related to Gene Ontology Biological Processes (GoBP), Molecular Function (GoMF) and Cellular Component (GoCC) that are significantly decreased S-SIT after 10 µM S107 treatment for 72 h (the treatment was applied immediately after stimulation). Protein groups exceeding 400 proteins were excluded. The median values of S-SIT − S-SIT S107 difference were calculated, and a score affected to the amplitude of the difference. The positive scores display the pathways significantly inhibited by S107 treatment. n = 5 per group. Benjamini–Hochberg corrected t test. Source data are provided as a Source Data file.
Inhibition of RyR1 Ca2+ leak may also decrease SERCA-mediated Ca2+ uptake and ATP utilization in the cytosol, and that could be a confounding factor for the observed mitochondrial changes in our S-SIT and S-SIT S107 myotubes. Sarcolipin (SLN) is a small molecule that is known to bind to SERCA inhibiting its activity. SERCA uses the energy derived from the hydrolysis of ATP to transport Ca2+ ions across the SR membrane61. SLN binding to SERCA promotes the uncoupling of the SERCA pump and slippage of Ca2+ into the cytoplasm instead of the SR lumen62. Expression of SERCA1 (the main isoform of SERCA expressed in C2C12 myotubes63) and its main regulator in the skeletal muscle, SLN64, was not different between SIT and S-SIT S107 myotubes at 72 h post stimulation (Supplementary Fig. 4b, c). SERCA/SLN co-immunoprecipitation assay showed no significant difference between S-SIT and S-SIT S107 myotubes compared to controls (Supplementary Fig. 4a, d). However, others have shown that SLN decreases SERCA Ca2+ uptake but does not alter ATP hydrolysis65, thus implicating SLN as an uncoupler of SERCA (it continues to hydrolyse ATP but less Ca2+ is transported to the SR lumen)66. We, therefore, prepared microsomes (SR-enriched fractions) (Supplementary Fig. 4e, f) to directly measure SERCA1 ATPase activity. Our results showed increased ATPase activity in S-SIT myotubes compared to control and S-SIT S107 myotubes at 72 h post stimulation (Supplementary Fig. 4g, h), suggesting increased ATP utilization at S-SIT myotube SR. Interestingly, ATP levels in whole-cell lysates showed a similar increase in S-SIT and S-SIT S107 myotubes compared to controls (Supplementary Fig. 4i). The high levels of ATP in S-SIT myotubes, despite the increased ATP utilization at the SR, suggest a higher metabolic state in S-SIT myotubes compared to control and S-SIT S107 myotubes.
S-SIT increases mitochondrial Ca2+ content
Ca2+ is a well-known second messenger involved in many physiological processes in numerous tissues including the skeletal muscle67,68. Ca2+ released through the SR can be taken up by mitochondria through the outer mitochondrial membrane via the voltage-dependent anion channel69,70 to cross the inner mitochondrial membrane via the MCU71. Such mitochondrial Ca2+ flux has been linked to improved mitochondrial bioenergetics72. We, therefore, investigated whether the Ca2+ leak through RyR1 in response to S-SIT induced mitochondrial Ca2+ uptake to trigger positive mitochondrial adaptations. We used the Rhod-2/AM probe and time-lapse confocal live imaging to investigate mitochondrial Ca2+ uptake14 in response to a single bout of S-SIT or S-MICT. Control myotubes loaded with Rhod-2/AM probe did not show any mitochondrial Ca2+ increase in response to light exposure (Fig. 5a), while mitochondrial Ca2+ significantly increased in both S-MICT and S-SIT myotubes (Fig. 5b, c, f). Mitochondrial Ca2+ during S-MICT returned rapidly towards basal levels, while it remained elevated after S-SIT (Fig. 5b, c, g), suggesting Ca2+ accumulation in the mitochondria after S-SIT. S-SIT myotubes pre-treated with 10 μM S107 did not show mitochondrial Ca2+ accumulation (Fig. 5d), indicating that Ca2+ leak through RyR1 is the main source of the observed mitochondrial Ca2+ uptake after S-SIT. A similar result was obtained when S-SIT myotubes were pre-treated with 20 μM of the MCU inhibitor mitoxantrone (MTX)73 (Fig. 5e, g). These results suggest that part of the Ca2+ leaking through RyR1 is directed to the mitochondria through MCU, which supports the decreased SR Ca2+ observed immediately after a SIT session (Fig. 2g–j).
a–e Normalized Rhod-2 fluorescence imaging in C2C12 myotubes. A 300 s time-lapse confocal recording of a bout of slightly modified S-SIT and S-MICT protocols: b S-MICT and c S-SIT. d, e Mitochondrial Ca2+ uptake in S-SIT myotubes pre-treated with 10 µM S107 for 1 h (d) or 20 µM mitoxantrone (MTX) (e). Horizontal black lines indicate the periods of stimulation; vertical red arrows indicate the amplitude of mitochondrial Ca2+ at the end of the recordings. f–g Maximal amplitude of normalized Rhod-2 fluorescence during stimulations (f) and 80 s after the end of S-MICT and S-SIT stimulations (g) in (a–e); n = 5 (CTRL, S-SIT and S-SIT S107) and 3 (S-MICT, S-SIT MTX) independent biological experiments. One-way ANOVA followed by Tukey’s multiple comparisons test. h–j Normalized Rhod-2 fluorescence imaging in mouse FDB intact single muscle fibres. Same protocols as described for (a–c), except that the voltage was 40 V. k–l Maximal amplitude of normalized Rhod-2 fluorescence in FDB muscle fibres in (h–j) during (k) and 80 s after the end (l) of S-MICT and S-SIT stimulations; n = 3 mice per group. One-way ANOVA followed by Tukey’s multiple comparisons test. m–n Representative immunoblots (m) and quantification (n) of phosphorylated PDH E1α at serine 293 related to total PDH E1α in myotubes 1 h after stimulation; 10 µM S107 or 20 µM MTX were applied after the stimulation for 1 h when indicated; n = 7 (CTRL, S-MICT, S-SIT and S-SIT S107) and 3 (S-SIT MTX) independent biological experiments. One-way ANOVA followed by Sidak’s multiple comparisons test. o, p Representative immunoblots (o) and quantification (p) of phosphorylated PDH E1α at serine 293 related to total PDH E1α in human muscles; n = 8 participants per group. Two-way ANOVA followed by Sidak’s multiple comparisons test. q Proposed schematic of RyR1 Ca2+ leak-activated mitochondrial PDH E1α dephosphorylation in response to S-SIT. Data are mean ± SD. *p ≤ 0.05, **p ≤ 0.01 and ***p ≤ 0.001. Source data are provided as a Source Data file.
To confirm S-SIT-induced mitochondrial Ca2+ accumulation in mature skeletal muscle, we performed mitochondrial Ca2+ uptake measurements in dissociated mouse FDB muscle fibres. We applied S-MICT and S-SIT to FDB muscle fibres and followed mitochondrial Ca2+ accumulation after the end of the stimulation as in the C2C12 myotubes. Control muscle fibres showed no increase in mitochondrial Ca2+ with light exposure (Fig. 5h). Like in C2C12 myotubes, S-MICT and S-SIT resulted in altered mitochondrial Ca2+ transients in FDB muscle fibres (Fig. 5i, j, k). Again, the amount of Ca2+ taken up by the mitochondria at the end of the S-SIT session was higher than at the end of the S-MICT session (Fig. 5l).
