Abstract
Senescent cell clearance is emerging as a promising strategy for treating age-related diseases. Senolytics are small molecules that promote the clearance of senescent cells; however, senolytics are uncommon and their underlying mechanisms remain largely unknown. Here, we investigated whether genomic instability is a potential target for senolytic. We screened small-molecule kinase inhibitors involved in the DNA damage response (DDR) in Zmpste24−/− mouse embryonic fibroblasts, a progeroid model characterized with impaired DDR and DNA repair. 4,5,6,7-tetrabromo-2-azabenzamidazole (TBB), which specifically inhibits casein kinase 2 (CK2), was selected and discovered to preferentially trigger apoptosis in Zmpste24−/− cells. Mechanistically, inhibition of CK2 abolished the phosphorylation of heterochromatin protein 1α (HP1α), which retarded the dynamic HP1α dissociation from repressive histone mark H3K9me3 and its relocalization with γH2AX to DNA damage sites, suggesting that disrupting heterochromatin remodeling in the initiation of DDR accelerates apoptosis in senescent cells. Furthermore, feeding Zmpste24-deficient mice with TBB alleviated progeroid features and extended their lifespan. Our study identified TBB as a new class senolytic compound that can reduce age-related symptoms and prolong lifespan in progeroid mice.
Similar content being viewed by others
Introduction
Aging is a major risk factor for many chronic diseases, such as cardiovascular diseases, degenerative diseases, and cancer. These chronic diseases seldom exist discretely in older people, which makes it challenging for clinical strategies to treat specific diseases and resolve various symptoms resulting from comorbid conditions. Interventions that impede the aging process itself offer an alternative approach that may delay the onset of age-related chronic diseases and thus prolong healthy longevity. Cellular senescence is a process that helps to prevent damaged cells from proliferating, thereby protecting cells from transformation and maintaining tissue homeostasis [1]. Senescence is also an important trigger of tissue remodeling during embryonic development and in response to tissue damage; senescence-related factors recruit immune cells to promote tissue regeneration [2]. However, the rate of senescent cell accumulation increases with age, resulting in pervasive damage and deficient cell clearance that disturbs tissue homeostasis. An increased burden of senescent cells in various tissues is a major factor contributing to aging and age-related diseases [3, 4]. Removing senescent cells as a therapeutic strategy was first verified by killing cells with high p16Ink4a expression in INK-ATTAC and p16-3MR transgenic mice, which delayed organ dysfunction and prolonged the lifespan in mice aged both prematurely and naturally [5]. Although this opened a new approach to anti-aging, only ‘late senescence’ cells that express a high level of p16Ink4a can be killed limiting the feasibility and potential application of this approach in humans.
A pharmacological target removal approach has subsequently become the new focus. Small-molecule compounds, termed senolytics, which increase the death rate of senescent cells, can alleviate inflammation and age-related organ dysfunction by targeting diverse survival pathways. Tens of senolytics have been reported. For instance, dasatinib, either alone or combined with quercetin, inhibits tyrosine kinase activity and initiates death in certain senescent cell types [6, 7]. Fisetin and quercetin are natural flavonoids that affect the PI3K/AKT/mTOR pathway; navitoclax (ABT-263) and ABT-737 inhibit the anti-apoptotic protein family Bcl2/BCL-xL, consequently triggering mitochondrial-mediated apoptosis [8, 9], and the cardiac glycosides ouabain has a similar effect. In addition, FOXO4-related peptide inhibits the interaction of p53 with FOXO4 to promote p53-medicated apoptosis [53]. CK2 depletion reduces the formation of γH2AX foci in radiation therapy and causes cancer cell death [54]. It has also been shown that inhibition of CK2 activity improves IR-induced human mesenchymal stem cells (hMSC) senescence [55]. TBB and CX4945, as specific CK2 inhibitors, can induce apoptosis in many types of cancer cells and are promising chemotherapeutic agents for chemo-refractory tumors [56, 57]. Therefore, it is possible that ‘senolytic’ and ‘senomorphic’ effects may coexist, each with opposing activity functions on the same target, and that both ‘toxicity to senescent cells’ and ‘alleviation of cellular senescence’ could attenuate aging processes at the systemic level. Nonetheless, DNA damage severity and drug initiation timing must be carefully evaluated. CK2-based senolytic strategies merit further exploration and development especially regarding applications to healthy aging. To summarize, we have identified several DDR kinase inhibitors, exemplified by TBB, with evident senolytic potential to specifically induce apoptosis in prematurely senescent cells. TBB feeding alleviates premature aging phenotypes and extends lifespan in progeroid mice by inhibiting CK2α phosphorylation of HP1α in response to DNA damage. Together with our previous findings, this new evidence highlights CK2α as a dual target for ‘senolytic’ and ‘senomorphic’ strategies in anti-aging intervention and the senolytic potential of TBB to treat progeroid syndromes.
