Abstract
Parathyroid hormone (PTH) and PTH-related peptide (PTHrP) are two endogenous hormones recognized by PTH receptor-1 (PTH1R), a member of class B G protein- coupled receptors (GPCRs). Both PTH and PTHrP analogs including teriparatide and abaloparatide are approved drugs for osteoporosis, but they exhibit distinct pharmacology. Here we report two cryo-EM structures of human PTH1R bound to PTH and PTHrP in the G protein-bound state at resolutions of 2.62 Å and 3.25 Å, respectively. Detailed analysis of these structures uncovers both common and unique features for the agonism of PTH and PTHrP. Molecular dynamics (MD) simulation together with site-directed mutagenesis studies reveal the molecular basis of endogenous hormones recognition specificity and selectivity to PTH1R. These results provide a rational template for the clinical use of PTH and PTHrP analogs as an anabolic therapy for osteoporosis and other disorders.
Similar content being viewed by others
Introduction
Parathyroid hormone (PTH) and PTH-related protein (PTHrP) are two endogenous peptide hormones that play key and distinct biological roles in skeletal development, calcium- and phosphate-regulating actions, and bone turn over [1, 2]. Lack of functional PTH results in hypoparathyroidism [3], while overexpression of PTHrP derived from tumor cells is the most common reason for cancer-associated hypercalcemia [4]. Both PTH and PTHrP also have potent cardiovascular effects, including stimulating heart rate increase independent of autonomic reflexes [5]. Both PTH and PTHrP exert their effects on bone by activating the PTH type 1 receptor (PTH1R), a class B G protein-coupled receptor (GPCR), primarily via Gs-cAMP signaling [1, 6]. High-resolution structures of PTH1R-Gs complexes bound to these endogenous hormones are important for both pharmacological research as well as clinical development aiming at PTH1R.
Osteoporosis is a disease of decreased bone mass, microarchitectural deterioration, and fragility fractures. It is widespread and can affect all ethnic groups. Osteoporosis becomes a major clinical problem in older women and men, especially in postmenopausal women [7]. Clinical trials were proved that both PTH and PTHrP can stimulate bone anabolism in patients with osteoporosis [6, 8]. Teriparatide, recombinant human parathyroid hormone 1–34 (PTH1–34) [9], and abaloparatide (ABL), an investigational analog of human PTHrP1–34 [6, 10], are two synthetic analogs of PTH and PTHrP. Although they exhibit distinct pharmacology, they were the only two approved drugs by FDA for the treatment of osteoporosis [6]. Recent studies suggest that PTH and PTHrP differ in their relative capacities to bind to two pharmacologically distinguishable high-affinity PTH1R conformations, R0 (the G protein-uncoupled PTH1R conformation) and RG (the G protein-coupled PTH1R conformation) [11, 12]. PTH1–34 can bind to the R0 conformation with the higher affinity than PTHrP1–36 and prolonged cAMP responses [12]. Whereas PTHrP1–36 binds weakly to R0, but it preferentially binds to the RG, promoted by the overexpression of a high affinity of Gαs and produce shorter cAMP responses [6, 12]. Hattersley et al. [13] found that abaloparatide has similar high affinity to the RG as teriparatide, and abaloparatide shows more selectivity for the RG and a shorter cAMP signaling response compared to teriparatide. Abaloparatide and teriparatide different selectivity for the RG conformation can explain the different increase in bone mineral density (BMD) and different incidence of hypercalcemia in the clinical trials treated by abaloparatide and teriparatide [6]. Such conformational selectivity of receptor binding can be a key regulator of that ligand’s biological activity, which provides an important clue to optimize conformational selective PTH and PTHrP analogs for a new treatment option for PTH-related disorders [14].