We then examined whether RyR1 Ca2+ leak elicits enough Ca2+ domains to trigger mitochondrial Ca2+ uptake through the MCU in our in vitro models. Mitochondrial Ca2+ uptake is known to occur in specific conditions, which require local microdomains of elevated Ca2+ between the SR and the mitochondria74. Despite the consensus that MCU needs a Ca2+ concentration in the micromolar range, there is evidence that mitochondria also can take up Ca2+ at nanomolar concentrations75,76. We hypothesized that such Ca2+ domains are formed in response to leaky RyR1 and can trigger mitochondrial Ca2+ uptake. We first checked the pattern of RyR1 opening-induced mitochondrial Ca2+ accumulation in our myotubes by treating them with 2.5 mM caffeine while monitoring mitochondrial Ca2+ with the Rhod-2/AM probe. Caffeine induced a rapid, large mitochondrial Ca2+ increase, which progressively decreased, upon which the addition of 10 μM of rapamycin (which dissociates calstabin1 from RyR114) induced a small additional mitochondrial Ca2+ uptake (Supplementary Fig. 4j). When the myotubes were first treated with 10 μM rapamycin, they also showed a rapid mitochondrial Ca2+ increase, but with a lower amplitude than that of caffeine treatment (Supplementary Fig. 4k). This mitochondrial Ca2+ uptake showed a progressive increase that reached a plateau, and the addition of caffeine elicited no further increase in mitochondrial Ca2+ (Supplementary Fig. 4k). These results suggest that rapamycin-induced leaky RyR1 is sufficient to elicit a progressive and sustained mitochondrial Ca2+ uptake. Overall, our observations in C2C12 myotubes and mouse FDB fibres indicate that, in response to S-SIT, a leaky RyR1 channel leads to Ca2+ uptake by the mitochondria. While electrical stimulation is known to elicit mitochondrial Ca2+ uptake72, we here show a specific mitochondrial Ca2+ accumulation following S-SIT, after the electrical stimulation ended, which suggests a role of RyR1 Ca2+ leak rather than Ca2+ release in response to electrical stimulation.
Mitochondrial Ca2+ uptake induced by RyR1 Ca2+ leak dephosphorylates PDH and underpins OXPHOS complex I increase in response to S-SIT
Several mitochondrial enzymes are Ca2+ sensitive. Among them, PDH is a gateway enzyme for carbohydrate-derived pyruvate entry into the TCA cycle for complete oxidation77. PDH catalyses pyruvate decarboxylation to acetyl-CoA, and the reaction leads to a reduction of NAD+ to NADH. PDH activity is covalently regulated by phosphorylation on four residues of the PDH E1 subunit: Ser293, Ser295, Ser300 and Ser232. PDH kinases inactivate PDH by phosphorylation, whereas PDH phosphatases activate PDH by dephosphorylation78. PDH activity is regulated in human muscle in response to exercise79. The following arguments prompted us to investigate PDH phosphorylation in response to SIT: (i) skeletal muscle PDH activity is dependent on exercise intensity80 and MCU−/− mice showed marked impairment in their ability to perform strenuous work81; (ii) MCU knockout induced an increase in PDH phosphorylation similarly to PDH phosphatases knockout linking mitochondrial Ca2+ uptake to PDH phosphorylation levels82.
We, therefore, measured PDH phosphorylation levels as a readout of the increased mitochondrial Ca2+ uptake in S-SIT myotubes. PDH phosphorylation level on Ser293 (P PDH E1α Ser293) was significantly decreased in S-SIT myotubes at 1 h post stimulation, but not in S-MICT myotubes (Fig. 5m, n), reflecting SIT-induced mitochondrial Ca2+ uptake and PDH activation. PDH dephosphorylation was also observed in our human muscle biopsies collected post SIT—but not MICT (Fig. 5o, p), indicating again that our S-SIT model mirrors the effects of a single SIT session in human muscle. To investigate the potential role of RyR1 Ca2+ leak in the process, we treated S-SIT myotubes with 10 μM S107 immediately after the stimulation and for 1 h before measurement of PDH phosphorylation levels. The decreased PDH phosphorylation was blunted (Fig. 5m, n), pointing to a causal role of RyR1 Ca2+ leak in PDH dephosphorylation in response to S-SIT. S-SIT myotube treatment with 20 μM MTX immediately after the stimulation and for 1 h also restored PDH phosphorylation levels, confirming the role of mitochondrial Ca2+ uptake in the process (Fig. 5m, n). It follows that SIT induces RyR1 PTMs and calstabin1 dissociation from the RyR1, which leads to a leaky RyR1, mitochondrial Ca2+ uptake and PDH dephosphorylation (Fig. 5q).
We finally investigated whether RyR1 Ca2+ leak-induced mitochondrial Ca2+ uptake plays a role in the late (72 h post) mitochondrial adaptations in response to S-SIT. For this purpose, we transfected S-SIT myotubes with small interfering RNAs (siRNAs) directed against MCU immediately after the stimulation and measured OXPHOS proteins 72 h later. We first confirmed that the si-MCU was effective to decrease MCU expression and mitochondrial Ca2+ uptake (Fig. 6a, b) in differentiated myotubes without altering the expression of the main proteins involved in SR Ca2+ transients (RyR1, SERCA, and calstabin1) (Fig. 6a). S-SIT myotubes transfected with the si-MCU immediately after the stimulation showed decreased MCU protein levels 72 h after the stimulation as compared to scrambled siRNA transfection (si-CTRL and S-SIT si-CTRL) (Fig. 6d). S-SIT si-CTRL myotubes showed significantly increased OXPHOS CI and IV proteins, whereas S-SIT si-MCU specifically prevented OXPHOS CI modifications (Fig. 6c, e–i). Since we downregulated MCU after S-SIT, our results suggest that mitochondrial Ca2+ uptake after completion of S-SIT (which we linked to the leaky RyR1) is responsible for the increase in mitochondrial OXPHOS CI expression. These results are in agreement with recent work showing an association between decreased MCU levels and a specifically decreased OXPHOS CI protein in cardiac tissues obtained from Barth syndrome patients83. Altogether, these observations point to a new role of MCU and mitochondrial Ca2+ in the regulation of OXPHOS CI that opens an area for further investigations.
a Immunoblots of MCU, RyR1, SERCA1, GAPDH and calstabin1 in si-MCU (siRNAs directed against MCU) compared to si-CTRL (negative control siRNAs) myotubes. b Original recordings of normalized Rhod-2 fluorescence imaging in si-MCU compared to si-CTRL myotubes at 72 h post transfection. c Representative immunoblots of mitochondrial OXPHOS and MCU proteins expression in S-SIT si-MCU myotubes at 72 h post stimulation (cells were transfected immediately after the stimulation with the siRNAs against MCU) compared to S-SIT si-CTRL myotubes (cells were transfected immediately after the stimulation with the negative control siRNAs) and si-CTRL (non-stimulated myotubes transfected with the negative control siRNAs). All the cropped parts of OXPHOS proteins are part of the same blot that is shown in Supplementary Fig. 6. d–i Quantifications of the immunoblots in (c) related to total protein and expressed as % of si-CTRL; n = 6 independent biological experiments per group. One-way ANOVA followed by Tukey’s multiple comparisons test (d, e, h). Data are mean ± SD. *p ≤ 0.05, **p ≤ 0.01 and ***p ≤ 0.001. Source data are provided as a Source Data file.
Discussion
We report RyR1 PTMs followed by calstabin1 dissociation in human muscle in response to a single session of SIT—but not MICT, and in C2C12 myotubes submitted to S-SIT and not S-MICT. This was associated with greater mitochondrial adaptations in human SIT muscle and in S-SIT myotubes. Our in vitro models allowed us to causally link RyR1 Ca2+ leak to mitochondrial remodeling and improved mitochondrial function, since the latter were blunted by the RyR stabilizer S107. In healthy individuals, intense exercise-induced acute RyR1 Ca2+ leak in muscle can therefore be considered beneficial since it triggers mitochondrial proteins increase. This places acute RyR Ca2+ leak in conditions of health apart from sustained RyR Ca2+ leak observed in pathological conditions.