We propose a model (Fig. 7) that CK2α is particularly important for maintaining the DNA damage response and repair of heterochromatin. In normal cells, CK2α promotes the decondensed structure of heterochromatin by phosphorylating the HP1α in case of DNA damage, thus facilitating the DNA damage response and repair. However, due to the low activity of CK2 in progeria cells, treatment with CK2 inhibitor TBB (TBB itself does not act as an inducer of DNA damage) to further inhibit CK2 activity will lead to increased accumulation of DNA damage, thus leading to apoptosis of progeria cells, acting as a senolytics effect and delaying the phenotype of premature aging mice.
In normal cells undergoing DNA damage, CK2α regulates heterochromatin relaxation by phosphorylating HP1α, which promotes its dynamic dissociation with H3K9me3 and relocalization at DNA double-strand break sites (DSB), which facilitates DNA damage response and repair. Reducing CK2 activity leads to the accumulation of DNA damage, which leads to cellular senescence. Since CK2 activity is low in Zmpste24-deficient cells (progeria cells), CK2 inhibition by TBB aggravates extant DDR defects in senescent cells and results in apoptosis. CK2 activity determines the fate of cells undergoing varying degrees of DNA damage.
Materials and methods
Reagents and antibodies
Rabbit anti-CK2α (ab236664), anti-ATM (ab78), anti-γH2AX (ab81299), anti-Bcl2 (ab32124), anti-Kap-1 (ab10484), anti-p-KAP-1 (Ser824, ab70369), anti-phospho-(Ser/Thr) Phe (ab17464), anti-H3K9me3 (ab8898) were obtained from Abcam (Cambridge, UK). Anti-lamin A/C (sc-20681), anti-p21 (sc-6246), and anti-p16 (sc-1661) were purchased from Santa Cruz Biotechnology. Rabbit anti-γH2AX (05-636), anti-p-ATM (Ser1981) (05-740), anti-HP1α (05–689) were sourced from EMD Millipore. Mouse anti-p-ATM (Ser1981) (#5883), anti-cleaved caspase-3 (#9661) and Rabbit anti-phospho-CK2 Substrate [(pS/pT)DXE] (#8738) were purchased from Cell Signaling Technology (Beverly, MA). Mouse anti-HA, anti-Flag were obtained from Sigma-Aldrich. Alexa Fluor 488 goat anti-rabbit IgG (#A11008), Alexa Fluor 594 goat anti-mouse IgG (#A11005), ImaGene GreenTM C12FDG Kit (#I2904) were purchased from Thermo Fisher Scientific. Mouse anti-actin, anti-GAPDH were obtained from Beyotime. Anti-pT50-HP1α monoclonal antibodies were prepared by Abmart generating from a specific phosphorylated peptide (peptide sequence C-SEEHN(pT)WEPEK). The chemical library were purchased from Targetmol Chemicals Inc. Cellular Senescence Detection Kit was purchased from Cell Signaling Technology (#9860). FITC Annexin V Apoptosis Detection Kit I (#556547), and APC Annexin V Apoptosis Detection Kit I (#550475) were purchased from BD Pharmingen.
Cell culture
The Zmpste24−/− mouse model has been described previously [22]. Primary MEFs were extracted from fetuses of 13.5-day-old pregnant female Zmpste24+/− mice (C57BL background). Cells were cultured in DMEM/HIGH glucose medium containing 10% (vol/vol) fetal bovine serum (Gibco) and 1% penicillin/streptomycin incubated at 37 °C in a 5% CO2 humidified atmosphere.
Drug preparation
All drugs (compounds in the DDR-related kinases inhibitor library- Custom compound library from Targetmol, USA.) were either prepared or provided as 10 mM stock solutions in DMSO. The compounds are list in Supplementary Table S1. They were then diluted in a culture medium to obtain a suitable working solution. Culture medium with DMSO at the same concentration was used as a negative control.