Up to now, the most effective treatment of osteoporosis is based on daily injections teriparatide or abaloparatide. These therapies are both costly and inconvenient for patients. There are no orally available PTH1R agonist drugs for osteoporosis. The detailed structural basis and mechanistic understanding of the receptor activation by endogenous peptide hormones are critical for aiding the rational development of new drug therapies targeting PTH1R. The only available structure of the full-length PTH1R is in complex with a synthetic long-acting PTH (LA-PTH), which reveals extensive interactions of LA-PTH with PTH1R [2]. Therefore, we determined two additional cryo-electron microscopy (cryo-EM) structures of the active PTH1R-Gs complexes bound to PTH and PTHrP, providing insights into the pharmacological mechanisms of the activation of PTH1R by different endogenous peptide hormones.
Materials and methods
Constructs
The human PTH1R (residues 27–502) with G188A and K484R mutations was cloned into pFastBac vector from Invitrogen (Carlsbad, CA, USA) with the haemagglutinin signal peptide (HA) followed by a TEV protease cleavage site and a double MBP (2MBP) and His tag to facilitate expression and purification [2]. To facilitate a stable complex, the above PTH1R construct was added the LgBiT subunit from Promega (Madison, WI, USA) at the C terminus of PTH1R [15]. It is different from the construct used for mammalian expression system in the cryo-EM structures of PTH1R bound to PTH and PTHrP recently reported by Kobayashi et al. [16]. Bovine dominant-negative Gαs (DNGαs) has two mutations G226A and A366S for increasing the dominant-negative effect to form a stable heterotrimer complex with Gβγ. The α-helical domain (AHD F68–S205) of DNGαs was replaced with human Gαi (AHD Y61–T182) for binding the Fab_G50 [2]. Based on the published DNGαs, a modified bovine Gαs (mDNGαs), its N terminus (M1–K25) and α-helical domain (AHD F68–L203) of Gαs were replaced with the N terminus (M1–M18) and AHD (Y61–K180) of the human Gαi, which can bind scFv16 and Fab_G50 [17]. The residues N254–T263 of Gαs were deleted and eight mutations (G49D, E50N, L63Y, A249D, S252D, L272D, I372A, and V375I) were added. It is also different from mini-Gs expressed in E. coli BL21 cells used by Kobayashi et al. [16]. To facilitate the folding of the G protein, DNGαs and mDNGαs were co-expressed with GST-Ric-8B, respectively [18]. Rat Gβ1 was fused with a His16 tag at the N terminus and with a SmBiT subunit (peptide 86) from Promega (Madison, WI, USA) [19], following a 15-amino acid (5′-GSSGGGGSGGGGSSG-3′) linker at its C terminus. In addition, to clone the constructs into the pcDNA3.0 vector from Promega (Madison, WI, USA) for cAMP accumulation, all constructs were cloned using ClonExpressII one step cloning kit (Vazyme Biotech Co., Ltd, Nan**g, China).
Protein expression
To facilitate a stable complex and purification, the PTH1R and G proteins were co-expressed in Sf9 insect cells from Invitrogen (Carlsbad, CA, USA). The Sf9 cell line growing to a density of 3.5 × 106 cells/mL in ESF 921 cell culture medium from Expression Systems (Davis, CA, USA) is good for expression. We infected the cells with five separate virus preparations at a ratio of 1:2:2:2:2, including PTH1R (27–502)-LgBiT-2MBP, DNGαs or mDNGαs, His16-Gβ1-peptide 86, Gγ2, and GST-Ric-8B. The infected cells were cultured at 27 °C for 48 h, the cells were harvested by centrifugation and washed with PBS once. The cell pellets were frozen at −80 °C for further usage.
Expression and purification of Nb35
Nanobody-35 (Nb35) was expressed in E. coli BL21 cells, the cultured cells were grown in 2TB media with 100 μg/mL ampicillin, 2 mM MgCl2, 0.1% glucose at 37 °C for 2.5 h until OD600 of 0.7–1.2 was reached. Then the culture was induced with 1 mM IPTG at 37 °C for 4–5 h, and harvested and frozen at −80 °C for further purification. Nb35 was purified by nickel affinity chromatography and followed by size-exclusion chromatography using HiLoad 16/600 Superdex 75 column from GE Healthcare (Boston, MA, USA) or following overnight dialysis against 20 mM HEPES, pH 7.4, 100 mM NaCl, 10% glycerol. The quality of purified protein was verified by SDS-PAGE and stored at −80 °C.
