Abstract
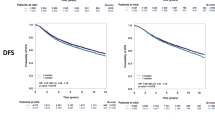
In breast cancer, dysregulated TP53 expression signatures are a better predictor of chemotherapy response and survival outcomes than TP53 mutations. Our previous studies have shown that high levels of Δ40p53 are associated with worse disease-free survival and disruption of p53-induced DNA damage response in breast cancers. Here, we further investigated the in vitro and in vivo implications of Δ40p53 expression in breast cancer. We have shown that genes associated with cell differentiation are downregulated while those associated with stem cell regulation are upregulated in invasive ductal carcinomas expressing high levels of Δ40p53. In contrast to p53, endogenous ∆40p53 co-localised with the stem cell markers Sox2, Oct4, and Nanog in MCF-7 and ZR75-1 cell lines. ∆40p53 and Sox2 co-localisation was also detected in breast cancer specimens. Further, in cells expressing a high ∆40p53:p53 ratio, increased expression of stem cell markers, greater mammosphere and colony formation capacities, and downregulation of miR-145 and miR-200 (p53-target microRNAs that repress stemness) were observed compared to the control subline. In vivo, a high ∆40p53:p53 ratio led to increased tumour growth, Ki67 and Sox2 expression, and blood microvessel areas in the vehicle-treated mice. High expression of ∆40p53 also reduced tumour sensitivity to doxorubicin compared to control tumours. Enhanced therapeutic efficacy of doxorubicin was observed when transiently targeting Δ40p53 or when treating cells with OTSSP167 with concomitant chemotherapy. Taken together, high Δ40p53 levels induce tumour growth and may promote chemoresistance by inducing a stemness phenotype in breast cancer; thus, targeting Δ40p53 in tumours that have a high Δ40p53:p53 ratio could enhance the efficacy of standard-of-care therapies such as doxorubicin.
Similar content being viewed by others
Introduction
Nearly all deaths from breast cancer are a result of resistance to treatment and the subsequent development of metastases [1]. Understanding the mechanisms that contribute to deregulated p53 activities may reveal novel avenues for increasing the sensitivity to commonly used therapies in breast cancer.
In breast cancer, dysregulated p53 expression signatures are a better predictor of outcome and chemotherapy response than TP53 mutation [2, 3], suggesting that alternative molecular mechanisms may compromise p53 function. In recent years, the complexity of p53 signalling has become increasingly apparent with the discovery that p53 is expressed as 12 isoforms whose expression is associated with clinical features and outcomes of human cancers [69]. GO (biological process) overrepresentation analysis was performed on DEGs (up or down) using enrichGO from clusterProfiler (v.4.4.4) [70] and visualised using enrichplot (v.1.16.1) [71]. Detected genes (from Array list) were used as the background list with false discovery rate (FDR) correction (adjusted p < 0.05). The most statistically significant pathways were summarised using the default settings calculated by ‘pairwise_termsim()’ on the GO results and displayed in a Tree plot with default hierarchical clustering by ‘treeplot()’. Heatmap** was performed using pheatmap (v.1.0.12) [72] on normalised counts from Partek filtered for DEGs with row (gene) normalisation by z-score followed by Euclidean hierarchical clustering of both columns (samples) and rows (genes). GO chord plot was plotted by https://www.bioinformatics.com.cn/en, an open-source data visualisation platform.
Cell lines
The normal human epithelial breast cell line MCF-10A and the oestrogen receptor-positive human breast cancer cell lines MCF-7 and ZR75-1, expressing wild-type p53 (WTp53), were generously provided by A/Professor Nikki Verrills and Dr Rick Thorne, respectively. The cell lines were authenticated by the Australian Genome Research Facility as previously described [18]. MCF-7 cells stably overexpressing Δ40p53 via the lentiviral LeGO vector and MCF-7 and MCF-10A knockdown sublines, -shNT (non-targeting control), -shΔ40p53, and -shp53α, were established by transduction of cells with lentiviral vectors [18]. Each of the MCF-7 cell sublines and MCF-7 and ZR75-1 parental cells were maintained in DMEM (Dulbecco modified Eagle’s medium), supplemented with 10% foetal bovine serum (FBS), insulin (10 μg/mL), and L-glutamine (2 mM) (Life Technologies, Mulgrave, VIC, Australia). MCF-10A cell sublines were maintained in DMEM/F12 media supplemented with 10% horse serum, insulin (10 μg/mL), L-glutamine (2 mM), epidermal growth factor (20 ng/mL), hydrocortisone (0.5 μg/mL), cholera toxin (1 ng/mL) (Life Technologies). The medium used for the MCF-7 and MCF-10A cell sublines was further supplemented with puromycin (1 μg/mL) (Sigma-Aldrich, Castle Hill, NSW, Australia) for the maintenance of positive clones. Cells were maintained in humidified 5% CO2 at 37 ˚C and were routinely tested for mycoplasma according to the manufacturer’s recommendations (MycoAlert PLUS, Lonza, Basel, Switzerland).