Senotherapeutics asssay
MEFs (5 × 103) at passage 6 were seeded in 96-well plates at least 6 h prior to treatment. After adding the drugs, the MEFs were incubated for 24–48 h. For fluorescence analysis of SA-β-Gal activity, cells were washed once with phosphate buffered saline, C12FDG (10 µM) was added to the culture medium, and the cells were incubated for 2 h. C12FDG fluorescence intensity was detected by Enspire 2300 Multimode Plate Reader (PerkinElmer, MA, USA). All samples were analyzed in duplicate with 3–5 fields per well and mean values and standard deviations were calculated accordingly.
Cell viability assay
MEFs were seeded on 96-well plates for 24 h, and then incubated with test compounds for another 48 h at 37 °C and 5% CO2. Cell viability was assessed from the CellTiter 96® AQueous One Solution Cell Proliferation Assay (cat#G3581, Promega, WI, USA). Pipet 20 µl of CellTiter 96® AQueous One Solution Reagent into each well of the 96-well assay plate containing the samples in 100 µl of culture medium, incubating the plate at 37 °C for 2 h in a humidified, 5% CO2 atmosphere. Record the absorbance at 490 nm using Enspire 2300 Multimode Plate Reader (PerkinElmer, MA, USA).
Plasmid construction and siRNAs
The full-length HP1α gene was subcloned into pcDNA3.1-3xFLAG vectors. CK2 was amplified from a cDNA library of HEK293 cells and cloned into pKH3-3xHA vectors. HP1α mutants were generated using a site-directed mutagenesis kit (New England Biolabs, USA). Transient transfection with these plasmids was performed using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions. Small interfering RNA (siRNA) against Ck2α or mock siRNA were synthesized by GenePharma (Shanghai, China). Transfection was performed using Lipofectamine RNAiMAX Transfection Reagent (Invitrogen) according to the manufacturer’s instructions. The primers used are listed in Supplementary Table S2.
Western blot analysis
Total protein was isolated with modified RIPA buffer (0.1 M Tris-HCl, pH 7.5, 0.1 M NaCl, 1 mM EDTA, 1 mM DTT), phosphatase inhibitor, and protease inhibitor cocktail (Roche, Germany), quantified with a BCA protein assay kit (Invitrogen) and separated by SDS polyacrylamide gel electrophoresis followed by western blotting with the respective antibodies.
Co-immunoprecipitation assay
For co-immunoprecipitation, cells that were subjected to the indicated treatments were lysed in lysis buffer (250 mM NaCl, 20 mM Tris-HCl pH 8.0, 2 mM EDTA, 10% glycerol, and 0.1% NP-40), supplemented with EDTA-free protease inhibitor (Roche). Whole-cell extracts were then incubated with the indicated antibody overnight at 4 °C. Bead-bound immunoprecipitates were washed with lysis buffer, boiled in a loading buffer, then analyzed by western blotting analysis.
Flow cytometry
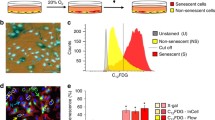
Senescent MEF cells from passage 6 were seeded at 70–80% confluence in 6-well culture plates and cultured for 24 h at 37 °C. For MEF, 5 × 105 cells/well were used. Following the addition of the drugs, lysosomal alkalinization was induced by pretreating cells with 100 nM bafilomycin A1 for 1 h in a fresh cell culture medium at 37 °C. C12FDG (10 μM) solution was then added to the cell culture medium for 2 h. The cells were harvested by trypsinization and resuspended in 1× Annexin V buffer containing 5 µl of Annexin V-APC and 5 µl 7-AAD per 1 × 105 cells/100 µl. The cells were analyzed by flow cytometry within 1 h. To estimate relative SA-β-Gal activity, a two-parameter display of forward scatter versus side scatter was set up, excluding subcellular debris. The events within this region were shown in a green fluorescence histogram, where the Y-axis indicates cell number and the X-axis indicates C12FDG fluorescence intensity on a log scale.