Complex purification
The complexes were purified according to previously described methods [2, 20]. The cell pellets were resuspended in 20 mM HEPES pH 7.4, 100 mM NaCl, 10 mM MgCl2, 10 mM CaCl2, 2 mM MnCl2, 10% glycerol, 0.1 mM TCEP, 15 μg/mL Nb35, 25 mU/mL apyrase from Sigma (St. Louis, MO, USA), 12 µM PTH1–34 (Synpeptide Co., Ltd, Nan**g, China) or 12 µM PTHrP1–36 (TGpeptide Biotechnology Co., Ltd, Nan**g, China), supplemented with 1 mL protease inhibitor cocktail (TargetMol, Boston, MA, USA) into 100 mL suspension. The lysate was incubated for 1 h at room temperature and then solubilized by 0.5% (w/v) lauryl maltose neopentylglycol (LMNG, Anatrace, Maumee, OH, USA) supplemented with 0.1% (w/v) cholesteryl hemisuccinate TRIS salt (CHS, Anatrace, Maumee, OH, USA) for 2 h at 4 °C. The supernatant of the solubilized membranes was collected by centrifugation at 65,000 × g for 40 min, then incubated with Amylose resin (Smart-lifesciences, Changzhou, China) for 2 h at 4 °C. The resin was loaded onto a gravity flow column and washed with 20 column volumes of 20 mM HEPES, pH 7.4, 100 mM NaCl, 10% glycerol, 5 mM CaCl2, 5 mM MgCl2, 1 mM MnCl2, 0.01% (w/v) LMNG, 0.01% glyco-diosgenin (GDN, Anatrace, Maumee, OH, USA) and 0.004% (w/v) CHS, 2 µM PTH1–34 or 2 µM PTHrP1–36, and 25 μM TCEP. After washing, the protein was cut with TEV protease on column overnight at 4 °C. Next day the flow through was collected and concentrated, then PTH-PTH1R-Gs or PTHrP-PTH1R-Gs flow through was loaded onto a Superdex200 10/300 GL column from GE Healthcare (Boston, MA, USA), with the buffer consisting of 20 mM HEPES, pH 7.4, 100 mM NaCl, 2 mM MgCl2, 0.00075% (w/v) LMNG, 0.00025% GDN, 0.0005% (w/v) digitonin from Biosynth (Staad, Switzerland), 0.0002% (w/v) CHS, 2 µM PTH1–34 or 2 µM PTHrP1–36, and 100 μM TCEP. The complex fractions were collected and concentrated for electron microscopy experiments.
Cryo-EM data acquisition
For the preparation of cryo-EM grids, 2.5 μL of the purified PTH-PTH1R-Gs and PTHrP-PTH1R-Gs complexes at a concentration of ~8.4 mg/mL and ~5.1 mg/mL were respectively applied to the glow-discharged Au 300 mesh holey carbon grids (Quantifoil R1.2/1.3, Germany). The grids were blotted and then plunge-frozen in liquid ethane using a Vitrobot Mark IV from Thermo Fisher Scientific (Waltham, MA, USA) at 4 °C.
Cryo-EM images of PTH-PTH1R-Gs were collected on a Titan Krios G4 equipped with a Gatan K3 direct electron detector with super-resolution mode and EPU from Thermo Fisher Scientific (Waltham, MA, USA) were used to acquire cryo-EM movies at Advanced Center for Electron Microscopy at Shanghai Institute of Materia Medica, Chinese Academy of Sciences. A total of 8002 movies were recorded with pixel size of 0.824 Å at a dose of 50 electron per Å2 for 36 frames. The defocus range of this dataset was −0.8 µm to −1.8 µm. For dimer complex, another 5054 movies were obtained with same parameters.