Immunofluorescence
Immunofluorescence was performed as previously described [20]. Cells were incubated for 1 h with primary antibodies: mouse-anti-human-Nanog (20 μg/ml; Life Technologies #MA1-017), mouse-anti-human-Sox2 (5 μg/ml; Life Technologies #MA1-014), mouse-anti-human-Oct4 (2 μg/ml; Life Technologies #MA1-104), rabbit-anti-human-Zeb1 (1 μg/ml; Bethyl Laboratories, Montgomery, TX, USA #A301-922A), rabbit-anti-human-p53 7F5 (1:800; Cell Signaling Technology, Danvers, MA, USA #2527), rabbit-anti-human-Δ40p53 KJC40 (detects all Δ40p53 isoforms, mainly Δ40p53α [18, 25]; 5 μg/ml; developed by J.C. Bourdon, The University of Dundee, Scotland), and/or rabbit-anti-human-GM130 (1:3200; Cell Signaling Technology #12480). Then, the cells were incubated for 1 h with secondary antibodies: goat-anti-mouse-Alexa Fluor 594 (1:30; Life Technologies #R37121), goat-anti-rabbit Alexa Fluor 594 (4 μg/ml; Life Technologies #A11037), and/or goat-anti-rabbit-Alexa Fluor 488 (4 μg/ml; Life Technologies #A11034). Each well was then stained with DAPI (300 nM in PBS) to detect cell nuclei. For cell spheroid immunofluorescence, following fixation, spheroids were sectioned with a cryostat and the slides were processed as described above. For breast cancer specimens immunofluorescence, FFPE IDC slides from our previous study looking at p53 isoform expression [25] were processed as previously described [73] with minor modifications (rinsing solution: 0.25% Triton-X-100 in phosphate-buffered saline (PBS) and blocking solution: 3% FBS in PBS). Three slides (S#1–3: ER+/PR+/Her2- IDCs) were incubated for 1 h with mouse-anti-human-CD38 (1:100; Leica Microsystems Pty Ltd, Mt Waverley, VIC, Australia #CD38-290-L-CE) and rabbit-anti-human-Δ40p53 KJC40 (8 μg/ml) antibodies and five slides of (S#4–7: ER+/PR+/Her2-and S#8: ER-/PR-/Her2+ IDCs) were incubated for 1 h with mouse-anti-human-Sox2 (5 μg/ml; Life Technologies #MA1-017) and rabbit-anti-human-Δ40p53 KJC40 (8 μg/ml) antibodies. Slides were then washed three times with the rinsing solution and incubated for 1 h with goat-anti-mouse-Alexa-Fluor 594 (1:30) and goat-anti-rabbit-Alexa 488 (4 μg/ml) secondary antibodies. After the final rinsing steps, mounting medium with DAPI was added to the slides. Images were obtained using a Cytation3 cell imager (BioTek, Winooski, VT, USA) using 10x and 40x objectives. For co-localisation analysis in MCF-7 and ZR75-1 cells and IDCs slides, four images were collected per well (approximately 30 cells were evaluated per triplicate) and ten microscope fields were collected per slide, respectively. Images were collected maintaining exposure and contrast settings. Images were analysed using Gen5 software for fluorescence intensity and ImageJ (Coloc 2) for co-localisation, which performs a pixel intensity correlation of regions of interest. Spearman’s rank correlation was calculated between Δ40p53 or p53 and Nanog, Sox2, or Oct4. The identification of the images was blinded to the investigator.
RNA extraction
Total RNA was extracted from cell lines using TRIzol RNA purification reagent (Life Technologies) following manufacturer’s recommendations. The RNA yield was determined by the Qubit RNA BR (broad range) Assay Kit (Life Technologies) on a Qubit 2.0 Fluorometer (Life Technologies), following manufacturer’s recommendations.
Reverse transcription quantitative polymerase chain reactions (RT-qPCR)
Total RNA of 148 IDCs (500 ng) and cell line samples (300 ng) was reverse transcribed into complementary DNA (cDNA) using the High-Capacity Reverse Transcription kit with RNase inhibitor (Life Technologies), as per the manufacturer’s instructions. No template RNA and no reverse transcriptase controls were included. TaqMan Advanced Master Mix (Life Technologies) and TaqMan Gene Expression assays for KI67 (Mm01278617_m1), NANOG (Hs02387400_g1), OCT4 (At02611156_m1), SOX2 (Hs04234836_s1), ZEB1 (Hs00232783_m1), VIM (Hs00185584_m1), CDH1 (Hs01023894_m1), SNAI1 (Hs00195591_m1), SNAI2 (Hs00161904_m1), and Δ40p53 (as previously described [6]) were used. β-actin (Hs01060665_g1) and GAPDH (Hs02786624_g1) were used as endogenous controls. Relative expression was calculated using the 2−ΔΔCt method [74]. Gene expression analysis of miRNAs was performed using TaqMan Advanced miRNA cDNA Synthesis Kit according to manufacturer’s recommendations (Life Technologies). TaqMan Advanced Master Mix (Life Technologies) and TaqMan Gene Expression assays for hsa-miR-145-5p (477916_mir), hsa-miR-200b-3p (477963_mir), and hsa-miR-200c-3p (478351_mir) were used. Hsa-miR-16-5p (477860_mir) was used as an endogenous control [75].
Gene expression analysis of single cells
MCF-7-LeGO and -Δ40p53 single cells were captured using Fluidigm integrated fluidic circuits with preamplification using TaqMan Assays on the Fluidigm C1 system (Fluidigm, South San Francisco, CA, USA) as per the manufacturer’s instructions. Single-cell qPCR of stem cell markers: SOX2 (Hs04234836_s1), OCT4 (At02611156_m1), NANOG (Hs02387400_g1), ELF5 (Hs01063022_m1), ITGA6 (Hs01041011_m1), ITGB1 (Hs00559595_m1), and FOXM1 (Hs01073586_m1), and the endogenous control GAPDH (Hs02786624_g1) was performed using Biomark HD System with 96.96 Dynamic Array integrated fluidic circuits (Fluidigm, USA). Data was analysed using Singular Analysis Toolset build under R and data are visualised using violin plot.