In a previous study, we reported RyR1 fragmentation in response to a single session of SIT in humans, which presumably led to a leaky RyR126. Follow-up research suggested that RyR1 fragmentation depends on subject susceptibility and training status, bringing the notion of responders and non-responders28. Here, we focused on calstabin1 dissociation from the RyR1 as it is a strong and consistent signature of leaky RyR1 channels53. Our working hypothesis was that SIT-induced acute RyR1 Ca2+ leak caused muscle mitochondrial remodeling leading to improved respiratory function. Our results mechanistically suggest that Ca2+ leak through RyR1 in response to S-SIT is taken up by the mitochondria and triggers the dephosphorylation of PDH as (i) S-SIT myotubes accumulate more mitochondrial Ca2+ as compared to S-MICT myotubes, a phenotype abolished by S107 treatment. In addition, (ii) S-SIT myotubes and human SIT muscle showed less PDH phosphorylation as compared to S-MICT and MICT, respectively, and (iii) S107 treatment and MCU inhibition with MTX restored PDH phosphorylation levels in S-SIT. In the skeletal muscle, the SR and mitochondria are spatially coupled69 and Ca2+ released from the SR can be taken up by the mitochondria72,84,85. Muscle cell depolarization enhances mitochondrial Ca2+ uptake. This is linked to mitochondrial bioenergetics since RyR1 inhibition during muscle cell depolarization decreases ATP-linked O2 consumption72. In both our S-MICT and S-SIT models, mitochondrial Ca2+ amplitude increased during electrical stimulation and then decreased after the stimulation. While the decrease was almost complete in S-MICT, S-SIT myotubes showed a small but sustained mitochondrial Ca2+ plateau that could not be linked to muscle cell depolarization, and which likely activated the mitochondrial Ca2+-dependent enzyme PDH. The early PDH dephosphorylation observed after S-SIT could not be directly linked to the late mitochondrial adaptations (increased OXPHOS proteins and respiration after 72 h), especially as our respiration data were collected using saturating substrate conditions. However, a role of RyR1 Ca2+ leak in mitochondrial NADH-linked adaptations is supported by (i) our proteomic data showing alteration of protein groups linked to PDH activity upon treatment with S107 (Supplementary Data 1 and 2) and (ii) decreased NADH levels in S-SIT S107 myotubes. MCU downregulation in myotubes blunted the increase in OXPHOS CI proteins (NDUFB8) in response to S-SIT, also pointing to a specific role of RyR1 Ca2+ leak in S-SIT-induced adaptations. These observations combined with the S-SIT S107 data strongly support a beneficial role of RyR1 Ca2+ leak and mitochondrial Ca2+ uptake in the mitochondrial adaptations in response to S-SIT.
Jain et al.86 previously reported that high-fat feeding increased mitochondrial ROS production triggering RyR1 S-nitrosylation and CaMK phosphorylation in the muscle cell, which was associated with increased mitochondrial biogenesis. We found different patterns of RyR1 PTMs but similar CaMK phosphorylation levels in response to SIT and MICT. During muscle contraction, the large Ca2+ release from the SR strongly and quickly induces CaMK phosphorylation and activity87, which supports the similar increase in CaMKII phosphorylation levels in SIT and MICT. Moreover, Wright et al.38 previously reported a CaMK-dependent p38 activation and PGC-1α expression in response to rising cytosolic Ca2+. This also supports our data showing similar increases in CaMKII phosphorylation levels and PGC-1α expression in response to SIT and MICT. Despite these similarities between SIT and MICT, our study points to a specific causality between RyR1 Ca2+ leak-induced mitochondrial Ca2+ uptake and activation of NADH-linked mitochondrial adaptations in response to a single session of SIT.
It has been advanced that HIIT-induced mitochondrial adaptations would require several days/bouts of exercise to induce beneficial adaptions88. In a recent study, Skelly et al. reported acute induction of mitochondrial biogenesis markers in SIT and MICT human muscles fibres irrespective to muscle fibre type, while a long-term training revealed differential muscle fibre-dependent adaptations89. An independent study by Trewin et al.31 reported changes in mitochondrial protein levels and mitochondrial respiration in response to a single session of HIIT and MICT, despite a lower workload in HIIT. Moreover, a rapid PGC-1α protein nuclear localization has been reported in response to SIT compared to MICT90. In our study, although both SIT and MICT showed increases in PGC-1α protein in human muscle and cells, only SIT showed significant increases in OXPHOS CI and CII proteins. As PGC-1 nuclear localization is important for its function, this may be differently regulated in SIT and MICT. Furthermore, whether the acute mitochondrial inductions we observed would translate into long-term adaptations remains unknown. Several studies have compared metabolic adaptations to several weeks of SIT vs. MICT and have reported similarly increased aerobic capacity (VO2max) and mitochondrial content23,91,92. However, the cellular stress and the resulting metabolic signals for mitochondrial biogenesis were shown to depend on exercise intensity, with limited work suggesting that the increases of mitochondrial content are superior after SIT compared to MICT, at least when matched-work comparisons are made within the same individuals91.
Regular exercise leads to widespread changes in numerous cells, tissues and organs (e.g. skeletal and cardiac muscle, brain, liver, fat) conferring multiple health-promoting benefits17. Physical fitness level in healthy middle-aged men is a strong predictor of mortality and small improvements in physical fitness are associated with a significantly lowered risk of mortality16.
Even though today’s recommendations suggest that any episode of any physical activity conveys benefits17, it is important to determine which time-efficient combinations of exercise (with different type, volume and intensity) are optimal to obtain a given outcome as the response might differ according to the fitness status. For maximal health benefits, especially for healthy fit people, even if the means towards that goal imply regular strenuous physical activity, it should perhaps be not too hard93. A recent human study compared cardio-metabolic health parameters and muscle adaptations upon 12 weeks of HIIT, resistance training or a combination of both. Only HIIT improved cardio-metabolic health parameters, while both HIIT and combined training increased aerobic capacity and skeletal muscle mitochondrial respiration22. Proteomics showed increased proteins groups of the electron transport chain, OXPHOS, TCA cycle and glycolysis after HIIT training22.
Using a maximum intensity variant of interval training, we found that inhibiting RyR1 Ca2+ leak with S107 after a single session of S-SIT decreases protein groups involved in glycolysis, acetyl-CoA metabolism and electron transport chain, muscle contraction and ion (including Ca2+) transport in S-SIT myotubes, thus revealing the importance of RyR1 Ca2+ leak in the mechanisms triggering improved muscle mitochondrial function in response to exercise. This suggests that triggering acute RyR1 Ca2+ leak causes mitochondrial remodeling, but may also play a key role in the myriad of other adaptations to exercise training in the muscle.
This study also has some limitations. In order to mechanistically discover the causal link between RyR1 Ca2+ leak and muscle beneficial adaptations in response to SIT, we developed in vitro SIT and MICT models to generate data that were combined with observations made on humans. Since the focus of this study is mitochondrial adaptations linked to Ca2+ leak through the RyR1, the in vitro models were designed to appropriately reflect the main observed changes in human muscle. While the combined models helped to decipher the mechanisms of the mitochondrial adaptations triggered by RyR1 Ca2+ leak, our in vitro models do not properly represent actual SIT and MICT models in humans. Moreover, our results do not rule out that other mechanisms contribute to the observed adaptations as Ca2+ may not be the only mechanism involved in the process.
