Abstract
Food digestion requires the cooperation of different digestive organs. The differentiation of digestive organs is crucial for larvae to start feeding. Therefore, during digestive organogenesis, cell identity and the tissue morphogenesis must be tightly coordinated but how this is accomplished is poorly understood. Here, we demonstrate that WD repeat domain 5 (Wdr5)-mediated H3K4 tri-methylation (H3K4me3) coordinately regulates cell differentiation, proliferation and apoptosis in zebrafish organogenesis of three major digestive organs including intestine, liver, and exocrine pancreas. During zebrafish digestive organogenesis, some of cells in these organ primordia usually undergo differentiation without apoptotic activity and gradually reduce their proliferation capacity. In contrast, cells in the three digestive organs of wdr5−/− mutant embryos retain progenitor-like status with high proliferation rates, and undergo apoptosis. Wdr5 is a core member of COMPASS complex to implement H3K4me3 and its expression is enriched in digestive organs from 2 days post-fertilization (dpf). Further analysis reveals that lack of differentiation gene expression is due to significant decreases of H3K4me3 around the transcriptional start sites of these genes; this histone modification also reduces the proliferation capacity in differentiated cells by increasing the expression of apc to promote the degradation of β-Catenin; in addition, H3K4me3 promotes the expression of anti-apoptotic genes such as xiap-like, which modulates p53 activity to guarantee differentiated cell survival. Thus, our findings have discovered a common molecular mechanism for cell fate determination in different digestive organs during organogenesis, and also provided insights to understand mechanistic basis of human diseases in these digestive organs.
Similar content being viewed by others
Introduction
The vertebrate digestive track and associated organs (pancreas, liver and gall bladder) develop from a primitive gut tube that is derived from definitive endoderm [1,2,3]. In zebrafish, after gastrulation, a sparse layer of endoderm cells migrate medially and form a solid rod along the midline at 20 hours post-fertilization (hpf). A clear lumen can be observed in most of the gut by 42 hpf, whereas the liver and pancreatic buds are clearly identifiable at the fore-part of the rod at 50 hpf [1, 4,5,6,7]. After organ bud formation, some of cells start to differentiate. At 5 days post-fertilization (dpf), zebrafish larvae begin to feed, which requires the cooperation of different digestive organs. Therefore, it is necessary for these organ buds to coordinately develop a functional digestive system on time. It has been identified that many signal molecules (FGF, BMP, Wnt, retinoic acid (RA), Hedgehog, TGF-β and Notch, etc) [8], and transcription regulators (Gata4/5/6, FoxA1/2/3, Cdx2, Sox9, Hnf1α/β, Hnf4α, Neurogenin3, Nkx2.2, NeuroD, Pax4/6, Pdx1, Ptf1a, Hnf6, OC2, Yap1 and C/EBPα, etc) play multiple stage-specific roles in the process of different digestive organ differentiation [9], whereas some other components such as Def, Npo, Elys, Lhr-1, Sec13 and Bms1, etc, function as a pan-digestive factor to regulate the cell proliferation in the entire digestive system [8, 10,11,12,13,14]. However, it is not known if there is a mechanism to coordinately regulate cell differentiation in different digestive organs during organogenesis. It is also unclear how differentiated cells gradually lose their proliferation capacity and remain survival, which is one of fundamental questions in organ development.
WD repeat domain 5 (Wdr5) is best characterized as an adaptor protein of the COMPASS complex that catalyze Histone3 lysine 4 di- and tri-methylation (H3K4me2,3) [15, 16], but emerging evidence demonstrates that it has functions outside this complex including as a component of the NSL (non-specific lethal complex) for H4 lysine 16 acetylation (H4K16ac) [15, 17, 18], interacting with Oct4, Myc, p53 or lncRNAs to facilitate their target gene expression, controlling expression of genes related to protein synthesis, re-initiating transcription upon exit from mitosis, and promoting faithful assembly of mitotic spindle [19,20,21,22,Full size image
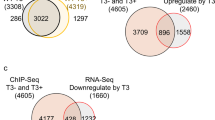
Among 2895 genes with decreased H3K4me3 peaks in wdr5−/− mutant, 953 genes were overlapped with down-regulated genes in RNA-seq (Fig. 2d). KEGG pathway analysis of these shared down-regulated genes showed that most of top 10 items were associated with different metabolic processes and phototransduction (Fig. 2e). Among 239 down-regulated endodermal differentiation genes in RNA-seq, H3K4me3 peaks were decreased in 69 genes (for instance, apom and ebp), not changed in 52 genes (sdr16c5b, tprg1 etc), not detected in 117 genes (fabp10a, fabp2, prss59.1, gc, etc), and increased only in one gene (Fig. 2d, f, g and Supplementary Fig. 4g, h). The results suggested that the decrease of H3K4me3 level in the endodermal organs of wdr5−/− mutant embryos (Fig. 2a, c) was correlated to the expression downregulation of many endodermal differentiation genes.
Among 120 digestive organogenesis regulators, H3K4me3 peaks were not changed in 65 genes, increased in 30 genes, not detected in 23 genes, decreased only in 2 genes (Supplementary Fig. 4e). H3K4me3 peaks were not detected in the only one downregulated endodermal transcriptional factor cdx1 gene (Supplementary Fig. 4f). The data demonstrated that Wdr5-mediated H3K4me3 promotes endodermal organ differentiation not by up-regulating the expression of these digestive organogenesis regulators.