Immunoblotting
Proteins were separated by sulphate dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) as previously described [18]. The membrane was blocked with Casein Blocking Buffer (Millennium Science, Mulgrave VIC, Australia) at room temperature for 1 h. The following primary antibodies were diluted in blocking buffer: pan-p53 rabbit-anti-human-CM-1 (1 μg/ml; The University of Dundee, Scotland), rabbit-anti-human-Nanog (1 μg/ml; Cell Signaling Technology #D73G4), rabbit-anti-human-Sox2 (1 μg/ml; Cell Signaling Technology #D609), rabbit-anti-human-β-catenin (1 μg/ml; Abcam, Melbourne, VIC, Australia #ab32572), rabbit-anti-human-Δ40p53 KJC40 (2.5 μg/ml), and mouse-anti-human-GAPDH (1 μg/ml; Calbiochem, San Diego, CA, USA #CB1001), and added to the membrane overnight (4 °C, rocking). Diluted secondary antibodies (1–5 μg/ml; LI-COR Biosciences, Lincoln, NE, USA) in blocking buffer were added and allowed to bind on a rocker for at least 1 h at room temperature. Bands were visualised and quantitated using an Odyssey CLx fluorescent imager (LI-COR Biosciences) relative to the loading control (GAPDH). Uncropped membranes are shown in the original western blots Supplemental file.
Flow cytometry and fluorescence-activated cell sorting (FACS) analysis
1 × 106 cells of each MCF-7-LeGO and -Δ40p53 sublines were resuspended in 100 μL of ice-cold 2% FBS in PBS containing allophycocyanin (APC)-conjugated mouse-anti-human CD44 (20 μL; clone G44-26; BD Biosciences, Becton Dickinson Pty Ltd, Macquarie Park, NSW, Australia #550392), BD Horizon Brilliant Violet 421 (BV421)-conjugated mouse anti-human CD24 monoclonal antibody (20 μL; clone ML5; BD Biosciences #562789), and 7-amino-actinomycin D (7-AAD) for 20 min at 4 °C in the dark. The cells were then rinsed twice with 2% FBS in PBS. CD44 and CD24 levels were determined using a BD FACS Aria III flow cytometer (BD Biosciences). Unstained cells were used as negative controls.
Mammospheres formation assay
For mammosphere formation, 1 × 103 cells/well were plated in 24-well ultra-low attachment plates (Corning, NY, USA) with MammoCult medium enriched with MammoCult proliferation supplement (10% v/v), hydrocortisone (0.48 µg/ml), and heparin (4 µg/ml) (StemCell Technologies, Vancouver, BC, Canada) and cultured for 7 days. After 7 days of culture the size and number of formed mammospheres were quantified using a Cytation3 cell imager (BioTek).
Colony formation and cell spheroid assays
For the colony formation assays, 1 × 103 cells/well were seeded onto 6-well plates. Cells were grown for 14 days, at which time visible colonies were apparent. Cells were fixed with ice-cold methanol and stained with 0.5% crystal violet solution. Colony number and size were calculated using the cellSens Standard software (Olympus, Notting Hill, VIC, Australia). For the cell spheroid assay, cells were seeded in 96-well ultra-low attachment plates (Corning) at 4 × 103 cells/well and the plates were centrifuged for 5 min at 200 × g to allow the formation of cell spheroids. The spheroids were imaged after 7 days of culture using a Cytation3 cell imager (BioTek).
Acini formation assay
Three-dimensional acinar assay in extracellular matrix (ECM; Sigma-Aldrich) was performed as previously described [76]. Briefly, 45 µl of ECM was carefully dispensed into 8-well chamber slides and allowed to solidify for at least 30 min in a standard cell culture incubator. MCF-10A cell sublines were trypsinised, washed, and resuspended in complete medium. 5 × 103 cells/well were seeded onto 8-well chamber slides (200 µl/well), and the chamber slides were returned to the cell culture incubator for 30 min for the cells to set onto the precoated ECM. 4% ECM in 200 µl of complete media (for a final concentration of 2% ECM/well) was carefully dispensed into each well. The media containing 2% ECM was replenished every 2 days. Every 7 days of culture, the size and number of formed acinar structures were quantified using a Cytation3 cell imager (BioTek). A threshold of 20 µm was set to filter out cell debris as the size of a MCF-10A cell is ~20 µm. All acini were processed for phalloidin staining after 21 days in culture. Briefly, acini were fixed with 3.7% formaldehyde (Sigma-Aldrich) in PBS for 10 min. The acini were subsequently incubated with TRITC-conjugated phalloidin (10 μg/ml; Sigma-Aldrich) for 20 min. The nuclei were stained with DAPI and mounted using ProLong Gold Antifade mount media (Life Technologies) on glass slides. Confocal microscopy was performed using a Leica DMRE upright fluorescent microscope (Leica, Wetzlar, Hesse, Germany) fitted with a blue argon (488 nm—FITC excitation) and a green helium neon (568 nm—TXR excitation) laser and a 20x objective.
Luciferase lentiviral particles transfection
For in vivo imaging purposes, MCF-7 sublines were transfected with luciferase lentiviral particles according to the manufacturer’s recommendation (GenTarget Inc., San Diego, CA, USA). Briefly, cells were seeded at 2 × 105 cells/well in 6-well plates until they reached 50% of confluence. Next, 500 µL of complete medium and 50 µL of lentiviral particles were added to each well. Media were replenished every 3–4 days with media containing the selection antibiotic, G-418 (400 µg/mL; Sigma-Aldrich), until resistant colonies were identified.