Immunofluorescence staining
Cells were mounted on glass slides and fixed in 4% paraformaldehyde. The cells were then permeabilized with PBS containing 0.1% Triton X-100 for 15 min, and blocked with 1% bovine serum in PBS for 30 min at room temperature. The cells were then incubated with various primary antibodies overnight at 4 °C, and detected by Alexa Fluor conjugated secondary antibodies (Alexa −488, −594; 1: 500, Life Technologies) for 1 h at room temperature in the dark. The cells were then co-stained with 4’, 6-diamidino-2-phenylindole (DAPI) (1:10000 dilution) to visualize the nuclei. Images were captured under a confocal laser scanning microscope system (CarlZeiss). A representative image for each condition is shown.
RNA isolation and quantitative RT-PCR
Total RNA was isolated using Trizol reagent (Invitrogen, 15596-026). RNA (2 μg) and reverse transcribed into cDNA using 5× Primescript RT Master Mix (Takara). Real-time quantitative polymerase chain reaction (RT-qPCR) was performed using 2× SYBR Green Mix (Takara) in a Bio-Rad detection system. Each sample was run in triplicate and gene expression levels were normalized to Gapdh. The primers are listed in Supplementary Table 2.
TUNEL staining
TUNEL staining was assessed with a DeadEnd™ Fluorometric TUNEL System (G3250, Promega, WI, USA) according to the manufacturer’s instructions, and then images were captured under a confocal laser scanning microscope system (CarlZeiss).
SA-β-Gal staining and apoptosis assays
SA-β-Gal staining was assessed with a cellular senescence assay kit according to the manufacturer’s instructions, and then photographed. Blue-stained MEFs were counted in more than 300 randomly chosen cells. Apoptosis was analyzed by flow cytometry as described previously [58].
Animal studies
The Zmpste24−/− mice used have been described elsewhere [22]. Zmpste24+/+ and Zmpste24−/− mice were assigned to the vehicle or the TBB treatment groups, which were fed TBB mixed into their drinking water at a final concentration of 50 μM/L. Survival of TBB-treated or vehicle-treated Zmpste24−/− mice and wild-type controls was recorded, and their body weight was monitored weekly. The survival curves of untreated and TBB-fed mice were compared by Kaplan–Meir analysis.
For bone density analysis, 4-month-old male mice were euthanized by decapitation. The thigh bone was fixed in 4% PFA at 4 °C overnight. The relevant data were collected by micro-CT (Scanco Medical, CT100). Each experimental group included at least three mice. All experiments using mice were performed in accordance with protocols approved by the Committee on the Use of Live Animals in Teaching and Research of Shenzhen University.
Statistical analysis
All experiments were carried out with at least three replicates. All data values are expressed as mean ± standard deviation (SD) or mean ± standard error of the mean (SEM). For multiple comparison, one-way or two-way ANOVA were applied as indicated. Statistical analysis was done by Student’s t test, with a p < 0.05 considered statistically significant.
Data availability
The data are available within the article, Supplementary Information or Original Data file.
References
Krizhanovsky V, Yon M, Dickins RA, Hearn S, Simon J, Miething C, et al. Senescence of activated stellate cells limits liver fibrosis. Cell. 2008;134:657–67.
Prata L, Ovsyannikova IG, Tchkonia T, Kirkland JL. Senescent cell clearance by the immune system: Emerging therapeutic opportunities. Semin Immunol. 2018;40:101275.
Munoz-Espin D, Canamero M, Maraver A, Gomez-Lopez G, Contreras J, Murillo-Cuesta S, et al. Programmed cell senescence during mammalian embryonic development. Cell. 2013;155:1104–18.
van Deursen JM. The role of senescent cells in ageing. Nature. 2014;509:439–46.
Baker DJ, Wijshake T, Tchkonia T, LeBrasseur NK, Childs BG, van de Sluis B, et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature. 2011;479:232–6.
Zhu Y, Tchkonia T, Pirtskhalava T, Gower AC, Ding H, Giorgadze N, et al. The Achilles’ heel of senescent cells: from transcriptome to senolytic drugs. Aging Cell. 2015;14:644–58.
Hickson LJ, Langhi Prata LGP, Bobart SA, Evans TK, Giorgadze N, Hashmi SK, et al. Senolytics decrease senescent cells in humans: preliminary report from a clinical trial of Dasatinib plus Quercetin in individuals with diabetic kidney disease. EBioMedicine. 2019;47:446–56.
Yosef R, Pilpel N, Tokarsky-Amiel R, Biran A, Ovadya Y, Cohen S, et al. Directed elimination of senescent cells by inhibition of BCL-W and BCL-XL. Nat Commun. 2016;7:11190.