Wdr5-mediated H3K4me3 is required for digestive organ differentiation
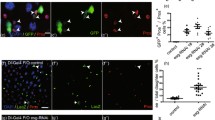
Previous studies have demonstrated that Ser91, Phe133, and Tyr191 are key residues of human WDR5 to directly interact with the N-terminal of Histone 3 (H3) [29, 30]. The protein sequence alignment showed zebrafish Wdr5 shares a high similarity with human or mouse WDR5, and especially the three residues of Ser91, Phe133, and Tyr191 are also conserved among the three species (Supplementary Fig. 5a). To evaluate whether the interaction between Wdr5 and H3 plays an essential role in the differentiation of endodermal organs, we generated two transgenetic lines: Tg(hsp70:HA-Wdr5WT) and Tg(hsp70:HA-Wdr5S91K,F133A,Y191F) (Fig. 3a). Western blot showed that HA-Wdr5WT or HA-Wdr5S91K,F133A,Y191F was induced in corresponding transgenetic lines by heatshock treatments (Fig. 3d). Co-immunoprecipitation (Co-IP) showed that the mutations in three residues of zebrafish Wdr5 did not affect the complex formation with H3 and H4K16ac, but obviously impaired the H3K4me3 (Fig. 3b). The induction of HA-Wdr5WT obviously increased the H3K4me3 level in wdr5−/− embryos at 3 dpf, but not in wdr5+/− embryos. However, the induction of HA-Wdr5S91K,F133A,Y191F did not changed the H3K4me3 level in either wdr5−/− or wdr5+/− embryos at 3 dpf (Fig. 3d), suggesting that the three residues are also important for Wdr5 to facilitate H3K4me3 in zebrafish.
a Two transgenic lines of Tg(hsp70:HA-Wdr5WT) and Tg(hsp70:HA-Wdr5S91K,F133A,Y191F) were treated with heat shock. Two transgenes of Tg(hsp70:HA-Wdr5WT) and Tg(hsp70:HA-Wdr5S91K,F133A,Y191F) were generated in wdr5+/− background. The upper diagram showing how the heat shock treatment was performed. The embryos crossed from each transgenic line were treated with four times of heat shock (39 °C for half an hour) at 1, 1.5, 2, and 2.5 dpf, respectively. The treated embryos were sampled at 3 dpf. Representative images were taken before the genoty**. b Co-IP analysis of the interaction between HA-Wdr5 and H3, H3K4me3 or H4K16ac in two transgenetic lines: Tg(hsp70:HA-Wdr5WT) and Tg(hsp70:HA-Wdr5S91K,F133A,Y191F) under heatshock condition. HA beads were used for immunoprecipitation. β-Actin and β-Tubulin were used as the interaction negative and positive controls, respectively. More than 300 embryos of 1 dpf from each line were treated with heat shocks as described in Fig. 3a. c Co-IP of the interaction between Flag-N-Setd1a or Flag-Rbbp5 with the HA-Wdr5WT or HA-Wdr5S91K,F133A,Y191F in 293 T cells. The upper diagram showing Flag-tagged N terminal of zebrafish Setd1a protein used in Co-IP. Different plasmids were transfected or co-transfected into 293 T cells as indicated. HA beads were used for immunoprecipitation. A zebrafish Wdr5 antibody was used to detect HA-Wdr5WT or HA-Wdr5S91K,F133A,Y191F. Flag antibody was used to detect Flag-Rbbp5 and Flag-N-Setd1a. d Western blots detected by anti-Wdr5, anti-HA, anti-H3K4me3 or anti-H3 antibody in different samples as indicated. Two transgenic lines Tg(hsp70:HA-Wdr5WT) and Tg(hsp70:HA-Wdr5S91K,F133A,Y191F) were generated in wdr5+/− background. More than 200 embryos of 1 dpf from each line were treated with heat shocks as described in Fig. 3a. Treated embryos were genotyped before protein extraction. At least 30 embryos in each group were pooled together for protein extraction. e WISH of fabp10a in transgenic wdr5−/− and sibling embryos treated with heat shocks as described in Fig. 3a. Each experiment was repeated for three times with similar results and a representative was showed here. n indicates the number of zebrafish embryos in each group.
Studies from human cell lines have demonstrated that WDR5 is an adaptor protein in the COMPASS complex and can directly interact with N-terminal of SETD1A or RBBP5 [31, 32]. N terminal of zebrafish Setd1a shared a high similarity with human SETD1A (Supplementary Fig. 5b, c). To investigate if mutations in the three residues of Wdr5S91K,F133A,Y191F interrupt the interaction between Wdr5 and Setd1a or Rbbp5, we co-transfected zebrafish HA-Wdr5 and Flag-N-Setd1a or Flag-Rbbp5 into 293 T cells, Co-IP showed that both HA-Wdr5WT and HA-Wdr5S91K,F133A,Y191F were able to interact with Rbbp5 (Fig. 3c). However, only HA-Wdr5WT, but not HA-Wdr5S91K,F133A,Y191 interacted with N-Setd1a (Fig. 3c). The results suggest that these three residues are important for Wdr5 to form complex with Setd1a, but not with Rbbp5. Furthermore, the abnormal phenotypes of wdr5−/− embryos at 3 dpf including curved body, small eyes, the expression of digestive organ differentiation genes: apom, ebp, fabp10a, fabp2, prss59.1 were almost completely rescued by the induction of HA-Wdr5WT, but not by the induction of HA-Wdr5S91K,F133A,Y191F (Fig. 3a, e and Supplementary Fig. 5d).
Taken together, these data demonstrate that the interaction between Wdr5 and Setd1a is required for Wdr5 to mediate H3K4me3, which plays an essential role in zebrafish endodermal organ differentiation.