Engraftment of NOD scid gamma mice
Six-week-old female NOD scid gamma (NSG) mice were obtained from the Animal Resources Centre (ARC, Murdoch, WA, Australia), and were kept at the Hunter Medical Research Institute Bioresources Facility at 22 ± 2 °C, with water and food ad libitum, and under a 12:12 h light and dark photoperiod. All experimental procedures were reviewed, approved, and carried out according to the Animal Care and Ethics Committee of the University of Newcastle (approval number: A-2020-016). G*Power 3.1 [77] was used to perform calculations on sample size, effect size, and statistical power. The minimal significance (α) and statistical power (1-β) were set at 0.05 and 0.80, respectively. Calculations were carried out for two groups by using Student’s t-distribution. The NSG mice were orthotopically injected with luciferase-labelled MCF-7 sublines (-shNT, -shΔ40p53, LeGO, or Δ40p53) at 2 × 106 cells suspended in 50:50 matrigel matrix phenol red-free high concentration (Corning)-PBS into the mammary fat pad under isoflurane anaesthesia. Simultaneously, the mice were implanted subcutaneously at the back of the neck with 17β-oestradiol pellets (60-day release, 0.36 mg/pellet; Innovative Research of America, Sarasota, FL, USA). The animals were monitored daily and body weight was recorded twice a week with a digital balance. The tumour take rate was found to be 50%. Tumour burden was assessed twice a week using digital calliper measurements (tumour volume = (length × width × depth)/2). For in vivo imaging, subcutaneous injections of D-luciferin (Sapphire Bioscience, Redfern, NSW, Australia) at a dose of 150 mg/kg were administered to animals under isoflurane anaesthesia. The luminescence signal was recorded using an in vivo imaging system (Xenogen IVIS 100 bioluminescent in-vivo imaging system, PerkinElmer, Waltham, MA, USA). Once tumours were established (50–100 mm3), the treatment regimen was started as detailed below.
In vivo treatment
Treatment 1: Three different doses of DOX were administered via repeated weekly intravenous injections to mice bearing the luciferase-labelled MCF-7-shNT subline to select the best tolerated dose that resulted in a reduction in tumour size. Randomly allocated mice were treated by tail vein intravenous injection with either vehicle (saline) (n = 3), or three different doses of DOX (1 mg/kg, 2 mg/kg, or 3 mg/kg; n = 3 in each treatment group) under isoflurane anaesthesia once a week for 3 weeks. The animals were followed up for 7 days after the last treatment, or until they reached ethical end-point (moribund animal and/or loss of >10% of initial body weight). All animals were euthanised by carbon dioxide asphyxiation (Supplementary Fig. 9A).
Treatment 2: Mice engrafted with luciferase-labelled sublines were randomly divided into eight groups (n = 5–9 mice/group): -shNT (vehicle or DOX-treated), -shΔ40p53 (vehicle or DOX-treated), LeGO (vehicle or DOX-treated), and Δ40p53 (vehicle or DOX-treated). DOX (2 mg/kg) (Supplementary Fig. 9A) or vehicle (saline) were administered once a week for 3 weeks, by intravenous tail vein injection under isofluorane anaesthesia. The identification of the mice (i.e., subline engrafted) was blinded to the investigator during treatment. The animals were monitored daily for clinical and behavioural changes and biweekly for body weight changes (Supplementary Fig. 9B). The animals were followed up for 7 days after the last treatment, or until they reached ethical end-point (moribund animal and/or loss of >10% of initial body weight). All animals were euthanised by carbon dioxide asphyxiation. After euthanasia, the tumours and spleens were harvested and preserved in buffered formalin solution (10%, pH 7.4; Sigma-Aldrich).
Histological analysis
Specimen processing and H&E staining were performed by the Hunter Medical Research Institute Core Histology Facility (Newcastle, NSW, Australia) according to established protocols. Immunohistochemistry (IHC) was performed by the NSW Regional Biospecimen & Research Services (Newcastle, NSW, Australia) using a Ventana Discovery Automated Immunostainer (Roche Medical Systems, Tuscon, AZ, USA) as previously described [25]. Tissue sections (4 µm/section) were deparaffinised and incubated in a Ventana solution for antigen retrieval at pH 9. After antigen retrieval, slides were incubated for 12 min with a peroxidase inhibitor (Roche Medical Systems) followed by incubation with mouse-anti-human-Sox2 (2.5 μg/ml; Life Technologies #MA1-014) or rabbit-anti-human-Ki67 (0.07 μg/ml; Life Technologies #MA5-14520) primary antibodies for 32 min at 37 °C (for slides of positive controls used during antibody optimisation see Supplementary Fig. 10). The pre-diluted anti-mouse hapten (HQ) or anti-rabbit HQ secondary antibodies (Roche Medical Systems) were added and slides were incubated with anti-HQ-horseradish peroxidase (HRP) (Roche Medical Systems), and visualised using diaminobenzidine (DAB) chromogen detection kit (Roche Medical Systems). All slides were manually counterstained with Mayers hematoxylin. Slides were scanned at 40x magnification using an Aperio AT2 scanner (Leica, Wetzlar, Germany), and analysed with HALO Software (Halo imaging analysis software, Indica Labs, Corrales, NM, USA) using the CytoNuclear v2.0.8 analysis mode. H-scores and the percentage of positive cells [7] were quantified. For microvessel area quantification, an analysis mask was created to recognise blood vessels based on the H&E staining (morphology/colour). Tissue artefacts were excluded from the analysis.