Chang Y, Li Y, Ye N, Guo X, Li Z, Sun G, et al. Atorvastatin inhibits the apoptosis of human umbilical vein endothelial cells induced by angiotensin II via the lysosomal-mitochondrial axis. Apoptosis. 2016;21:977–96.
Zhang C, **e Y, Chen H, Lv L, Yao J, Zhang M, et al. FOXO4-DRI alleviates age-related testosterone secretion insufficiency by targeting senescent Leydig cells in aged mice. Aging (Albany NY). 2020;12:1272–84.
Fuhrmann-Stroissnigg H, Ling YY, Zhao J, McGowan SJ, Zhu Y, Brooks RW, et al. Identification of HSP90 inhibitors as a novel class of senolytics. Nat Commun. 2017;8:422.
Samaraweera L, Adomako A, Rodriguez-Gabin A, McDaid HM. A novel indication for panobinostat as a senolytic drug in NSCLC and HNSCC. Sci Rep. 2017;7:1900.
Liebl MC, Hofmann TG. Cell fate regulation upon DNA damage: p53 Serine 46 kinases pave the cell death road. Bioessays. 2019;41:e1900127.
Childs BG, Baker DJ, Kirkland JL, Campisi J, van Deursen JM. Senescence and apoptosis: dueling or complementary cell fates? EMBO Rep. 2014;15:1139–53.
Lopez-Otin C, Blasco MA, Partridge L, Serrano M, Kroemer G. Hallmarks of aging: an expanding universe. Cell. 2023;186:243–78.
Liu Y, Rusinol A, Sinensky M, Wang Y, Zou Y. DNA damage responses in progeroid syndromes arise from defective maturation of prelamin A. J Cell Sci. 2006;119:4644–9.
Manju K, Muralikrishna B, Parnaik VK. Expression of disease-causing lamin A mutants impairs the formation of DNA repair foci. J Cell Sci. 2006;119:2704–14.
Liu B, Wang J, Chan KM, Tjia WM, Deng W, Guan X, et al. Genomic instability in laminopathy-based premature aging. Nat Med. 2005;11:780–5.
Roos WP, Kaina B. DNA damage-induced cell death by apoptosis. Trends Mol Med. 2006;12:440–50.
Tsuzuki T, Fujii Y, Sakumi K, Tominaga Y, Nakao K, Sekiguchi M, et al. Targeted disruption of the Rad51 gene leads to lethality in embryonic mice. Proc Natl Acad Sci USA 1996;93:6236–40.
Anerillas C, Herman AB, Rossi M, Munk R, Lehrmann E, Martindale JL, et al. Early SRC activation skews cell fate from apoptosis to senescence. Sci Adv. 2022;8:eabm0756.
Pendas AM, Zhou Z, Cadinanos J, Freije JM, Wang J, Hultenby K, et al. Defective prelamin A processing and muscular and adipocyte alterations in Zmpste24 metalloproteinase-deficient mice. Nat Genet. 2002;31:94–9.
Navarro CL, Cadinanos J, De Sandre-Giovannoli A, Bernard R, Courrier S, et al. Loss of ZMPSTE24 (FACE-1) causes autosomal recessive restrictive dermopathy and accumulation of Lamin A precursors. Hum Mol Genet. 2005;14:1503–13.
Krishnan V, Chow MZ, Wang Z, Zhang L, Liu B, Liu X, et al. Histone H4 lysine 16 hypoacetylation is associated with defective DNA repair and premature senescence in Zmpste24-deficient mice. Proc Natl Acad Sci USA 2011;108:12325–30.
Malavolta M, Pierpaoli E, Giacconi R, Costarelli L, Piacenza F, Basso A, et al. Pleiotropic effects of tocotrienols and quercetin on cellular senescence: introducing the perspective of senolytic effects of phytochemicals. Curr Drug Targets. 2016;17:447–59.
Du G, Zhao Z, Chen Y, Li Z, Tian Y, Liu Z, et al. Quercetin attenuates neuronal autophagy and apoptosis in rat traumatic brain injury model via activation of PI3K/Akt signaling pathway. Neurol Res. 2016;38:1012–9.
Kang HT, Park JT, Choi K, Kim Y, Choi HJC, Jung CW, et al. Chemical screening identifies ATM as a target for alleviating senescence. Nat Chem Biol. 2017;13:616–23.