Overactivation of Wnt/β-Catenin signal is responsible for the increase of cell proliferation in wdr5 −/− mutant digestive organs
Transcriptome analysis revealed that cell cycle was among the up-regulated items in the wdr5−/− mutant embryos (Supplementary Fig. 2d). Among 6724 differentiated expression genes (DEGs) between WT and wdr5−/− embryos, 123 cell cycle related genes were upregulated and only 46 cell cycle related genes were downregulated (Supplementary Fig. 6a). Western blot showed that total phospho-Histone3 (pH3, a mitosis marker) level significantly increased in the wdr5−/− mutant embryos at 3 and 5 dpf (Fig. 4a). Immunostaining displayed that the percentages of pH3 positive cells also significantly increased in the liver and intestine of wdr5−/− mutant embryos at 3 and 5 dpf (Fig. 4b and Supplementary Fig. 6b, c). The results demonstrated that cell proliferation was increased in the digestive organs of wdr5−/− mutant embryos. In searching for the reasons of the increased proliferation, we found that the expression of β-Catenin target genes (such as: myca, mycb, mycn and ccnd1 highly correlated to cell proliferation) was significantly up-regulated in wdr5−/− mutant embryos from transcriptome analysis (Fig. 4c). Western blot showed that β-Catenin protein was increased in wdr5−/− mutant embryos at 3 dpf (Fig. 4d). Previous studies have demonstrated that the Wnt /β-Catenin signal promotes hepatoblast and intestinal stem cell proliferation [33,34,35]. To investigate if the over activation of β-Catenin plays a role for the increased proliferation in wdr5−/− mutant endodermal organs, we treated wdr5−/− mutant embryos with Salinomycin sodium salt (SAL), a Wnt signal inhibitor (inhibiting phosphorylation of Lrp6-Wnt receptor) [36, 37]. The results showed that the treatment of SAL not only decreased the accumulation of β-Catenin and pH3, and the expression of β-Catenin-target genes (myca and ccnd1) in wdr5−/− mutant embryos down to the levels of those in WT embryos, but also significantly reduced the percentage of pH3 positive cells in the liver and intestine of wdr5−/− mutant embryos at 3 and 5 dpf, whereas the treatment of SAL did not have obvious effects on the accumulation of β-Catenin and pH3 in WT embryos at 3 and 5 dpf, suggesting that the over activation of β-Catenin was responsible for the increased proliferation in wdr5−/− mutant endodermal organs (Fig. 4e–g and Supplementary Fig. 6d, e).
a Western blots of pH3, Wdr5, β-Actin, H3 in WT and wdr5−/− mutant embryos at 3 and 5 dpf. Relative intensity of pH3 was normalized with H3. b Cryosections of WT and wdr5−/− mutant embryos at 3 and 5 dpf were immunostained by anti-pH3 (in red) anti-Bhmt (a liver specific marker in green). The nuclear was stained with DAPI (in blue). L: liver; I: intestine. Framed area was magnified in bottom panel. Scale bar: 40 μm. The percentage of pH3 positive cells in each sample was calculated as the number of pH3 positive cells divided by total cell number in different organs from continuous cryosections. Also see Supplementary Fig. 6b. c Transcript TPM of β-Catenin targeted genes (myca, mycb, mycn and ccnd1) in RNA-seq from WT and wdr5−/− mutant embryos at 3 dpf. d Western blots of β-Catenin, Wdr5, β-Actin, in WT and wdr5−/− mutant embryos at 3 dpf. Relative intensity of β-Catenin was normalized with β-Actin. e Western blots of β-Catenin, pH3, Wdr5, β-Actin and H3 in WT and wdr5−/− mutant embryos with different treatment as indicated. The wdr5−/− mutant and WT embryos at 2.3 or 4 dpf were treated with SAL or DMSO. The protein was extracted from treated embryos at 3 and 5 dpf. Also see Supplementary Fig. 6d, e. f Relative expression level of myca and ccnd1 in WT and wdr5−/− mutant embryos at 3 dpf with SAL or DMSO. The treatment was described in Fig. 4e. g Cryosections of WT and wdr5−/− mutant embryos at 3 dpf treated with SAL or DMSO were immunostained by anti-pH3 (in red) and anti-Bhmt (in green). The nuclear was stained with DAPI (in blue). L: liver; I: intestine. Framed area was magnified in bottom panel. Scale bar: 40 μm. Statistical analysis on the percentage of pH3 positive cells in the liver or intestine of wdr5−/− embryos at 3 dpf treated with SAL or DMSO was showed in the right panel. The treatment was described in Fig. 4e. Each experiment was repeated for three times with similar results and a representative was showed here. n indicates the number of zebrafish embryos in each group. Data are mean±S.D. Two-tailed t-test was applied for each individual comparison (*p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; n.s no significance).
Wdr5-mediated H3K4me3 upregulates apc expression to inhibit Wnt/β-Catenin signal, resulting cell cycle termination in differentiated cells of digestive organs
To find out why β-Catenin protein increased in wdr5−/− mutant embryos, we examined the expression of upstream regulators of β-Catenin including ctnnb1 encoding β-Catenin (Supplementary Fig. 7a), wnt ligands and members of β-Catenin degradation complex from transcriptome analysis. Interestingly, we found that the expression of Adenomatous Polyposis Coli (apc), a member of β-Catenin degradation complex, decreased up to 35%, while the H3K4me3 peaks in apc locus were also decreased about 33% in wdr5−/− mutant embryos (Fig. 5a). WISH showed that the expression of apc enriched in head and endodermal organs of WT embryos at 3 dpf (Fig. 5b). However, no expression enrichment was observed in endodermal organs of wdr5−/− mutant embryos at 3 dpf (Fig. 5b). The results suggested that downregulation of apc expression was due to the decrease of Wdr5-mediated H3K4me3 in endodermal organs. A previous study showed that a zebrafish apc+/− heterozygous mutant embryos developed enlarged livers through increased proliferation [38]. To evaluate whether this apc haploid insufficiency also results increased proliferation in other digestive organs, we used another apc mutant allele (Supplementary Fig. 7c). The homozygous mutation of apc−/− led to early embryo lethality. Interestingly, similar to the wdr5−/− mutant embryos, the levels of total β-Catenin and pH3 were also increased in apc+/− mutant embryos at 3 dpf (Fig. 5c), whereas the expression of apc in both of apc+/− and wdr5−/− embryos was significantly lower than that in WT embryos at 3 dpf (Supplementary Fig. 7d). The percentages of pH3 positive cells were significantly increased in liver and intestine of apc+/− mutant embryos at 3 dpf (Fig. 5d).