H&E slides of the IDC cohort (n = 47) were used to assess TILs. TILs were visualised using HALO Software and manually quantified according to [78].
siRNA transfection
Two siRNAs targeting intron 2 of TP53 were used to knockdown Δ40p53: Δ40p53 #1 (5′-AGACCTGTGGGAAGCGAAA-3′) and Δ40p53 #2 (5′-GCGAAAATTCCATGGGACT-3′) in MCF-7 cells. A non-targeting siRNA was used as a control (D-001810-01-20) (Horizon Discovery, Dharmacon, Millenium Science). For confluence assays, 1 × 104 cells/well were seeded into 96-well plates. For downstream protein expression analysis, 3 × 105 cells/well were seeded in 6-well plates. siRNAs were diluted with 1 x siRNA buffer and serum-reduced media (Opti-MEM) and mixed with DharmaFECT transfection reagent-1 (Millennium Science) diluted in Opti-MEM (Life Technologies). The mixture was diluted in pre-warmed media to achieve a final concentration of 25 nM siRNA. Cells were then treated with DOX (1 µM). For protein analysis, cells were harvested following 24 h of treatment.
Cell confluence and death assay
For real-time cell death assessment and confluence analysis, propidium iodide (PI) at the final concentration of 2.5 µg/mL (Sigma-Aldrich) was added to the wells as previously described [79]. Cells were then placed into an incubator connected to an IncuCyte imaging system (Sartorius, Göttingen, Germany). Images were analysed using the IncuCyte Zoom software. Cell confluence and PI-positive cells normalised to confluence were calculated using integrated software algorithms.
OTSSP167 and DOX treatment
Cells were treated with DOX (1 µM) and/or the Melk inhibitor, OTSSP167 (40 nM) (Selleckchem, Houston, TX, USA). The effects of the treatments were assessed by mammosphere and cell spheroid formation assays as described above. For the mammosphere assay, the size and number of formed mammospheres were quantified after 7 days of treatment. For the cell spheroid assay, spheroids were imaged every other day and on the seventh day of treatment, half of the media was replenished and additional treatment was added to the wells. After 14 days of treatment, spheroid size was recorded and spheroid viability was evaluated using CellTiter-Glo 3D assay as per manufacturer’s recommendations (Promega, Madison, WI, USA). Images of the created mammospheres, spheroids, and luminescence was recorded using a Cytation3 cell imager (BioTek) and analysed using Gen5 software.
Statistical analysis
All continuous variables were tested for normal distribution. Unpaired student t-tests or Mann–Whitney tests were performed for two comparisons. For multiple comparisons, one-way ANOVA corrected for multiple comparisons using the Dunnett’s or Tukey’s tests or two-way ANOVA corrected for multiple comparisons using the Sidak’s or Tukey’s tests were performed. All results are the mean of three independent experiments, and error bars represent the standard deviation (SD) or the standard error of the mean (SEM). Spearman rank correlation analysis was used to compare the relative mRNA expression of Δ40p53 with Ki67 in tumour tissues. All statistical analyses were performed using GraphPad Prism v. 9.0 (GraphPad Software, La Jolla, CA, USA). An adjusted p-value of <0.05 was considered statistically significant.
Data availability
Data from HumanGene1.0 Arrays were deposited in the NCBI Gene Expression Omnibus database with the accession number GSE61725. Other data generated in this study are available within the supplementary data files or upon request from the corresponding author.
References
Liang Y, Zhang H, Song X, Yang Q. Metastatic heterogeneity of breast cancer: molecular mechanism and potential therapeutic targets. Semin Cancer Biol. 2020;60:14–27.
Miller LD, Smeds J, George J, Vega VB, Vergara L, Ploner A, et al. An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. PNAS. 2005;102:13550–5.
Coutant C, Rouzier R, Qi Y, Lehmann-Che J, Bianchini G, Iwamoto T, et al. Distinct p53 gene signatures are needed to predict prognosis and response to chemotherapy in ER-positive and ER-negative breast cancers. Clin Cancer Res. 2011;17:2591–601.
Bourdon JC, Fernandes K, Murray-Zmijewski F, Liu G, Diot A, **rodimas DP, et al. p53 isoforms can regulate p53 transcriptional activity. Genes Dev. 2005;19:2122–37.
Arsic N, Slatter T, Gadea G, Villain E, Fournet A, Kazantseva M, et al. Δ133p53β isoform pro-invasive activity is regulated through an aggregation-dependent mechanism in cancer cells. Nat Commun. 2021;12:5463.
Avery-Kiejda KA, Morten B, Wong-Brown MW, Mathe A, Scott RJ. The relative mRNA expression of p53 isoforms in breast cancer is associated with clinical features and outcome. Carcinogenesis. 2014;35:586–96.
Kazantseva M, Eiholzer RA, Mehta S, Taha A, Bowie S, Roth I, et al. Elevation of the TP53 isoform Δ133p53β in glioblastomas: an alternative to mutant p53 in promoting tumor development. J Pathol. 2018;246:77–88.
Steffens Reinhardt L, Zhang X, Wawruszak A, Groen K, De Iuliis GN, Avery-Kiejda KA. Good cop, bad cop: defining the roles of Δ40p53 in cancer and aging. Cancers 2020;12:1659.
Eiholzer RA, Mehta S, Kazantseva M, Drummond CJ, McKinney C, Young K, et al. Intronic TP53 polymorphisms are associated with increased Δ133TP53 transcript, immune infiltration and cancer risk. Cancers. 2020;12:2472.
Tadijan A, Precazzini F, Hanžić N, Radić M, Gavioli N, Vlašić I, et al. Altered expression of shorter p53 family isoforms can impact melanoma aggressiveness. Cancers. 2021;13:5231.
Morten BC, Wong-Brown MW, Scott RJ, Avery-Kiejda KA. The presence of the intron 3 16 bp duplication polymorphism of p53 (rs17878362) in breast cancer is associated with a low Delta40p53:p53 ratio and better outcome. Carcinogenesis. 2016;37:81–6.
Zang Y, Shi Y, Liu K, Qiao L, Guo X, Chen D. Δ40p53 is involved in the inactivation of autophagy and contributes to inhibition of cell death in HCT116-Δ40p53 cells. Oncotarget. 2017;8:12754–63.