Pierre F, Chua PC, O’Brien SE, Siddiqui-Jain A, Bourbon P, Haddach M, et al. Pre-clinical characterization of CX-4945, a potent and selective small molecule inhibitor of CK2 for the treatment of cancer. Mol Cell Biochem. 2011;356:37–43.
Sarno S, Reddy H, Meggio F, Ruzzene M, Davies SP, Donella-Deana A, et al. Selectivity of 4,5,6,7-tetrabromobenzotriazole, an ATP site-directed inhibitor of protein kinase CK2 (‘casein kinase-2’). FEBS Lett. 2001;496:44–8.
Liu J, Yin X, Liu B, Zheng H, Zhou G, Gong L, et al. HP1alpha mediates defective heterochromatin repair and accelerates senescence in Zmpste24-deficient cells. Cell Cycle. 2014;13:1237–47.
Liu B, Zhou Z. Activation of SIRT1 by resveratrol requires lamin A. Aging. 2013;5:94–5.
Liu B, Wang Z, Zhang L, Ghosh S, Zheng H, Zhou Z. Depleting the methyltransferase Suv39h1 improves DNA repair and extends lifespan in a progeria mouse model. Nat Commun. 2013;4:1868.
Hiragami-Hamada K, Shinmyozu K, Hamada D, Tatsu Y, Uegaki K, Fujiwara S, et al. N-terminal phosphorylation of HP1{alpha} promotes its chromatin binding. Mol Cell Biol. 2011;31:1186–1200.
Nishibuchi G, Nakayama J. Biochemical and structural properties of heterochromatin protein 1: understanding its role in chromatin assembly. J Biochem. 2014;156:11–20.
Ayoub N, Jeyasekharan AD, Bernal JA, Venkitaraman AR. HP1-beta mobilization promotes chromatin changes that initiate the DNA damage response. Nature. 2008;453:682–6.
Nishibuchi G, Machida S, Osakabe A, Murakoshi H, Hiragami-Hamada K, Nakagawa R, et al. N-terminal phosphorylation of HP1alpha increases its nucleosome-binding specificity. Nucleic Acids Res. 2014;42:12498–511.
Leung KK, Shilton BH. Quinone reductase 2 is an adventitious target of protein kinase CK2 inhibitors TBBz (TBI) and DMAT. Biochemistry. 2015;54:47–59.
Lolli G, Cozza G, Mazzorana M, Tibaldi E, Cesaro L, Donella-Deana A, et al. Inhibition of protein kinase CK2 by flavonoids and tyrphostins. A structural insight. Biochemistry. 2012;51:6097–107.
Rao F, Cha J, Xu J, Xu R, Vandiver MS, Tyagi R, et al. Inositol pyrophosphates mediate the DNA-PK/ATM-p53 cell death pathway by regulating CK2 phosphorylation of Tti1/Tel2. Mol Cell. 2014;54:119–32.
Khoronenkova SV, Dianova II, Ternette N, Kessler BM, Parsons JL, Dianov GL. ATM-dependent downregulation of USP7/HAUSP by PPM1G activates p53 response to DNA damage. Mol Cell. 2012;45:801–13.
Ciccia A, Huang JW, Izhar L, Sowa ME, Harper JW, Elledge SJ. Treacher Collins syndrome TCOF1 protein cooperates with NBS1 in the DNA damage response. Proc Natl Acad Sci USA 2014;111:18631–6.
Li Z, Li J, Kong Y, Yan S, Ahmad N, Liu X. Plk1 phosphorylation of Mre11 antagonizes the DNA damage response. Cancer Res. 2017;77:3169–80.
McCabe N, Hanna C, Walker SM, Gonda D, Li J, Wikstrom K, et al. Mechanistic rationale to target PTEN-deficient tumor cells with inhibitors of the DNA damage response kinase ATM. Cancer Res. 2015;75:2159–65.
Liu XG, Liang NC, Liu W, She J, Mo LE. [Inhibitory effect and kinetic analysis of sodium quercetin-7,4'-disulphate on recombinant human protein kinase CK2 holoenzyme]. Yao Xue Xue Bao. 2002;37:165–8.