a Graphs showing H3K4me3 and RNA peaks at apc locus in WT and wdr5−/− mutant embryos at 3 dpf. b WISH of apc in WT and wdr5−/− mutant at 3 dpf. L: liver; I: intestine; P: pancreas. c Western blots of β-Catenin, pH3, Wdr5, β-Actin and H3 in WT, wdr5−/− and apc+/− embryos at 3 dpf. Relative intensities of β-Catenin and pH3 in WT, wdr5−/− and apc+/− from the western blots were normalized with H3 was showed in right panel. d Cryosections of WT and apc+/− embryos at 3 dpf were immunostained by anti-pH3 (in red) and anti-Bhmt (in green). The nuclear was stained with DAPI (in blue). L: liver; I: intestine. Framed area was magnified in bottom panel. Scale bar: 40 μm. Statistical analysis on the percentages of pH3 positive cells in the liver or intestine between WT and apc+/− embryos at 3 dpf was showed in the right panel. e Western blots of pH3 and H3 in WT embryos at 1 dpf, 3 dpf and 5 dpf. f Western blots of β-Catenin and β-Actin in WT embryos at 3 dpf and 5 dpf. g Statistical analysis on the percentages of pH3 positive cells in liver and intestine of WT between 3 dpf and 5 dpf. h Relative expression of apc in WT embryos from 1–5 dpf. Each experiment was repeated for three times with similar results and a representative was showed here. n indicates the number of zebrafish embryos in each group. Data are mean±S.D. Two-tailed t-test was applied for each individual comparison (*p < 0.05; ***p < 0.001, ****p < 0.0001, n.s no significance).
To evaluate if Wdr5 promotes apc expression to terminate digestive organ proliferation through H3K4me3, we performed western blot to examine the levels of β-Catenin protein and pH3, and quantitative Real-Time PCR (qRT-PCR) to analyze the transcripts of apc and β-catenin in two transgenetic lines: Tg(hsp70:HA-Wdr5WT) and Tg(hsp70:HA-Wdr5S91K,F133A,Y191F) under wdr5−/− genetic background. The qRT-PCR showed that the induction of HA-Wdr5WT, but not HA-Wdr5S91K,F133A,Y191F, up-regulated the expression of apc, while both transgenes had a little effect on the expression of β-catenin (Supplementary Fig. 7e). Western blot displayed that only HA-Wdr5WT down-regulated the levels of β-Catenin protein and pH3 (Supplementary Fig. 7f). Taken together, the data demonstrated that Wdr5-mediated H3K4me3 repressed cell proliferation through promoting the expression of apc to degrade β-Catenin protein.
In addition, we observed that the expression of apc was gradually increased from 1 to 5 dpf (Fig. 5h), and the enrichment of apc expression in endodermal organs was also gradually increased from 1 to 5 dpf in WT embryos (Supplementary Fig. 7b), while the levels of total β-Catenin and pH3 were decreased from 3 to 5 dpf in WT embryos (Fig. 5e, f). The percentage of pH positive cells in either liver or intestine of WT embryos was greatly reduced from 3 to 5 dpf (Fig. 5g). The results indicated that on the one hand, Wdr5-mediated H3K4me3 promoted the expression of digestive organ differentiation genes; on the other hand, Wdr5-mediated H3K4me3 decreased the cell proliferation in differentiated cells by promoting apc expression.
The increase of apoptotic activity in wdr5 −/− mutant is largely dependent on the activation of p53
Next question is why the digestive organs in wdr5−/− mutant embryos are smaller, even though their cell proliferation rate is higher, than those in WT embryos. Our previous studies have demonstrated that Δ113p53 is an N-terminal truncated p53 isoform that lacks exon 1 to 4 [10]. It is a p53 target gene and transcribed from an alternative promoter in intron 4 [10]. Transcriptome analysis showed that the expression of exon 1-4 of p53 gene was decreased and the expression of exon 5-12 of p53 was increased in wdr5−/− mutant embryos, which was confirmed by qRT-PCR (Fig. 6a, b). Furthermore, the accumulation of H3K4me3 was decreased around exon1 and increased in intron4 (Fig. 6a). The results demonstrated that the transcription of the full-length p53 was decreased and the transcription of its target gene Δ113p53 was increased. The expression of other p53 target genes (including p21, mdm2, bax, gadd45ba) was also increased in wdr5−/− mutant embryos (Supplementary Fig. 8a). Western blot revealed that p53 protein obviously increased in wdr5−/− mutant embryos at 3 dpf (Fig. 6c). These results were consistent with KEGG analysis, which showed that the p53 signal pathway in wdr5−/− mutant embryos was among the items of upregulated genes in both RNA-seq and H3K4me3 ChIP-seq (Supplementary Figs. 2d and 4d). To investigate whether the activation of p53 played a role in small digestive organs of wdr5−/− mutant embryos, we generated p53−/−;wdr5−/− double mutant. The abnormal phenotypes in wdr5−/− mutant (curved body, small eyes) were partially rescued in the p53−/−;wdr5−/− mutant (Supplementary Fig. 8b). TUNEL assay showed that the percentages of apoptotic cells in liver, intestine and pancreas at 3 dpf were significantly increased in wdr5−/− mutant, compared with those in WT embryos, whereas the apoptotic activities in the three digestive organs were significantly reduced in p53−/−;wdr5−/− mutant, compared with wdr5−/− mutant (Fig. 6d and Supplementary Fig. 8c). Furthermore, the total cell number of liver and the size of foxa3 expression (a pan marker for liver, pancreas and intestine) at 3 dpf were partially rescued in p53−/−;wdr5−/− mutant, compared with wdr5−/− mutant, while western blot showed that β-Catenin and pH3 in p53−/−;wdr5−/− mutant remained similar levels as those in wdr5−/− mutant (Fig. 6e, f and Supplementary Fig. 8d, e). The results demonstrated that one of main reasons for the small digestive organs in wdr5−/− mutant was the activation of apoptotic activity, which was triggered by the p53 protein accumulation.