Horikawa I, Park KY, Isogaya K, Hiyoshi Y, Li H, Anami K, et al. Δ133p53 represses p53-inducible senescence genes and enhances the generation of human induced pluripotent stem cells. Cell Death Differ. 2017;24:1017–28.
Gong H, Zhang Y, Jiang K, Ye S, Chen S, Zhang Q, et al. p73 coordinates with Δ133p53 to promote DNA double-strand break repair. Cell Death Differ. 2018;25:1063–79.
Campbell H, Fleming N, Roth I, Mehta S, Wiles A, Williams G, et al. ∆133p53 isoform promotes tumour invasion and metastasis via interleukin-6 activation of JAK-STAT and RhoA-ROCK signalling. Nat Commun. 2018;9:254.
Gong L, Pan X, Abali GK, Little JB, Yuan ZM. Functional interplay between p53 and Δ133p53 in adaptive stress response. Cell Death Differ. 2020;27:1618–32.
Levandowski CB, Jones T, Gruca M, Ramamoorthy S, Dowell RD, Taatjes DJ. The Δ40p53 isoform inhibits p53-dependent eRNA transcription and enables regulation by signal-specific transcription factors during p53 activation. PLoS Biol. 2021;19:e3001364.
Zhang X, Groen K, Morten BC, Steffens Reinhardt L, Campbell HG, Braithwaite AW, et al. The effect of p53 and its N-terminally truncated isoform, Δ40p53, on breast cancer migration and invasion. Mol Oncol. 2021;16:447–65.
Guo Y, Rall-Scharpf M, Bourdon J-C, Wiesmüller L, Biber S. p53 isoforms differentially impact on the POLι dependent DNA damage tolerance pathway. Cell Death Dis. 2021;12:941.
Steffens Reinhardt L, Zhang X, Groen K, Morten BC, De Iuliis GN, Braithwaite AW, et al. Alterations in the p53 isoform ratio govern breast cancer cell fate in response to DNA damage. Cell Death Dis. 2022;13:907.
Avery-Kiejda KA, Zhang XD, Adams LJ, Scott RJ, Vojtesek B, Lane DP, et al. Small molecular weight variants of p53 are expressed in human melanoma cells and are induced by the DNA-damaging agent cisplatin. Clin Cancer Res. 2008;14:1659–68.
Powell DJ, Hrstka R, Candeias M, Bourougaa K, Vojtesek B, Fahraeus R. Stress-dependent changes in the properties of p53 complexes by the alternative translation product p53/47. Cell Cycle. 2008;7:950–9.
Ungewitter E, Scrable H. Delta40p53 controls the switch from pluripotency to differentiation by regulating IGF signaling in ESCs. Genes Dev. 2010;24:2408–19.
Takahashi R, Giannini C, Sarkaria JN, Schroeder M, Rogers J, Mastroeni D, et al. p53 isoform profiling in glioblastoma and injured brain. Oncogene. 2013;32:3165–74.
Steffens Reinhardt L, Groen K, Morten BC, Bourdon J-C, Avery-Kiejda KA. Cytoplasmic p53β isoforms are associated with worse disease-free survival in breast cancer. Int J Mol Sci. 2022;23:6670.
Yeong J, Lim JCT, Lee B, Li H, Chia N, Ong CCH, et al. High densities of tumor-associated plasma cells predict improved prognosis in triple negative breast cancer. Front Immunol. 2018;9:1209.
Ghatak D, Das Ghosh D, Roychoudhury S. Cancer stemness: p53 at the wheel. Front Oncol. 2020;10:604124.
Chakrabarti R, Hwang J, Andres Blanco M, Wei Y, Lukačišin M, Romano RA, et al. Elf5 inhibits the epithelial-mesenchymal transition in mammary gland development and breast cancer metastasis by transcriptionally repressing Snail2. Nat Cell Biol. 2012;14:1212–22.
Li X, Li S, Li B, Li Y, Aman S, **a K, et al. Acetylation of ELF5 suppresses breast cancer progression by promoting its degradation and targeting CCND1. NPJ Precis Oncol. 2021;5:20.
Wafai R, Williams ED, de Souza E, Simpson PT, McCart Reed AE, Kutasovic JR, et al. Integrin alpha-2 and beta-1 expression increases through multiple generations of the EDW01 patient-derived xenograft model of breast cancer—insight into their role in epithelial mesenchymal transition in vivo gained from an in vitro model system. Breast Cancer Res. 2020;22:136.
Drápela S, Bouchal J, Jolly MK, Culig Z, Souček K. ZEB1: a critical regulator of cell plasticity, DNA damage response, and therapy resistance. Front Mol Biosci. 2020;7:36.
Ganesan R, Mallets E, Gomez-Cambronero J. The transcription factors Slug (SNAI2) and Snail (SNAI1) regulate phospholipase D (PLD) promoter in opposite ways towards cancer cell invasion. Mol Oncol. 2016;10:663–76.
Rajarajan D, Kaur B, Penta D, Natesh J, Meeran SM. miR-145–5p as a predictive biomarker for breast cancer stemness by computational clinical investigation. Comput Biol Med. 2021;135:104601.
Xu N, Papagiannakopoulos T, Pan G, Thomson JA, Kosik KS. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell. 2009;137:647–58.
Eggers JC, Martino V, Reinbold R, Schäfer SD, Kiesel L, Starzinski-Powitz A, et al. microRNA miR-200b affects proliferation, invasiveness and stemness of endometriotic cells by targeting ZEB1, ZEB2 and KLF4. Reprod Biomed Online. 2016;32:434–45.