In conclusion, SIT is a time-efficient exercise intervention that can counteract the effects of physical inactivity, one of the top health risk factors worldwide. In the present study, we contribute to the understanding of the molecular mechanisms underlying its beneficial effects in muscle. We draw two main conclusions: (i) high-intensity exercise triggers RyR1 post-translational changes leading to an acute leaky state and (ii) this is causally linked to improved mitochondrial remodeling and function of the activated muscle. On a conceptual level, we conclude that acute RyR1 Ca2+ leak in muscle can be considered beneficial and shows clear differences with the sustained RyR Ca2+ leaks observed in pathological conditions. These original mechanistic observations pave the way for new research to further investigate the role of acute RyR Ca2+ leak in other physiological conditions. Whether such acute RyR Ca2+ leak may also play a beneficial role in pathological conditions remains to be investigated.
Methods
Human experiments: general procedures
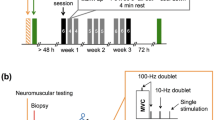
The human general procedures were followed as previously described26 with slight modifications. The study was validated by the Commission d'éthique de la recherche sur l'être humain du Canton de Vaud (protocol 2017-00303) and performed in accordance with the Declaration of Helsinki. Sixteen male recreationally active subjects gave written informed consent before participation. The participants refrained from physical activity and caffeine consumption for 24 and 12 h before the experimental session, respectively. Participants were familiarized with electrical stimulation and voluntary contraction procedures at least 48 h before the first experimental session. In the familiarization session, they performed an incremental test to exhaustion on a cycle ergometer (Lode Excalibur Sport, Lode, Groningen, Netherlands). The test started at a power of 1 W and was increased by 1 W every 2 s. The participants were instructed to maintain a cadence between 60 and 80 r.p.m. and when they were unable to maintain 60 r.p.m. the test was stopped. The mean of the VO2 values in the last 30 s of the test was used to determine VO2peak. Gas exchange was measured with a stationary gas analyser (Quark CPET, COSMED, Rome, Italy). The participants (n = 8 per group) were allocated into two groups (SIT and MICT) based on their VO2peak so that mean VO2peak was similar for each group (Supplementary Fig. 1a).
MICT consisted of 1 h of cycling on a cycle ergometer at 65% of the maximal aerobic power reached during the incremental VO2max test. Each experiment was preceded by a standard warm-up on the cycle ergometer (5 min at 100 W). SIT was comprised of 30 s all-out cycling bouts at 0.7 N.m/kg body mass on a cycle ergometer (Lode Excalibur Sport, Lode, Groningen, Netherlands), with 4 min recovery periods (rest) between bouts.
Knee extensor neuromuscular function of the right (dominant) leg was tested before (Pre), immediately post (Post) and 24 h after (24 h Post) exercise. The tests consisted of a 5 s MVC with a superimposed 100 Hz doublet (paired stimuli) evoked via supramaximal electrical stimulation of the femoral nerve (twitch interpolation technique), followed by supramaximal stimulations of a relaxed muscle evoked at 2 s intervals: a doublet at 100 Hz, 10 Hz, and a single stimulus to obtain the compound muscle action potential (M-wave).
Our study complies in full with the STROBE statement. The schematic of Figs. 1a, 5q are made using the BioRender application: https://app.biorender.com.
Muscle biopsies
Needle biopsies were taken from the left (non-dominant leg) vastus lateralis muscle before, ∼10 min and 24 h after exercise, using previously described and validated procedures94. Briefly, after skin sterilization and local anaesthesia, a 1–2-mm-long skin cut was made with the tip of a scalpel. Biopsies were collected using an automatic biopsy device (Bard Biopsy Instrument, Bard Radiology, Covington, GA, USA). A 14-gauge disposable trocar mounted in the device was inserted through the cut, perpendicular to the muscle fibres, until the fascia was pierced. Three samples (∼15 mg each) were collected from one puncture site at each time point. Muscle samples were immediately frozen in liquid nitrogen and stored at −80 °C until analysis.
C2C12 cell culture
C2C12 mouse skeletal myoblasts were obtained from the American Type Culture Collection and grown in a proliferation medium (PM) composed of Dulbecco’s modified Eagle’s medium (DMEM; Thermo Fisher Scientific, Basel, Switzerland) supplemented with 10% foetal bovine serum (FBS) (Thermo Fisher Scientific, Basel, Switzerland), 100 IU/ml penicillin, 100 μg/ml streptomycin (Thermo Fisher Scientific, Basel, Switzerland) and 1% non-essential amino acids (Thermo Fisher Scientific, Basel, Switzerland), and maintained at 37 °C in a humidified atmosphere with 5% CO2. To induce differentiation, myoblasts were grown to 80–90% confluence, the proliferation medium was then replaced with a differentiation medium (DM), consisting of DMEM supplemented with 2% horse serum (Thermo Fisher Scientific, Basel, Switzerland).
Electrical stimulation of C2C12 myotubes
Well-differentiated C2C12 myotubes (day 6 or 7 post differentiation) in 6-well plates (Corning, NY, USA) containing 4 ml of DM were electrically stimulated (C-Pace EM stimulator, IONOPTIX LLC, MA, USA) as follows: 1 h repeated stimulation at 14 V, 2 Hz and 2 ms pulse duration (S-MICT) or 6 × 30 s pulse (5 s on, 1 s off) at 14 V, 50 Hz and 2 ms pulse duration with 4 min rest (S-SIT) (Fig. 2a).
The DM was replaced before and after the electrical stimulation and the cells were harvested at the appropriate time point. Different treatment conditions from the same plate per independent culture were used and the different wells from the same condition per culture were considered as technical replicates.
Myotube siRNA transfection
C2C12 myotubes at day 6 of differentiation were electrically stimulated using the S-SIT protocol. Stimulated myotubes were immediately transfected with 50 nM final concentration of silencer select negative control siRNAs (si-CTRL) (Thermo Fisher Scientific, Basel, Switzerland) or silencer select pre-designed siRNAs directed against MCU (si-MCU) (Thermo Fisher Scientific, Basel, Switzerland) using Lipofectamine RNAimax protocol according to the manufacturer (Thermo Fisher Scientific, Basel, Switzerland). The myotubes were collected at day 3 post transfection for western blot analysis.
Animals
C57BL/6J (B6) mice obtained from Janvier were provided by the animal facility from the Department of Biomedical Sciences, University of Lausanne. The animals were maintained in a temperature-controlled animal facility with a 12-h light/12-h dark cycle and had access to food and water according to the Swiss Animal Protection Ordinance (OPAn). The 8-week female mice were used in this study. Our protocol was approved by the Animal Ethics Committee of Lausanne (commission cantonale pour l’expérimentation animale) with the number VD3489.
Mouse FDB muscle fibre dissociation
The FDB muscles were isolated from 8-week female C57BL/6J (B6) mice and incubated for 38 min at 37 °C in an oxygenated Krebs-HEPES solution (in mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, CaCl2 1.8, final pH 7.3) containing 0.2% collagenase type IV (Sigma-Aldrich Corp., St. Louis, MO, USA). Muscles were then washed twice in DMEM/Ham’s F12 (Sigma-Aldrich Corp., St. Louis, MO, USA) supplemented with 2% FBS (Sigma-Aldrich, St. Louis, MS, USA) and mechanically dissociated by repeated passages through fire-polished Pasteur pipettes of progressively decreasing diameter. For Ca2+ measurements, dissociated fibres were plated on 35‐mm‐diameter glass-bottom MatTek dishes (MatTek, Ashland, MA, USA) coated with Matrigel (BD Bioscience, San Jose, CA, USA). Culture dishes were kept in an incubator, with 5% CO2 at 37 °C for 2 h to allow the fibres to attach95.