a Graphs showing H3K4me3 and RNA peaks at p53 locus in WT and wdr5−/− mutant embryos at 3 dpf. b The confirmation of the expression levels of full-length p53 transcript (Exon 1-4, only in full-length p53) and Δ113p53 transcript (Exon 5–12 in both full-length p53 and Δ113p53) by qRT-PCR. c Western blots detected by anti-p53 (for both p53 50 kD and Δ113p53 35kD), anti-Wdr5, or anti-β-Actin antibody in WT and wdr5−/− mutant embryos at 3 dpf. d Cryosections of WT, wdr5−/− and p53−/−;wdr5−/− embryos at 3 dpf were analyzed by TUNEL assay (in red). The nuclear was stained with DAPI (in blue). Statistical analysis on the percentages of apoptotic cells in the liver, pancreas or intestine between genotypes was showed in the right panel. L: liver; P: exocrine pancreas; I: intestine. e Cryosections of WT, wdr5−/− and p53−/−;wdr5−/− embryos at 3 dpf were immunostained by anti-Bhmt (in green). The nuclear was stained with DAPI (in blue). Framed area was magnified in bottom panel. Scale bar: 40 μm. Total cell number of liver was counted from the Bhmt-positive cells in continuous cryosections. Statistical analysis on the total cell numbers of liver between genotypes was showed in the right panel. f WISH of WT, wdr5−/− and p53−/−;wdr5−/− embryos at 3 dpf with foxa3. g WISH of WT, wdr5−/− and p53−/−;wdr5−/− embryos at 3 dpf with ebp, mgst3a, tkfc, apom and anxa4 as indicated. Each experiment was repeated for three times with similar results and a representative was showed here. n indicates the number of zebrafish embryos in each group. Data are mean ± S.D. Two-tailed t-test was applied for each individual comparison (**p < 0.01; ***p < 0.001, ****p < 0.0001; n.s no significance).
Interestingly, the expression of digestive organ differentiation genes (such as apom, ebp, mgst3a, tkfc and anxa4), that were downregulated in wdr5−/− mutant from both RNA-seq and H3K4me3 ChIP-seq (Fig. 2g and Supplementary Fig. 8f), was not restored in p53−/−;wdr5−/− mutant (Fig. 6g). The results indicated that small undifferentiated digestive organs in wdr5−/− mutant were caused by the apoptosis of undifferentiated cells, but not by the apoptosis of differentiated cells. The expression of the differentiation genes depended on both their specific transcription factors and the H3K4me3 modification on their transcriptional start site. If a cell under differentiated status was not able to differentiate, the cell would undergo apoptosis.
Wdr5-mediated H3K4me3 promotes differentiated cell survival partially by upregulating the expression of xiap-like gene
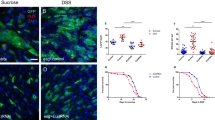
The above results showed that the transcription of p53 was decreased and its protein was increased in wdr5−/− mutant, suggesting that p53 protein negative regulators might be downregulated in wdr5−/− mutant. Therefore, we searched for downregulated anti-apoptotic genes in wdr5−/− mutant from both RNA-seq and H3K4me3 ChIP-seq, and found that only 5 anti-apoptotic genes in this category including choroideremia (chm), zgc:171740, prothymosin α type b (ptmab), cone-rod homeobox (crx) and selenoprotein W, 2a (selenow2a) (Fig. 7a). Previous studies have demonstrated that chm, ptmab, crx and selenow2a promote cell survival during zebrafish development. However, their results also suggest that the survival role of these four genes is unlikely to be realized by regulation of the p53 protein activity [39,40,41,42]. Interestingly, protein alignment revealed that the Zgc:171740 protein shares 21.8% and 23.9% similarities with human and mouse X-linked inhibitor of apoptosis proteins (XIAP) respectively (Fig. 7b). A previous study has showed that XIAP directly interacts with p53 protein to downregulate p53 protein stability and mitochondrial localization [43]. WISH showed that the expression of zgc:171740 was enriched in the liver, intestine and exocrine pancreas of WT embryos at 3 dpf, but disappeared in these digestive organs of wdr5−/− mutant embryos (Fig. 7c). Next, we generated a zebrafish zgc:171740 mutant (Fig. 7d). Western blot showed that the accumulation of p53 protein, but not Wdr5, obviously increased in zgc:171740−/− mutant embryos at 3 dpf (Fig. 7e). TUNEL assay also revealed that the percentage of apoptotic cells was significantly increased in zgc:171740 mutant intestine at 7 dpf (Fig. 7f, g). Although zgc:171740−/− mutant fish developed relative normal at early stage, most of them (80–90%) died around 30 dpf. Thus, we named zgc:171740 as xiap-like gene.
a Graphs showing H3K4me3 and RNA peaks at 5 anti-apoptotic gene loci (zgc:171740, chm, ptmab, crx and selenow2a) in WT and wdr5−/− mutant embryos at 3 dpf. Both H3K4me3 and RNA peaks at these 5 gene loci were decreased in wdr5−/− mutant embryos. b Amino acid sequence alignment between zebrafish Zgc:171740 (Danio rerio NP_001103196), human XIAP (Homo sapiens NP_001158) and ** (Danio_rerio.GRCz11) [63] and differential expression genes (DEGs) analysis (|log2FoldChange | ≥ 1 and Padj < 0.05) were performed by Annoroad company (Bei**g, China). KEGG analysis was further performed using DAVID Bioinformatics resources (https://david.ncifcrf.gov).