Cryo-EM images of the PTHrP-PTH1R-Gs complex were collected on a Titan Krios G4 at 300KV accelerating voltage equipped with Gatan K3 direct electron detector. A total of 7266 movies were recorded with pixel size of 0.824 Å at a dose of 50 electron per Å2 for 36 frames. The defocus range of this dataset was −0.8 µm to −1.8 µm.
Image processing
All dose-fractionated image stacks were subjected to beam-induced motion correction by RELION-4.0 [21]. The defocus parameters were estimated by CTFFIND 4.1 [22]. PTH-PTH1R-Gs complex and dimer complex, by auto-picking yielded 3,051,218 particles and 10,497,057 which were processed by reference-free 2D classification using Cryosparc [23]. With initial model, after several rounds of 3D classification using RELION, 407,467 particles and 55,858 particles were used to further refinement and polishing, yielding two reconstructions with global resolution of 2.62 Å and 3.76 Å respectively, and subsequently post-processed by DeepEMhancer [24].
For the PTHrP-PTH1R-Gs complex, movies were aligned with RELION-4.0 [21]. Initial contrast transfer function (CTF) fitting was performed with CTFFIND 4.1 [22] from Cryosparc [23]. Auto-picking and two-dimensional (2D) Classification were processed using Cryosparc, producing 7,700,728 particles for further processing. With initial model, two rounds of 3D classifications were carried out. A further one round of 3D classification was conducted with a mask on the receptor, in which 436,359 particles were subjected to 3D auto-refinement and polishing. A map with an indicated global resolution of 3.25 Å at a Fourier shell correlation (FSC) of 0.143 was generated from the final 3D refinement, and subsequently post-processed by DeepEMhancer [24].
Model building and refinement
The cryo-EM structure of the LA-PTH1R-Gs-Nb35 complex (PDB code: 6NBF) and two ECD crystal structures (PDB: 3C4M and PDB: 3H3G) were used as the start for model building and refinement against the electron microscopy map. The model was docked into the electron microscopy density map using Chimera [25], followed by iterative manual adjustment and rebuilding in COOT [26]. Real space and Rosetta refinements were performed using Phenix [27]. The model statistics were validated using MolProbity [28]. Fitting of the refined model to the final map was analyzed using model-versus-map FSC. To monitor the potential over-fitting in model building, FSCwork and FSCfree were determined by refining ‘shaken’ models against unfiltered half-map-1 and calculating the FSC of the refined models against unfiltered half-map-1 and half-map-2. The final refinement statistics are provided in Supplementary Table S2. Structural figures were prepared in Chimera and PyMOL (https://pymol.org/2/).
Molecular dynamics simulations
The cryo-EM structures of PTH-PTH1R-Gs complex and PTHrP-PTH1R-Gs complex were used for the construction of MD simulation systems. Missing ICL3 (5 residues for both systems) and short part before TM1 (6 residues for PTH-PTH1R only) were supplied according to the loop builder program in Molecular Operation Environment. Missing long N-terminal (56 residues for PTH-PTH1R, 51 residues for PTHrP-PTH1R) and ECL1 (31 residues for PTH-PTH1R, 30 residues for PTHrP-PTH1R) structures were constructed according to the AlphaFold2 models [29]. In using AlphaFold2 models, we locally aligned the model to cryo-EM structure with respect to the residues around the missing part until local Cα RMSD was less than 0.3 Å. The mutation G188A was modified back and the G proteins were removed. Using CHARMM-GUI, the models were inserted in POPC (palmitoyl-2-oleoyl-sn-glycerol-3-phosphocholine) lipids [30, 31]. The length of X and Y were both 130 Å. TIP3P waters and 0.15 M NaCl under a periodic boundary condition were also added to systems. The force fields for proteins and lipids were FF19SB and lipid17, respectively [31,32,33]. We used hydrogen mass repartitioning during system construction.