Cytosolic Ca2+ imaging using Fluo-4 AM in C2C12 myotubes
C2C12 myoblasts were plated on poly-d-lysine-coated 35‐mm‐diameter glass‐bottom MatTek dishes (MatTek, Ashland, MA, USA). At 80% of confluence, the myoblasts were differentiated by replacing the PM with DM and myotubes were used at day 5–7 of differentiation. Myotubes were stimulated using simulated MICT or SIT protocols (S-MICT and S-SIT); then, at the indicated time of measurement, they were loaded with the cytosolic Ca2+ indicator Fluo-4 AM (5 µM, Invitrogen, Basel, Switzerland) solubilized in a Krebs solution (in mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, CaCl2 1.8, final pH 7.3) for 20 min in the incubator. Cells were then rinsed twice with a Ca2+-free Krebs solution (in mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, 200 μM Na-EGTA, final pH 7.3). Fluo-4 fluorescence was monitored using a confocal microscope system (Zeiss LSM 5 Live, Oberkochen, Germany; ×40 oil immersion lens; the excitation wavelength was 488 nm and the emitted fluorescence was recorded between 495 and 525 nm). After recording the basal fluorescence, myotubes were stimulated with 2.5 mM (final concentration) caffeine to trigger Ca2+ release from the SR. Zen software 2012 version (Zeiss, Oberkochen, Germany) was used for the acquisition and data were exported to excel files for analysis. The use of the single excitation/emission Fluo-4 dye necessitated normalizing to pre-stimulation values to account for possible differences in dye loading and excitation strength.
SR Ca2+ measurements with D1ER sensor
SR Ca2+ was measured with the ratiometric genetically encoded Ca2+ sensor D1ER as previously described96. Briefly, C2C12-derived myotubes cultured on poly-d-lysine‐treated 35-mm-diameter glass‐bottom dishes (MatTek, Ashland, MA, USA) were transfected with 2 µg of D1ER plasmid, using Lipofectamine 3000 protocol according to the manufacturer (Thermo Fisher Scientific, Basel, Switzerland). Two days after transfection myotubes were washed four times, electrically stimulated and SR Ca2+ measurements were performed at 37 °C in Ca2+-free Krebs solution (in mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, 200 μm Na-EGTA, final pH 7.3). Glass coverslips were inserted in a thermostatic chamber (Life Imaging Services, Basel, Switzerland). Cells were imaged on a DMI6000 B inverted fluorescence microscope, using an HCX PL APO ×40/1.30 numerical aperture oil immersion objective (Leica Microsystems, Wetzlar, Germany) and an Evolve 512 back-illuminated CCD with 16 × 16-pixel camera (Photometrics, Tucson, AZ, USA). Cells were excited at 430 nm through a BP436/20 filter. The two emission images were acquired with BP480/40 and BP535/30 emission filters. Fluorescence ratios were calculated in MetaFluor 7.0 (Meta Imaging Series, Molecular Devices, San Jose, CA, USA) and analysed in Excel (Microsoft, Seattle, WA, USA) and GraphPad Prism 8.3.1 (GraphPad, San Diego, CA, USA). Images were taken every 2 s. Resting D1ER fluorescence (535/480) ratio values fluorescence intensity data were measured before (CTRL) and after the simulated MICT or SIT protocols (S-MICT and S-SIT).
SOCE assessment
C2C12 S-MICT and S-SIT myotubes in MatTek glass‐bottom dishes (MatTek, Ashland, MA, USA) were loaded with the cytosolic Ca2+ indicator Fluo-4 AM (5 µM, Invitrogen, Basel, Switzerland) solubilized in a Krebs solution (in mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, CaCl2 1.8, final pH 7.3) immediately after the stimulation for 20 min in the incubator. Cells were then rinsed twice with a Ca2+-free Krebs solution (in mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, 200 μM, Na-EGTA, final pH 7.3) and mounted to the confocal microscope system (Zeiss LSM 5 Live, Oberkochen, Germany; ×40 oil immersion lens; the excitation wavelength was 488 nm and the emitted fluorescence was recorded between 495 and 525 nm). The cells were stimulated with 1 μM thapsigargin (Sigma, St. Louis, USA) to deplete the SR Ca2+ stores in Krebs Ca2+-free solution. SOCE was investigated by adding the Krebs containing Ca2+ solution to the dishes to reach a final concentration of 1 mM Ca2+.
RyR1 immunoprecipitation and immunoblotting
RyR1 immunoprecipitation was performed as previously described14 with slight modifications. Briefly, muscle biopsies or C2C12 myotubes were isotonically lysed in an ice-cold lysis buffer composed of: 50 mM Tris-HCl (pH 7.4), 150 mM NaCl, 20 mM NaF and 1 mM Na3VO4, and protease inhibitors (100 µl per 5 mg muscle tissue or per myotube well of a 6-well plate). The lysates were sonicated and then centrifuged at 9300 × g at 4 °C for 10 min. The supernatant was collected, and protein concentration was quantified using a BCA Assay Kit (Thermo Fisher Scientific, Basel, Switzerland). An anti-RyR1 antibody (4 μg 5029 Ab, Prof. Marks’ lab, Columbia University, NY, USA) was used to immunoprecipitate RyR1 from 250 μg homogenate of human muscle or cells. The samples were incubated with the antibody in 0.5 ml of a modified RIPA buffer (50 mM Tris-HCl, pH 7.4, 0.9% NaCl, 5 mM NaF, 1 mM Na3VO4, 1% Triton X-100 and protease inhibitors) for 1 h at 4 °C. The immune complexes were incubated with protein A Sepharose beads (Sigma-Aldrich, St. Louis, MS, USA) at 4 °C for 1 h and the beads were washed three times with buffer. Proteins were separated on sodium dodecyl sulfate (SDS)–polyacrylamide electrophoresis gels (4–15% SDS-precast gradient gels to visualize both RyR1 and calstabin1 or 6% homemade gels for RyR1 alone and its post-translational modifications) and transferred onto nitrocellulose membranes for 1 h at 400 mA. After incubation with blocking solution (LI-COR Biosciences, Lincoln NE, phosphate-buffered saline (PBS) v/v) to prevent non-specific antibody binding, immunoblots were developed with rabbit anti-RyR1 (5029, Prof. Marks’ lab, 1:5000), rabbit anti-phospho-RyR1-pSer2844 (Prof. Marks’ lab, 1:5000), rabbit anti-Cys-NO antibody (Sigma-Aldrich, St. Louis, MS, USA, 1:2000) or rabbit anti-FKBP12 (Abcam, Cambridge, UK, 1:2500) antibodies. To determine channel oxidation, the carbonyl groups on the protein side chains were derivatized to 2,4-dinitrophenylhydrazone (DNP-hydrazone) by reaction with 2,4-dinitrophenylhydrazine. The DNP signal on RyR1 was determined by immunoblotting with an anti-DNP antibody (Sigma-Aldrich, St. Louis, MS, USA, 1:2000). Immunoreactive bands were visualized using infrared fluorescence (IR-Odyssey scanner, LI-COR, Lincoln, NE, USA). Band densities were quantified using Image Studio v.5.2.5 (LI-COR, Lincoln, NE, USA). Skeletal muscle cells were treated with 200 μM H2O2 (positive control for RyR oxidation), 250 μM NOC-12 (positive control of RyR nitrosylation) and 5 units PKA/reaction (positive control for RyR phosphorylation). Beads coupled with proteins without antibody were used as an IP negative control. IP purity was further tested by investigating SERCA protein contamination (using the rabbit SERCA-2a antibody, Abcam ab137020) as compared to non-IP samples (results shown in Supplementary Figs. 1 and 2).