Genes related to endodermal organ differentiation and regulators of endodermal organ development were searched from the database of ZFIN (http://www.zfin.org) and published literatures.
Chromatin immunoprecipitation followed by sequencing (ChIP-seq) analysis
Two independent samples from WT and wdr5−/− mutant embryos at 3 dpf were used for ChIP-seq analysis. ChIP-seq and basic analysis including peak calling, peak annotation and differential analysis (|log2FoldChange | ≥ 0.58 and Padj < 0.05) were performed by ActiveMotif Biotechnology company (Shanghai, China). KEGG analysis was further performed using DAVID Bioinformatics resources (https://david.ncifcrf.gov). Up-or down-regulated genes in both RNA-seq and ChIP-seq were found as |log2FoldChange | ≥ 1 in RNA-seq and |log2FoldChange | ≥ 0.58 in ChIP-seq with Padj < 0.05 in both analysis (too few results with |log2FoldChange | ≥ 1 in ChIP-seq).
Sampling of experiments
More than 5 pairs of mutant or WT fish were used to set one specific cross. For a cross between homozygous parents, more than 30 embryos were used for gene expression analysis in each treatment. For a cross between heterozygous parents, more than 50 embryos were used for each treatment and subjected for genoty**. Each experiment was repeated at least 3 times. A representative one was shown in the figures.
Statistical analysis
Significance of differences between means was analyzed using two-sided t-test. Sample sizes were indicated in the figures or figure legends. Plotted mean was calculated by GraphPad software. Data were shown as mean ± SD. P value below 0.05 marked as *, P value below 0.01 marked as **, P value below 0.001 marked as ***, and P value below 0.0001 marked as ****; ns means no significant difference.
Data availability
All relevant data are available from the authors and/or included in the manuscript or Supplementary information. RNA-seq data and ChIP-seq data in this paper have been deposited in the NCBI database (BioProject: PRJNA913404).
References
Ng A. Sevoflurane sedation in infants-a fine line between sedation and general anesthesia. Paediatr Anaesth. 2005;15:1–2.
Fukuda K, Kikuchi Y. Endoderm development in vertebrates: fate map**, induction and regional specification. Dev Growth Differ. 2005;47:343–55.
Wells JM, Melton DA. Vertebrate endoderm development. Annu Rev Cell Dev Biol. 1999;15:393–410.
Field HA, Ober EA, Roeser T, Stainier DY. Formation of the digestive system in zebrafish. I. Liver morphogenesis. Dev Biol. 2003;253:279–90.
Ober EA, Verkade H, Field HA, Stainier DY. Mesodermal Wnt2b signalling positively regulates liver specification. Nature. 2006;442:688–91.
Wallace KN, Akhter S, Smith EM, Lorent K, Pack M. Intestinal growth and differentiation in zebrafish. Mech Dev. 2005;122:157–73.
Wallace KN, Pack M. Unique and conserved aspects of gut development in zebrafish. Dev Biol. 2003;255:12–29.
Tsai SM, Liu DW, Wang WP. Fibroblast growth factor (Fgf) signaling pathway regulates liver homeostasis in zebrafish. Transgenic Res. 2013;22:301–14.
Ober EA, Field HA, Stainier DY. From endoderm formation to liver and pancreas development in zebrafish. Mech Dev. 2003;120:5–18.
Chen J, Ruan H, Ng SM, Gao C, Soo HM, Wu W, et al. Loss of function of def selectively up-regulates Delta113p53 expression to arrest expansion growth of digestive organs in zebrafish. Genes Dev. 2005;19:2900–11.
de Jong-Curtain TA, Parslow AC, Trotter AJ, Hall NE, Verkade H, Tabone T, et al. Abnormal nuclear pore formation triggers apoptosis in the intestinal epithelium of elys-deficient zebrafish. Gastroenterology. 2009;136:902–11.
Mayer AN, Fishman MC. Nil per os encodes a conserved RNA recognition motif protein required for morphogenesis and cytodifferentiation of digestive organs in zebrafish. Development. 2003;130:3917–28.
Wang Y, Luo Y, Hong Y, Peng J, Lo L. Ribosome biogenesis factor Bms1-like is essential for liver development in zebrafish. J Genet Genomics. 2012;39:451–62.
Niu X, Gao C, Jan Lo L, Luo Y, Meng C, Hong J, et al. Sec13 safeguards the integrity of the endoplasmic reticulum and organogenesis of the digestive system in zebrafish. Dev Biol. 2012;367:197–207.
Zhao X, Su J, Wang F, Liu D, Ding J, Yang Y, et al. Crosstalk between NSL histone acetyltransferase and MLL/SET complexes: NSL complex functions in promoting histone H3K4 di-methylation activity by MLL/SET complexes. PLoS Genet. 2013;9:e1003940.
Wysocka J, Swigut T, Milne TA, Dou Y, Zhang X, Burlingame AL, et al. WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell. 2005;121:859–72.