Chang C, Chao C, **a W, Yang J, **ong Y, Li C, et al. p53 regulates epithelial-mesenchymal transition and stem cell properties through modulating miRNAs. Nat Cell Biol. 2011;13:317–23.
Li W, Wang Y, Liu R, Kasinski AL, Shen H, Slack FJ, et al. MicroRNA-34a: potent tumor suppressor, cancer stem cell inhibitor, and potential anticancer therapeutic. Front Cell Dev Biol. 2021;9:640587.
Ravichandran Y, Goud B, Manneville J-B. The Golgi apparatus and cell polarity: roles of the cytoskeleton, the Golgi matrix, and Golgi membranes. Curr Opin Cell Biol. 2020;62:104–13.
Vantangoli MM, Madnick SJ, Huse SM, Weston P, Boekelheide K. MCF-7 human breast cancer cells form differentiated microtissues in scaffold-free hydrogels. PLoS ONE. 2015;10:e0135426.
Debnath J, Mills KR, Collins NL, Reginato MJ, Muthuswamy SK, Brugge JS. The role of apoptosis in creating and maintaining luminal space within normal and oncogene-expressing mammary acini. Cell. 2002;111:29–40.
Singh A, Settleman J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene. 2010;29:4741–51.
Zhang Z, Sun C, Li C, Jiao X, Griffin BB, Dongol S, et al. Upregulated MELK leads to doxorubicin chemoresistance and M2 macrophage polarization via the miR-34a/JAK2/STAT3 pathway in uterine leiomyosarcoma. Front Oncol. 2020;10:453.
Ren L, Deng B, Saloura V, Park JH, Nakamura Y. MELK inhibition targets cancer stem cells through downregulation of SOX2 expression in head and neck cancer cells. Oncol Rep. 2019;41:2540–8.
Bollu LR, Shepherd J, Zhao D, Ma Y, Tahaney W, Speers C, et al. Mutant P53 induces MELK expression by release of wild-type P53-dependent suppression of FOXM1. NPJ Breast Cancer. 2020;6:2.
Ji W, Arnst C, Tipton AR, Bekier ME 2nd, Taylor WR, Yen TJ, et al. OTSSP167 abrogates mitotic checkpoint through inhibiting multiple mitotic kinases. PLoS ONE. 2016;11:e0153518.
Joruiz SM, Beck JA, Horikawa I, Harris CC. The Δ133p53 isoforms, tuners of the p53 pathway. Cancers. 2020;12:3422.
Arsic N, Gadea G, Lagerqvist EL, Busson M, Cahuzac N, Brock C, et al. The p53 isoform Delta133p53beta promotes cancer stem cell potential. Stem Cell Rep. 2015;4:531–40.
Solomon H, Bräuning B, Fainer I, Ben-Nissan G, Rabani S, Goldfinger N, et al. Post-translational regulation of p53 function through 20S proteasome-mediated cleavage. Cell Death Differ. 2017;24:2187–98.
Haronikova L, Olivares-Illana V, Wang L, Karakostis K, Chen S, Fåhraeus R. The p53 mRNA: an integral part of the cellular stress response. Nucleic Acids Res. 2019;47:3257–71.
Sun X, Jiao X, Pestell TG, Fan C, Qin S, Mirabelli E, et al. MicroRNAs and cancer stem cells: the sword and the shield. Oncogene. 2014;33:4967–77.
Singh SK, Chen NM, Hessmann E, Siveke J, Lahmann M, Singh G, et al. Antithetical NFATc1-Sox2 and p53-miR200 signaling networks govern pancreatic cancer cell plasticity. EMBO J. 2015;34:517–30.
Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S, et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 2008;9:582–9.
Ren D, Wang M, Guo W, Zhao X, Tu X, Huang S, et al. Wild-type p53 suppresses the epithelial-mesenchymal transition and stemness in PC-3 prostate cancer cells by modulating miR‑145. Int J Oncol. 2013;42:1473–81.
Phang BH, Othman R, Bougeard G, Chia RH, Frebourg T, Tang CL, et al. Amino-terminal p53 mutations lead to expression of apoptosis proficient p47 and prognosticate better survival, but predispose to tumorigenesis. Proc Natl Acad Sci USA. 2015;112:E6349–E58.
Jain AK, Allton K, Iacovino M, Mahen E, Milczarek RJ, Zwaka TP, et al. p53 regulates cell cycle and microRNAs to promote differentiation of human embryonic stem cells. PLoS Biol. 2012;10:e1001268.
Lin T, Chao C, Saito S, Mazur SJ, Murphy ME, Appella E, et al. p53 induces differentiation of mouse embryonic stem cells by suppressing Nanog expression. Nat Cell Biol. 2005;7:165–71.
Huang Y-H, Luo M-H, Ni Y-B, Tsang JYS, Chan S-K, Lui PCW, et al. Increased SOX2 expression in less differentiated breast carcinomas and their lymph node metastases. Histopathology. 2014;64:494–503.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Khromova NV, Kopnin PB, Stepanova EV, Agapova LS, Kopnin BP. p53 hot-spot mutants increase tumor vascularization via ROS-mediated activation of the HIF1/VEGF-A pathway. Cancer Lett. 2009;276:143–51.
Melo Dos Santos N, de Oliveira GAP, Ramos Rocha M, Pedrote MM, Diniz da Silva Ferretti G, Pereira Rangel L, et al. Loss of the p53 transactivation domain results in high amyloid aggregation of the Delta40p53 isoform in endometrial carcinoma cells. J Biol Chem. 2019;294:9430–9.
Hafsi H, Santos-Silva D, Courtois-Cox S, Hainaut P. Effects of Delta40p53, an isoform of p53 lacking the N-terminus, on transactivation capacity of the tumor suppressor protein p53. BMC Cancer. 2013;13:134.