After preparation, the systems first encountered a minimization process of 5000 steepest descent cycles with a constraint on backbone atom, sidechain atom, and lipid coordinates. Then, the constraints were generally decreased in the separated 6 steps of the equilibration process provided by CHARMM-GUI. The first three of them had a timestep of 1 fs and the system was heated to 303.15 K, 1 atm in 375 ps. The last three of them had 2 fs timestep and decreased constrain in the coming 1.5 ns. We then performed the independent 400 ns × 6 MD simulations on Amber 20. During simulations, the Langevin thermostat and Berendsen barostat were used for temperature (300 K) and pressure (1 atm) control. Long-range electrostatic interactions were treated by the Particle mesh Ewald and a 10 Å cutoff was employed for short-range interactions. In the following analysis, tICA was calculated in MSMbuilder [34] and the representative structures were extracted by CPPTRAJ [35]. MMGBSA were calculated by Maestro, Schrödinger. The decomposition of binding free energy was calculated by Amber MMPBSA.py [36].
cAMP accumulation assay
PTH and PTHrP stimulated cAMP accumulation was measured by a LANCE Ultra cAMP kit from PerkinElmer (Boston, MA, USA). After 24 h culture, the transfected cells were seeded into 384-well microtiter plates at a density of 3000 cells per well in HBSS supplemented with 5 mM HEPES, 0.1% (w/v) BSA or 0.1% (w/v) casein and 0.5 mM 3-isobutyl-1-methylxanthine. The cells were stimulated with different concentrations of peptide agonists for 40 min at RT. Eu-cAMP tracer and ULightTM-anti-cAMP were then diluted by cAMP detection buffer and added to the plates separately to terminate the reaction. Plates were incubated at RT for 1 h and the fluorescence intensity was measured at 620 nm and 665 nm by an EnVision multilabel plate reader from PerkinElmer (Boston, MA, USA).
Statistical analysis
All functional data were displayed as means ± standard error of the mean (SEM). Statistical analysis was performed using GraphPad Prism 8.0 (GraphPad Software, San Diego, CA, USA). Experimental data were evaluated with a three-parameter logistic equation. The significance was determined with either two-tailed Student’s t-test or one-way ANOVA. P < 0.05 was considered statistically significant.
Results
Structural determination
Stable PTH1R-Gs protein complex can be assembled with LA-PTH, but not with PTH and PTHrP. To overcome this technical obstacle, we employed the NanoBiT tethering strategy to stabilize the PTH1R-Gs protein complex assembly with PTH and PTHrP (Supplementary Fig. S1). The PTH1R was co-expressed with Gαs, Gβ1, Gγ2 and GST-Ric-8B. Nb35 was added into membranes to stabilizing the receptor-G protein complex according to the method previously described [2] (Supplementary Fig. S1).
The complex structures of the Gs-coupled PTH1R bound to PTH and PTHrP were determined by cryo-EM to the resolutions of 2.62 Å and 3.25 Å, respectively (Fig. 1 and Supplementary Figs. S2 and S3, Supplementary Table S1). In addition, we applied extensive particle classifications that yielded a distinct subclass of a dimeric form of the PTH-PTH1R-Gs complexes, which was determined to a resolution of 3.76 Å (Supplementary Fig. S4). Because this dimeric form is mediated through no-physiological Nb35 and Gs interface, no further discussion is presented in the paper. Both endogenous peptide-PTH1R-Gs complexes are similar to other GPCR-G protein complex structures, the α-helical domain (AHD) of Gs is flexible and invisible. Apart from extracellular domain (ECD) of receptor, the majority of the amino acid side chains of receptor and heterotrimeric Gs protein were well resolved in the final models, which are refined against the EM density map with excellent geometry. Both peptides, PTH1–34 and PTHrP1–36 were clearly identified, thus providing reliable models for the mechanistic explanation of pharmacological actions of two endogenous peptide hormones at PTH1R (Supplementary Fig. S5). In addition, like LA-PTH-PTH1R-Gs complex, an ordered annular lipid band wrap** around the periphery of the receptor transmembrane domain (TMD) is visible in the cryo-EM map, among which several lipid molecules were built (Fig. 1b-e). These structural lipids possibly stabilize the PTH1R in its active state.