Dias J, Van Nguyen N, Georgiev P, Gaub A, Brettschneider J, Cusack S, et al. Structural analysis of the KANSL1/WDR5/KANSL2 complex reveals that WDR5 is required for efficient assembly and chromatin targeting of the NSL complex. Genes Dev. 2014;28:929–42.
Li X, Li L, Pandey R, Byun JS, Gardner K, Qin Z, et al. The histone acetyltransferase MOF is a key regulator of the embryonic stem cell core transcriptional network. Cell Stem Cell. 2012;11:163–78.
Guarnaccia AD, Tansey WP. Moonlighting with WDR5: a cellular multitasker. J Clin Med. 2018;7:21.
Li Q, Huang Y, Xu J, Mao F, Zhou B, Sun L, et al. p53 inactivation unmasks histone methylation-independent WDR5 functions that drive self-renewal and differentiation of pluripotent stem cells. Stem Cell Rep. 2021;16:2642–58.
Li Q, Mao F, Zhou B, Huang Y, Zou Z, denDekker AD, et al. p53 integrates temporal WDR5 inputs during neuroectoderm and mesoderm differentiation of mouse embryonic stem cells. Cell Rep. 2020;30:465–80.e6.
Thomas LR, Adams CM, Fesik SW, Eischen CM, Tansey WP. Targeting MYC through WDR5. Mol Cell Oncol. 2020;7:1709388.
Huang G, **ang Z, Wu H, He Q, Dou R, Lin Z, et al. The lncRNA BDNF-AS/WDR5/FBXW7 axis mediates ferroptosis in gastric cancer peritoneal metastasis by regulating VDAC3 ubiquitination. Int J Biol Sci. 2022;18:1415–33.
Ali A, Veeranki SN, Chinchole A, Tyagi S. MLL/WDR5 complex regulates Kif2A localization to ensure chromosome congression and proper spindle assembly during mitosis. Dev Cell. 2017;41:605–22.e7.
Wang F, Zhang J, Ke X, Peng W, Zhao G, Peng S, et al. WDR5-Myc axis promotes the progression of glioblastoma and neuroblastoma by transcriptional activating CARM1. Biochem Biophys Res Commun. 2020;523:699–706.
Ang YS, Tsai SY, Lee DF, Monk J, Su J, Ratnakumar K, et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell. 2011;145:183–97.
Vilhais-Neto GC, Fournier M, Plassat JL, Sardiu ME, Saraf A, Garnier JM, et al. The WHHERE coactivator complex is required for retinoic acid-dependent regulation of embryonic symmetry. Nat Commun. 2017;8:728.
Ma Z, Zhu P, Shi H, Guo L, Zhang Q, Chen Y, et al. PTC-bearing mRNA elicits a genetic compensation response via Upf3a and COMPASS components. Nature. 2019;568:259–63.
Dou Y, Milne TA, Ruthenburg AJ, Lee S, Lee JW, Verdine GL, et al. Regulation of MLL1 H3K4 methyltransferase activity by its core components. Nat Struct Mol Biol. 2006;13:713–9.
Ruthenburg AJ, Wang W, Graybosch DM, Li H, Allis CD, Patel DJ, et al. Histone H3 recognition and presentation by the WDR5 module of the MLL1 complex. Nat Struct Mol Biol. 2006;13:704–12.
Dharmarajan V, Lee JH, Patel A, Skalnik DG, Cosgrove MS. Structural basis for WDR5 interaction (Win) motif recognition in human SET1 family histone methyltransferases. J Biol Chem. 2012;287:27275–89.
Zhang P, Lee H, Brunzelle JS, Couture JF. The plasticity of WDR5 peptide-binding cleft enables the binding of the SET1 family of histone methyltransferases. Nucleic Acids Res. 2012;40:4237–46.
Vandernoot I, Haerlingen B, Gillotay P, Trubiroha A, Janssens V, Opitz R, et al. Enhanced canonical Wnt signaling during early zebrafish development perturbs the interaction of cardiac mesoderm and pharyngeal endoderm and causes thyroid specification defects. Thyroid. 2021;31:420–38.
Zhang Z, Rankin SA, Zorn AM. Different thresholds of Wnt-Frizzled 7 signaling coordinate proliferation, morphogenesis and fate of endoderm progenitor cells. Dev Biol. 2013;378:1–12.
Pinto D, Clevers H. Wnt control of stem cells and differentiation in the intestinal epithelium. Exp Cell Res. 2005;306:357–63.
Lu D, Choi MY, Yu J, Castro JE, Kipps TJ, Carson DA. Salinomycin inhibits Wnt signaling and selectively induces apoptosis in chronic lymphocytic leukemia cells. Proc Natl Acad Sci USA. 2011;108:13253–7.
Klose J, Eissele J, Volz C, Schmitt S, Ritter A, Ying S, et al. Salinomycin inhibits metastatic colorectal cancer growth and interferes with Wnt/β-catenin signaling in CD133(+) human colorectal cancer cells. BMC Cancer. 2016;16:896.
Goessling W, North TE, Lord AM, Ceol C, Lee S, Weidinger G, et al. APC mutant zebrafish uncover a changing temporal requirement for wnt signaling in liver development. Dev Biol. 2008;320:161–74.
Song KH, Woo SR, Chung JY, Lee HJ, Oh SJ, Hong SO, et al. REP1 inhibits FOXO3-mediated apoptosis to promote cancer cell survival. Cell Death Dis. 2017;8:e2536.
Emmanouilidou A, Karetsou Z, Tzima E, Kobayashi T, Papamarcaki T. Knockdown of prothymosin α leads to apoptosis and developmental defects in zebrafish embryos. Biochem Cell Biol. 2013;91:325–32.