Noy P, Gaston K, Jayaraman PS. Dasatinib inhibits leukaemic cell survival by decreasing PRH/Hhex phosphorylation resulting in increased repression of VEGF signalling genes. Leuk Res. 2012;36:1434–7.
Borgo C, D’Amore C, Sarno S, Salvi M, Ruzzene M. Protein kinase CK2: a potential therapeutic target for diverse human diseases. Signal Transduct Target Ther. 2021;6:183.
Lukas J, Lukas C, Bartek J. More than just a focus: the chromatin response to DNA damage and its role in genome integrity maintenance. Nat Cell Biol. 2011;13:1161–9.
Di Micco R, Sulli G, Dobreva M, Liontos M, Botrugno OA, Gargiulo G, et al. Interplay between oncogene-induced DNA damage response and heterochromatin in senescence and cancer. Nat Cell Biol. 2011;13:292–302.
White D, Rafalska-Metcalf IU, Ivanov AV, Corsinotti A, Peng H, Lee SC, et al. The ATM substrate KAP1 controls DNA repair in heterochromatin: regulation by HP1 proteins and serine 473/824 phosphorylation. Mol Cancer Res. 2012;10:401–14.
Kwon SJ, Park JH, Park EJ, Lee SA, Lee HS, Kang SW, et al. ATM-mediated phosphorylation of the chromatin remodeling enzyme BRG1 modulates DNA double-strand break repair. Oncogene. 2015;34:303–13.
Ao Y, Zhang J, Liu Z, Qian M, Li Y, Wu Z, et al. Lamin A buffers CK2 kinase activity to modulate aging in a progeria mouse model. Sci Adv. 2019;5:eaav5078.
Li WJ, Wang CW, Tao L, Yan YH, Zhang MJ, Liu ZX, et al. Insulin signaling regulates longevity through protein phosphorylation in Caenorhabditis elegans. Nat Commun. 2021;12:4568.
Turowec JP, Duncan JS, Gloor GB, Litchfield DW. Regulation of caspase pathways by protein kinase CK2: identification of proteins with overlap** CK2 and caspase consensus motifs. Mol Cell Biochem. 2011;356:159–67.
Zwicker F, Ebert M, Huber PE, Debus J, Weber KJ. A specific inhibitor of protein kinase CK2 delays gamma-H2Ax foci removal and reduces clonogenic survival of irradiated mammalian cells. Radiat Oncol. 2011;6:15.
Wang D, Jang DJ. Protein kinase CK2 regulates cytoskeletal reorganization during ionizing radiation-induced senescence of human mesenchymal stem cells. Cancer Res. 2009;69:8200–7.
Nitta RT, Bolin S, Luo E, Solow-Codero DE, Samghabadi P, Purzner T, et al. Casein kinase 2 inhibition sensitizes medulloblastoma to temozolomide. Oncogene. 2019;38:6867–79.
Qaiser F, Trembley JH, Kren BT, Wu JJ, Naveed AK, Ahmed K. Protein kinase CK2 inhibition induces cell death via early impact on mitochondrial function. J Cell Biochem. 2014;115:2103–15.
Shao L, Feng W, Li H, Gardner D, Luo Y, Wang Y, et al. Total body irradiation causes long-term mouse BM injury via induction of HSC premature senescence in an Ink4a- and Arf-independent manner. Blood. 2014;123:3105–15.
Acknowledgements
This study was supported by grants from the National Key R&D Program of China, MOST (2023YFC2509900); the National Natural Science Foundation of China (82071580, 81971321); the Shenzhen Municipal Commission of Science and Technology Innovation (JCYJ20200814152850001; JCYJ20210324094606017); SZU Top Ranking Project (86000000210).
Author information
Authors and Affiliations
Contributions
JZ, YA, and YM conducted the experiments; ZZ, BL, YA, and ZW contributed reagents/analytic tools; JZ, YA, PS, ZW, JW, JJ, HZ, BL, and ZW analyzed data; JZ, YA, BL, and ZW wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The animal study was approved by the Ethic Committee approval Institutional is Shenzhen Graduate School of Peking University and the approval code is FYXK (No. 2017-0172).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Anastasis Stephanou
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Sun, P., Wu, Z. et al. Targeting CK2 eliminates senescent cells and prolongs lifespan in Zmpste24-deficient mice. Cell Death Dis 15, 380 (2024). https://doi.org/10.1038/s41419-024-06760-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06760-0
- Springer Nature Limited