Ota A, Nakao H, Sawada Y, Karnan S, Wahiduzzaman M, Inoue T, et al. Delta40p53alpha suppresses tumor cell proliferation and induces cellular senescence in hepatocellular carcinoma cells. J Cell Sci. 2017;130:614–25.
Nutthasirikul N, Limpaiboon T, Leelayuwat C, Patrakitkomjorn S, Jearanaikoon P. Ratio disruption of the ∆133p53 and TAp53 isoform equilibrium correlates with poor clinical outcome in intrahepatic cholangiocarcinoma. Int J Oncol. 2013;42:1181–8.
Tu Q, Gong H, Yuan C, Liu G, Huang J, Li Z, et al. Δ133p53/FLp53 predicts poor clinical outcome in esophageal squamous cell carcinoma. Cancer Manag Res. 2020;12:7405–17.
Bouchie A. First microRNA mimic enters clinic. Nat Biotechnol. 2013;31:577.
Hong DS, Kang Y-K, Borad M, Sachdev J, Ejadi S, Lim HY, et al. Phase 1 study of MRX34, a liposomal miR-34a mimic, in patients with advanced solid tumours. Brit J Cancer. 2020;122:1630–7.
Kravchenko JE, Ilyinskaya GV, Komarov PG, Agapova LS, Kochetkov DV, Strom E, et al. Small-molecule RETRA suppresses mutant p53-bearing cancer cells through a p73-dependent salvage pathway. Proc Natl Acad Sci USA. 2008;105:6302–7.
Mathe A, Wong-Brown M, Morten B, Forbes JF, Braye SG, Avery-Kiejda KA, et al. Novel genes associated with lymph node metastasis in triple negative breast cancer. Sci Rep. 2015;5:15832.
Team RC. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2022. https://www.R-project.org/.
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. The Innovation. 2021;2:100141.
Yu G _enrichplot: Visualization of Functional Enrichment Result_. R package version 1.16.1; 2022. https://yulab-smu.top/biomedical-knowledge-mining-book/.
Kolde R. pheatmap: Pretty Heatmaps. 2019. https://CRAN.R-project.org/package=pheatmap
Zaqout S, Becker L-L, Kaindl AM. Immunofluorescence staining of paraffin sections step by step. Front Neuroanat. 2020;14:582218.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–8.
Davoren PA, McNeill RE, Lowery AJ, Kerin MJ, Miller N. Identification of suitable endogenous control genes for microRNA gene expression analysis in human breast cancer. BMC Mol Biol. 2008;9:76.
Lee GY, Kenny PA, Lee EH, Bissell MJ. Three-dimensional culture models of normal and malignant breast epithelial cells. Nat Methods. 2007;4:359–65.
Faul F, Erdfelder E, Buchner A, Lang AG. Statistical power analyses using G*Power 3.1: tests for correlation and regression analyses. Behav Res Methods. 2009;41:1149–60.
Salgado R, Denkert C, Demaria S, Sirtaine N, Klauschen F, Pruneri G, et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs Working Group 2014. Ann Oncol. 2015;26:259–71.
Szalai P, Engedal N. An image-based assay for high-throughput analysis of cell proliferation and cell death of adherent cells. Bio Protoc. 2018;8:e2835.
Acknowledgements
The authors would like to thank Ms Carolyn Allport for the exceptional assistance with the animals’ care, Dr Heather Murray, Dr Abdul Mannan, and Dr Severine Roselli for assistance with the establishment of the animal model, Dr Min Yuan Quah for assistance with the in vivo imaging, Dr Sean Burnard for assistance with bioinformatics analyses, the NSW Regional Biospecimen & Research Services team, especially Ms Clarke and Ms O’Brien, for performing the IHC, the HMRI Core Histology, especially Ms Clout and Ms Bielanowicz, for sample processing, and Dr Vilain for the assistance with the histological evaluation.
Funding
This work was funded by the Hunter Cancer Research Alliance (Biomarkers and Targeted Therapies Flagship), the Cancer Institute NSW, and the Hunter Medical Research Institute. LSR is supported by a University of Newcastle International Postgraduate Research Scholarship and a University of Newcastle Research Scholarship External. AW is supported by The Iwanowska Programme, The Polish National Agency for Academic Exchange PPN/IWA/2018/1/00005 grant. KAAK is supported by a Cancer Institute NSW Career Development Fellowship (CDF181205).
Author information
Authors and Affiliations
Contributions
LSR and KAAK conceived and designed the project. LSR acquired and analysed the data. XZ performed acini formation and gene expression assays. BCM performed the single-cell analysis. AW carried out the cell sorting. LSR and KG performed the in vivo experiments. LSR, KG, and KAAK interpreted the data. LSR wrote the paper. KG and KAAK supervised students working on the project. XZ, BCM, and AW provided feedback to the final draft of the manuscript. KAAK obtained funding. LSR, KG, and KAAK reviewed and edited the final draft of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The experiments with patient samples were conducted in accordance with the Helsinki Declaration with ethical approval from the Hunter New England Human Research Ethics Committee (approval number: 09/05/20/5.02) and the University of Newcastle Health and Safety Committee (approval number: R7/2021). The experiments with animals were reviewed, approved, and carried out according to the Animal Care and Ethics Committee of the University of Newcastle (approval number: A-2020-016).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Ute Moll
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Steffens Reinhardt, L., Groen, K., Zhang, X. et al. p53 isoform expression promotes a stemness phenotype and inhibits doxorubicin sensitivity in breast cancer. Cell Death Dis 14, 509 (2023). https://doi.org/10.1038/s41419-023-06031-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-023-06031-4
- Springer Nature Limited