Western blot analysis
A lysis buffer containing the following: 20 mM Tris/HCl (pH 6.8), 2 mM EDTA (pH 8), 137 mM NaCl, 10% glycerol, 10% Triton X-100, 10 mM glycerophosphate, 1 mM KH2PO4, 1 mM PMSF, 1 mM NaVO3, 50 mM NaF, 10 mM NaPPi, and a protease inhibitor mixture (Roche, Complete Mini, Basel, Switzerland) was used to resuspend myotube pellets (100 µl/well of a 6-well plate) or human muscle samples (100 µl/5 mg of tissue). The preparation was homogenized with pipette tips for cells or potter for muscles, incubated for 1 h at 4 °C and gently sonicated. Then, nuclei and debris were removed by centrifugation at 9300 × g at 4 °C for 10 min. Protein quantification was assessed using the BCA Kit (Thermo Fisher Scientific, Ecublens, Switzerland). Fifteen to 20 μg of protein were incubated with 2× Laemmli sample buffer containing SDS and 2-mercapto-ethanol (Bio-Rad, Hercules, CA, USA) for 3 min at 95 °C, electrophoresed 1 h on 4–15% SDS-precast gradient gels (Bio-Rad, Hercules, CA, USA) and wet transferred 1 h onto PVDF membranes. Membranes were stained with Red Ponceau (homemade) and total protein bands were quantified using the Image Studio software v.5.2.5 (LI-COR, Lincoln, NE, USA). Then, the Red Ponceau was washed out with PBS and the membranes were saturated 1 h at room temperature with PBS-LI-COR blocking buffer (LI-COR, Lincoln, NE, USA). Blots were incubated overnight with rabbit anti-FKBP12 (Abcam, Cambridge, UK, 1:2500), rabbit anti-SERCA1 (Abcam, Cambridge, UK, 1:1000), mouse anti-total OXPHOS (Abcam, Cambridge, UK, 1:1000), mouse anti-PGC-1-α (Sigma, St. Louis, USA, 1:1000), rabbit anti-GAPDH (Sigma, St. Louis, USA, 1:5000), rabbit anti-PDH E1α phospho-serine 293 (Abcam, Cambridge, UK, 1:1000), mouse anti-PDHα (Abcam, Cambridge, UK, 1:1000), rabbit anti-STIM1 (Sigma, St. Louis, USA, 1:1000), mouse anti-MCU (Sigma, St. Louis, USA, 1:1000), rabbit CaMKII phospho-threonine 286 (Cell Signaling, Leiden, Netherlands, 1:1000), rabbit CaMKII total (Cell Signaling, Leiden, Netherlands, 1:1000). Membranes were washed in PBS-buffered saline-Tween-20 and incubated for 1 h at room temperature with IRDye 680-conjugated donkey anti-mouse or anti-rabbit IgG (LI-COR, Lincoln, NE, USA, 1:10,000) and IRDye 800-conjugated donkey anti-mouse or anti-rabbit IgG (LI-COR, Lincoln, NE, USA, 1: 5000) in blocking buffer. Immunoreactive bands were visualized using infrared fluorescence (IR-Odyssey scanner, LI-COR, Lincoln, NE, USA). Band densities were quantified using Image Studio v 5.2.5 (LI-COR, Lincoln, NE, USA). Protein intensity signal was normalized to that of GAPDH (which was stable across samples and conditions) in human samples, while total protein staining (found as a more representative loading control for cells) was used to normalize protein content quantified in cells. All loading controls (GAPDH or total proteins) were investigated on the same gels of the protein of interest. Whole representative gels for OXPHOS proteins are provided in Supplementary Fig. 5 (for humans) and Supplementary Fig. 6 (for the cells).
The LI-COR system allows the detection of different proteins at the same time (revealed in different channels). When needed, the membranes were stripped using the appropriate LI-COR strip** solution (LI-COR, Lincoln, NE, USA). For human samples, all the protein quantifications were expressed as a percentage of the Pre-MICT values (kept as the reference for analysis of variance (ANOVA) testing). The protein quantifications for the cell samples were reported to that of CTRL or si-CTRL cells. Detailed information on the antibodies used in this study (supplier, catalogue number, clone name and number, lot number and method of validation) is provided in Supplementary data 3.
ATP, NAD and NADH measurements
ATP levels were measured in cell lysates using a luciferase-based assay (Sigma, St. Louis, USA) according to the manufacturer’s instructions. Luminescence was measured using a plate reader (VICTOR Multilabel Plate Reader, Perkin-Elmer) in the presence of the substrate D-luciferin. The light intensity as relative light units was considered as a direct measurement of intracellular ATP concentration.
NAD and NADH levels were determined using the NAD/NADH Assay Kit (Abcam, Cambridge, UK) according to the manufacturer’s instructions. Briefly, cell lysates were used for each enzyme recycling reaction. The fluorescence was measured in a fluorescence microplate (VICTOR Multilabel Plate Reader, Perkin-Elmer) at 540/590 nm. NAD and total NAD/NADH levels were calculated from a standard curve (µM).
Measurement of mitochondrial morphology in C2C12 cells
Myotubes were incubated in 200 nM MitoTracker red (Invitrogen, Basel, Switzerland) in Krebs solution (mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, CaCl2 1.8, final pH 7.3) and protected from light for 15 min at 37 °C. Rhod-2 fluorescence was detected by using a confocal laser scanning microscopy (inverted Zeiss LSM 710 confocal microscope, Oberkochen, Germany) technique to define single layers of cells at ×40 magnification. Mitochondrial shape descriptors and size measurement were determined using Fiji97, an enhanced version of ImageJ software (http://fiji.sc/) as previously described98,99.
Measurement of mitochondrial Ca2+ uptake
To investigate mitochondrial Ca2+ uptake, cultured C2C12 myotubes or dissociated FDB muscle fibres in MatTek glass‐bottom 6-well plates with a bottom coverslip (MatTek, Ashland, MA, USA) were loaded with 1 ml of Krebs solution (mM: NaCl 135.5, MgCl2 1.2, KCl 5.9, glucose 11.5, HEPES 11.5, CaCl2 1.8, final pH 7.3) containing 1 µM of the mitochondrial fluorescent indicator Rhod-2 AM (Invitrogen, Basel, Switzerland) for 1 h at room temperature. Cells were then washed twice with Krebs solution and Rhod-2 fluorescence was measured using a confocal microscope (inverted Zeiss LSM 710 confocal microscope, Oberkochen, Germany) and ×40 oil immersion lens, with excitation at 532 nm and the emitted signal collected through a bandpass filter (540–625 nm). In some conditions, 10 µM of S107 or 20 µM of MTX was mixed into the Rhod-2/AM incubation solution (for 1 h) and maintained during the acquisition. A 180 s bout of S-MICT stimulation pattern (180 s, 14 V, 2 ms, 2 Hz continuous stimulation) or 2 × 30 s bouts separated by 2 min of no stimulation of the S-SIT stimulation pattern (5 s on, 1 s off, 14 V, 2 ms, 50 Hz) protocols were applied to the myotubes using a 300-s time-series live acquisition. These slightly modified S-MICT and S-SIT protocols allowed us to follow the Ca2+ fluxes continuously and to avoid photobleaching. FDB fibres were excited as described for myotubes, with the exception that the voltage was set to 40 V. CTRL myotubes were only exposed to the laser light. Change in Rhod-2 fluorescence was calculated by reporting the peak of fluorescence to the baseline (normalized fluorescence). The amplitude of Rhod-2 normalized fluorescence measured 80 s after the end of the stimulation in each condition was used as an indicator of resting mitochondrial Ca2+ uptake in response to exercise.