Penglase S, Hamre K, Ellingsen S. Selenium prevents downregulation of antioxidant selenoprotein genes by methylmercury. Free Radic Biol Med. 2014;75:95–104.
Shen YC, Raymond PA. Zebrafish cone-rod (crx) homeobox gene promotes retinogenesis. Dev Biol. 2004;269:237–51.
Hyeon SJ, Park J, Yoo J, Kim SH, Hwang YJ, Kim SC, et al. Dysfunction of X-linked inhibitor of apoptosis protein (XIAP) triggers neuropathological processes via altered p53 activity in Huntington’s disease. Prog Neurobiol. 2021;204:102110.
Vastenhouw NL, Schier AF. Bivalent histone modifications in early embryogenesis. Curr Opin Cell Biol. 2012;24:374–86.
Kim H, Jang MJ, Kang MJ, Han YM. Epigenetic signatures and temporal expression of lineage-specific genes in hESCs during differentiation to hepatocytes in vitro. Hum Mol Genet. 2011;20:401–12.
Li J, Wu X, Zhou Y, Lee M, Guo L, Han W, et al. Decoding the dynamic DNA methylation and hydroxymethylation landscapes in endodermal lineage intermediates during pancreatic differentiation of hESC. Nucleic Acids Res. 2018;46:2883–900.
San B, Aben M, Elurbe DM, Voeltzke K, den Broeder MJ, Rougeot J, et al. Genetic and epigenetic regulation of zebrafish intestinal development. Epigenomes. 2018;2:23.
Hussain SZ, Sneddon T, Tan X, Micsenyi A, Michalopoulos GK, Monga SP. Wnt impacts growth and differentiation in ex vivo liver development. Exp Cell Res. 2004;292:157–69.
Verzi MP, Shivdasani RA. Wnt signaling in gut organogenesis. Organogenesis. 2008;4:87–91.
Lesko AC, Goss KH, Prosperi JR. Exploiting APC function as a novel cancer therapy. Curr Drug Targets. 2014;15:90–102.
Sansom OJ, Reed KR, Hayes AJ, Ireland H, Brinkmann H, Newton IP, et al. Loss of Apc in vivo immediately perturbs Wnt signaling, differentiation, and migration. Genes Dev. 2004;18:1385–90.
Zhang L, Shay JW. Multiple roles of APC and its therapeutic implications in colorectal cancer. J Natl Cancer Inst. 2017;109:djw332.
Fodde R. The APC gene in colorectal cancer. Eur J Cancer. 2002;38:867–71.
Tao T, Shi H, Guan Y, Huang D, Chen Y, Lane DP, et al. Def defines a conserved nucleolar pathway that leads p53 to proteasome-independent degradation. Cell Res. 2013;23:620–34.
Mudde ACA, Booth C, Marsh RA. Evolution of our understanding of XIAP deficiency. Front Pediatr. 2021;9:660520.
Jost PJ, Vucic D. Regulation of cell death and immunity by XIAP. Cold Spring Harb Perspect Biol. 2020;12:a036426.
Wahida A, Müller M, Hiergeist A, Popper B, Steiger K, Branca C, et al. XIAP restrains TNF-driven intestinal inflammation and dysbiosis by promoting innate immune responses of Paneth and dendritic cells. Sci Immunol. 2021;6:eabf7235.
Azabdaftari A, Uhlig HH. Paneth cell dysfunction and the intestinal microbiome in XIAP deficiency. Sci Immunol. 2021;6:eabm0293.
** Q, Gao Y, Shuai S, Chen Y, Wang K, Chen J, et al. Cdx1b protects intestinal cell fate by repressing signaling networks for liver specification. J Genet Genomics. 2022;49:1101–13.
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X, et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res. 2013;23:465–72.
Gong L, Pan X, Yuan ZM, Peng J, Chen J. p53 coordinates with Δ133p53 isoform to promote cell survival under low-level oxidative stress. J Mol Cell Biol. 2016;8:88–90.
Ye S, Zhao T, Zhang W, Tang Z, Gao C, Ma Z, et al. p53 isoform Δ113p53 promotes zebrafish heart regeneration by maintaining redox homeostasis. Cell Death Dis. 2020;11:568.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Acknowledgements
We thank all members in JC and JRP two labs for their technique help and discussions in this work; Ms. **angfeng Shen for her help in the fish facility.
Funding
This work is financially supported by the National Natural Science Foundation of China (32192400), the National Key R&D Program of China (2018YFA0801005), and the Starry Night Science Fund of Zhejiang University Shanghai Institute for Advanced Study (SN-ZJU-SIAS-004).
Author information
Authors and Affiliations
Contributions
JC and LJL supervised the research. JC and ZZ (Zhe Zhang) conceived the project and designed the experiments. ZZ and LWG prepared RNA samples for RNA-seq and ChIP-seq experiments and performed WISH. ZZ did the entire western blot. ZPM generated the wdr5−/− mutants. ZZ generated the zgc:171740−/− mutant and WISH. YPX provided assistance in the protein Co-IP assay. ZXW helped feed zebrafish and helped in genoty**. YC, MJH, and CG helped with analysis of the RNA sequencing and ChIP-seq data. JC, ZZ, and YC wrote the manuscript. JRP, ZHZ (Zhenhai Zhang), and YHS checked and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Z., Yang, C., Wang, Z. et al. Wdr5-mediated H3K4me3 coordinately regulates cell differentiation, proliferation termination, and survival in digestive organogenesis. Cell Death Discov. 9, 227 (2023). https://doi.org/10.1038/s41420-023-01529-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01529-4
- Springer Nature Limited