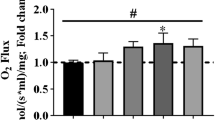
Mitochondrial O2 flux measurements using the Oroboros O2k
High-resolution respirometry measurements were made as previously reported59 and (Gnaiger_2019_MitoFit_Preprint_Arch_doi_10.26124_mitofit_190001). Fully differentiated C2C12 myotubes were stimulated with one session of the S-MICT or S-SIT protocols, and then washed and provided with warm fresh DM. The respiration measurements were performed 3 days after stimulation. In some conditions, cells were incubated in a DM containing 10 µM of S107 applied immediately after electrical stimulation for 72 h (S107 was also added to the MIRO5 buffer during data acquisition). The Oroboros O2k chambers were equilibrated using MIRO5 buffer (mM: EGTA 0.5, MgCl2 3, lactobionic acid 60, taurine 20, KH2PO4 10, HEPES 20, d-sucrose 110, bovine serum albumin fatty acid free 1 g/l, pH 7.3). Then, one well of a 6-well plate (3–4 mg of protein/ml) was trypsinized 3 min with Trypsin/EDTA 0.05% (Invitrogen, Basel, Switzerland), centrifuged for 5 min at room temperature and 212 × g. The pellet was gently washed with a MiR05 buffer and resuspended in 2.5 ml of MiR05 solution and 2 mL of cell suspension inserted in each chamber. Mitochondrial respiration rates were assessed at 37 °C using a slightly modified SUIT-008 protocol of the DatLab 7.3 software (Oroboros, Innsbruck, Austria) (Supplementary Fig. 2p). After stabilization of respiration, the cells were permeabilized using 1 µl of digitonin (50 µg/ml), then successively incubated with 5 µl of malate (2 mM), 10 µl of pyruvate (10 mM) to check leak respiration (MPL). MgCl2 (0.5 mol/mol ADP) associated with 20 µl ADP (500 mM) was then added to assess OXPHOS capacity supported by pyruvate and malate completed with 5 µl glutamate (10 mM), NP. The integrity of the mitochondria was checked after the addition of ADP by using 5 µl of cytochrome C (4 mM). To investigate the activity of NSP, 20 µl of succinate (10 mM) was added to the chamber. The ETS (NSE) was assessed by using a 1 µl titration protocol with FCCP (1 mM); then, the activity of complex I was inhibited by 1 µl of rotenone (1 mM) (SE) and residual oxygen consumption (ROX) was determined by addition of 1 µl of antimycin A (5 mM). An air calibration was performed on each experimental day. Protein quantification was performed in each condition on a representative well using the BCA assay (Thermo Fisher Scientific, Basel, Switzerland). The specific O2 flux per mass was obtained after normalization of ROX-corrected O2 fluxes by mg of protein in the chamber.
Mitochondrial SC quantification
Mitochondrial isolation and BN-PAGE were performed as described in detail elsewhere100. Fully differentiated myotubes were homogenized in 2 ml of cold sucrose isolation buffer with 40 strokes in a Wheaton glass tube at maximum speed. The mitochondrial fraction was isolated by collecting the supernatant after two rounds of centrifugation at 600 × g for 10 min and pelleted at 7000 × g for 10 min. The final pellet was resuspended in 200 µl of cold isolation buffer, and mitochondrial protein levels were quantified using the Lowry method. Fifty micrograms of mitochondrial extracts were solubilized with digitonin (8 g/g—digitonin/protein ratio) and centrifuged at 20,000 × g. The supernatant was collected and loaded into a NativePAGE 3–12% Bis-Tris-Gel (Invitrogen, Basel, Switzerland). After separation of the bands, the mitochondrial proteins were transferred onto a PVDF membrane using an iBlot gel Transfer Device (Invitrogen, Basel, Switzerland), followed by protein fixation with 8% acetic acid. The membrane was blocked, incubated with the OXPHOS primary antibodies (Abcam, Cambridge, UK) and anti-mitochondrial cytochrome C oxidase (Abcam, Cambridge, UK) and further incubated with the secondary mouse antibody (Novex, Thermo Fisher Scientific, Basel, Switzerland). Chromogenic substrate solution (Novex, Thermo Fisher Scientific, Basel, Switzerland) was used for the detection of the bands. After air-drying, the stained membrane was scanned.
Metabolite quantification
Sample preparation
Cells were scraped and extracted by the addition of 500 µl of MeOH:H2O (4:1) per dish. This solution containing lysed cells was further homogenized in the Cryolys Precellys 24 sample Homogenizer (2 × 20 s at 9300 × g, Bertin Technologies, Rockville, MD, USA) with ceramic beads. The bead beater was air-cooled down at a flow rate of 110 L/min at 6 bar. Homogenized extracts were centrifuged for 15 min at 4000 × g at 4 °C (Hermle, Gosheim, Germany) and the resulting supernatant was collected and evaporated to dryness in a vacuum concentrator (LabConco, Missouri, USA). Dried sample extracts were resuspended in MeOH:H2O (4:1, v/v) prior to liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) analysis according to the total protein content.
Protein quantification
The protein pellets were evaporated and lysed in 20 mM Tris-HCl (pH 7.5), 4 M guanidine hydrochloride, 150 mM NaCl, 1 mM Na2EDTA, 1 mM EGTA, 1% Triton, 2.5 mM sodium pyrophosphate, 1 mM beta-glycerophosphate, 1 mM Na3VO4 and 1 µg/ml leupeptin using the Cryolys Precellys 24 sample homogenizer (2 × 20 s at 10,000 r.p.m., Bertin Technologies, Rockville, MD, USA) with ceramic beads. BCA Protein Assay Kit (Thermo Scientific, Massachusetts, USA) was used to measure (A562 nm) total protein concentration (Hidex, Turku, Finland).
LC-MS/MS analysis
Extracted samples were analysed by hydrophilic interaction liquid chromatography coupled to tandem mass spectrometry (HILIC-MS/MS) in negative ionization mode using a 6495 triple quadrupole system interfaced with 1290 UHPLC System (Agilent Technologies)101.
Chromatographic separation was carried out using a SeQuant ZIC-pHILIC (100 mm, 2.1 mm ID and 5 μm particle size, Merck, Darmstadt, Germany) column. The mobile phase was composed of A = 20 mM ammonium acetate and 20 mM NH4OH in water at pH 9.7 and B = 100% acetonitrile. The linear gradient elution from 90% (0–1.5 min) to 50% B (8–11 min) down to 45% B (12–15 min). Finally, the initial chromatographic conditions were established as a post-run during 9 min for column re-equilibration. The flow rate was 300 μl/min, column temperature 30 °C and sample injection volume 2 µl. Electrospray ionization source conditions were set as follows: dry gas temperature 290 °C and flow 14 L/min, sheath gas temperature 350 °C, nebulizer 45 psi, and flow 12 L/min, nozzle voltage 0 V and capillary voltage −2000 V. Dynamic multiple reaction monitoring was used as acquisition mode with a total cycle time of 600 ms. Optimized collision energies for each metabolite were applied.
Raw LC-MS/MS data were processed using the Agilent Quantitative analysis software (version B.07.00, MassHunter Agilent Technologies). For absolute quantification, calibration curves and the stable isotope-labelled internal standards were used to determine the response factor. Linearity of the standard curves was evaluated for each metabolite using a 7-point range. In addition, peak area integration was manually curated and corrected when necessary.
A cut-off was set by the metabolomics platform at 20% change compared to CTRL, over which data variations are considered as relevant.
The raw data related to the metabolite quantification are presented in Supplementary Data 4.
Proteomics data acquisition
Protein digestion
Differentiated C2C12 myotubes were exposed to the SIT stimulation protocol and then divided into two groups with one immediately treated after the SIT session with 10 µM S107 for 72 h. After 72 h, cell pellets were collected and stored at −80 °C for later analysis. Replicate samples of SIT and SIT S107 (5 per group) were digested with the miST method (modified version of the in-StageTip method, ref. 102). Briefly, frozen cell pellets were resuspended in 100 µl miST lysis buffer (1% sodium deoxycholate, 100 mM Tris pH 8.6, 10 mM dithiothreitol) by vigorous vortexing. Resuspended samples were heated at 95 °C for 5 min, and 100 µg of protein were transferred into new tubes, based on tryptophan quantification103. Samples were then diluted 1:1 (v:v) with water containing 4 mM MgCl2 and benzonase (Merck #70746, 100× dilution of stock = 250 U/µl) and incubated for 15 min at room temperature to digest nucleic acids. Reduced disulfides were alkylated by adding 1/4 vol (25 µl) of 160 mM chloroacetamide (final 32 mM) and incubating at 25 °C for 45 min in the dark. Samples were adjusted to 3 mM EDTA and digested with 1 µg Trypsin/LysC mix (Promega #V5073) for 1 h at 37 °C, followed by a second 1 h digestion with a second and identical aliquot of proteases. To remove sodium deoxycholate, two sample volumes of isopropanol containing 1% trifluoroacetic acid (TFA) were added to the digests, and the samples were desalted on a strong cation exchange (SCX) plate (Oasis MCX; Waters Corp., Milford, MA, USA) by centrifugation. After washing with isopropanol/1%TFA, peptides were eluted in 250 µl of 80% MeCN, 19% water and 1% (v/v) ammonia.
Tandem mass tag (TMT) labelling
Eluates after SCX desalting were dried and resuspended in 100 µl water. Thirty microliters of digests were then aliquoted and dried again, before resuspension in 25 µl of 50 mM TEAB buffer, pH 8.0. For labelling, 0.2 mg of TMT reagent in 20 µl acetonitrile were added to the samples for 1 h at room temperature, after which excess reagent was quenched with 1 µl of 5% hydroxylamine for 15 min at room temperature.
An aliquot (0.8 µl) was injected before mixing to assess labelling completion (>98% peptide spectrum matches) by database search with TMT as variable modification (MASCOT software, www.matrixscience.com). After mixing, the TMT multiplex sample was dried and desalted on a SepPak micro C18 96-well plate (Waters Corp., Milford, MA, USA).
Peptide fractionation
The dried desalted eluate was dissolved in 4 M urea containing 0.1% ampholytes pH 3–10 (GE Healthcare, Ecublens, Switzerland). Then, 75% of the sample (225 µg) was fractionated by off-gel focusing as described104. The 24 peptide fractions obtained were desalted on a SepPak micro C18 96-well plate, dried and dissolved in 25 µl of 0.05% trifluoroacetic acid, 2% (v/v) acetonitrile for liquid chromatography-mass spectrometry/mass spectrometry (LC-MS/MS) analysis.
MS analysis
Data-dependent LC-MS/MS analysis of TMT samples was carried out on a Fusion Tribrid Orbitrap mass spectrometer (Thermo Fisher Scientific, Basel, Switzerland) interfaced through a nano-electrospray ion source to an Ultimate 3000 RSLCnano HPLC System (Dionex, Thermo Fisher Scientific, Basel, Switzerland). Peptides were separated on a reversed-phase custom packed 40 cm C18 column (75 μm ID, 100 Å, Reprosil Pur 1.9 µm particles, Dr. Maisch HPLC Gmbh, Ammerbuch-Entringen, Germany) with a 4–76% acetonitrile gradient in 0.1% formic acid (total time = 140 min). Full MS survey scans were performed at 120,000 resolution. A data-dependent acquisition method controlled by the Xcalibur 4.2 software (Thermo Fisher Scientific, Basel, Switzerland) was used to optimize the number of precursors selected (top speed) of charge 2+ to 5+ while maintaining a fixed scan cycle of 1.5 s. The precursor isolation window used was 0.7 Th.
Peptides were fragmented by higher energy collision dissociation with a normalized energy of 37%. MS2 scans were performed at a resolution of 50,000 in the Orbitrap cell, to resolve 10-plex TMT reporter ions. The m/z of fragmented precursors was then dynamically excluded from selection during 60 s.
MS data analysis
Data files were analysed with MaxQuant 1.6.3.4105,106, incorporating the Andromeda search engine106. Cysteine carbamidomethylation and TMT labelling (peptide N termini and lysine side chains) were selected as fixed modifications while methionine oxidation and protein N-terminal acetylation were specified as variable modifications. The sequence databases used for searching were the mouse (Mus musculus) Reference Proteome based on the UniProt database (www.uniprot.org, version of January 31, 2019, containing 54,211 sequences), and a “contaminant” database containing the most usual environmental contaminants and the enzymes used for digestion (keratins, trypsin, etc). Mass tolerance was 4.5 p.p.m. on precursors (after recalibration) and 20 p.p.m. on higher energy collision dissociation fragments. Both peptide and protein identifications were filtered at a 1% false discovery rate (FDR) relative to hits against a decoy database built by reversing protein sequences. For TMT analysis, the raw reporter ion intensities generated by MaxQuant (with a mass tolerance of 0.003 Da) and summed for each protein group were used in all following steps to derive quantitation.
Processing of quantitative data and statistical tests
The MaxQuant output table “proteinGroups.txt” was processed with the Perseus software107 to remove proteins matched to the contaminants database as well as proteins identified only by modified peptides or reverse database hits, and those without any quantitative values, yielding a first unfiltered list of 7229 identified proteins. Next, the table was filtered to retain only proteins identified by a minimum of three peptides (5870 protein groups left).
After log 2 transformation of all intensity values and normalization by median subtraction, a two-sample t test with Benjamini–Hochberg FDR correction (threshold at 0.05 on the adjusted p value;108 was performed between the SIT and SIT S107 groups and showed 365 significant proteins. Gene ontology annotation enrichment test was carried out with Perseus, using an average difference in log 2 scale between both groups, and applying a Benjamini–Hochberg FDR of 2%. The resulting score used in Fig. 3 and Supplementary Fig. S3 indicates how far is the centre of the distribution of values for the protein category considered relatively to the overall distribution of values109. The interval of this positional score is between −1 and 1.
Raw data deposition
All the mass spectrometry proteomics raw data together with MaxQuant output tables are available via the Proteomexchange Consortium via the PRIDE110 partner repository with the dataset identifier PXD018409.
Submission details:
Project Name: “Triggering acute RyR1 Ca2+ leak leads to improved mitochondrial remodeling and function”.
Project accession: PXD018409.
Project DOI: Not applicable.
The proteomics data generated in this study have been deposited with PRIDE under accession code PXD018409.
Analysed proteomics data are provided in Supplementary Data 1 and 2. SIE (sprint interval exercise) refers to SIT.
The Supplementary Method is provided in the Supplementary information file.
Chemicals
Fluo-4 AM, DMEM, CM-H2DCFDA, Rhod-2 AM, FBS, HS, NEAA, streptomycin–penicillin Lipofectamine 3000, OptiMEM solutions were purchased from Life Technologies (Thermo Fisher, Basel, Switzerland). Details on chemicals are provided in Supplementary Data 3.
Statistical analysis
Except for proteomic and metabolomic analyses, data are presented as mean ± SD. Statistical significance was determined using unpaired t tests to compare two groups or paired t tests to compare two time points. When data were normally distributed, analysis of variance (ANOVA) was performed to compare more than two groups. Two-way ANOVA was followed by Tukey’s or Sidak’s multiple comparisons post hoc tests to compare many groups or multiple time points when the interaction was significant. One-way ANOVA was followed by Tukey’s or Sidak’s multiple comparisons post hoc tests to compare different groups when the main effect was significant. When data were not normally distributed, Kruskal–Wallis ANOVA test followed by Dunn’s multiple comparisons were used to compare different groups. Data were analysed using GraphPad Prism version 8.4.2 and SigmaPlot version 11.0. The level of significance was fixed at p ≤ 0.05. All detailed p values are presented in Supplementary Data 5. The figures were mounted using Adobe Illustrator 23.0.3.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.