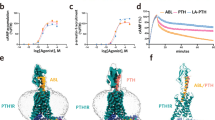
a Schematic diagram of two endogenous hormones, PTH (salmon) and PTHrP (steel blue) are recognized by PTH1R (light sea green) and exert their effects on bone by activating PTH1R, primarily via Gs-cAMP signaling. Gs is colored in medium purple, Gβ1 in light blue and Gγ2 in rosy brown. b Orthogonal views of the density map and c model for PTH-PTH1R-Gs complex. The colored cryo-EM density map is shown at the 0.0183 threshold and light gray surface indicates a micelle diameter of 10 nm. Cadet blue, PTH1R; salmon, PTH; medium purple, Gs; light blue, Gβ1; rosy brown, Gγ2; peru, Nb35; goldenrod, CLR; orchid, PLM. d Orthogonal views of the density map and e model for PTHrP-PTH1R-Gs complex. The colored cryo-EM density map is shown at the 0.136 threshold and light gray surface indicates a micelle diameter of 11 nm. Slate blue, PTH1R; steel blue, PTHrP; medium purple, Gs; light blue, Gβ1; rosy brown, Gγ2; peru, Nb35; goldenrod, CLR; orchid, PLM.
Overall structures of Gs-coupled PTH1R bound to PTH and PTHrP
Both PTH-PTH1R-Gs and PTHrP-PTH1R-Gs complex structures closely resemble that of the LA-PTH-PTH1R-Gs complex with the root mean square deviation (RMSD) value of 0.5 Å for whole complexes, 0.7 Å for Gs alone, 0.5 Å for receptor alone (Fig. 2). Notable conformational differences were observed in the position and orientation of the C-terminal fragment of peptide and the ECD of receptor. The C-terminal fragments of PTH and PTHrP in complexes are shifted away TM7 by ~9.6 Å and 2.6 Å (measured at Cα of H32P), respectively, relative to the same position of LA-PTH in the LA-PTH-bound structure (PDB: 6NBF) [2]. The receptor ECD-α1 helixes in the PTH- and PTHrP-bound structures are shifted away TM7 by ~10 Å and 7.1 Å (measured at Cα of K50ECD), respectively, relative to the same position of ECD-α1 helix in the LA-PTH-bound structure (Fig. 2a–c). These conformational changes allow the C-terminal fragment of hormone bind specificity to PTH1R, indicating PTH1R-associated peptide recognition specificity. Except ECD-α1 helix in PTH-PTH1R cryo-EM structure is shifted away PTH by ~2 Å, relative to X-ray structure (measured at Cα of K50ECD), the overall conformations of the PTH-receptor ECD and PTHrP-receptor ECD in PTH-PTH1R cryo-EM structures resemble the previously published crystal structures of PTH-ECD and PTHrP-ECD complexes (PDB: 3C4M and 3H3G) [11, 37], respectively. The structural basis for the binding specificity of PTH and PTHrP to PTH1R is relatively consistent with the crystal structures (Fig. 2d).
a–c Structural comparison of PTH-PTH1R-Gs (salmon, PTH; cadet blue, PTH1R; medium purple, Gs; light blue, Gβ1; rosy brown, Gγ2), PTHrP-PTH1R-Gs (steel blue, PTHrP; slate blue, PTH1R; purple, Gs; dark khaki, Gβ1; yellow green, Gγ2) and LA-PTH-PTH1R-Gs (PDB: 6NBF, sandy brown, LA-PTH; medium violet red, PTH1R; dark slate blue, Gs; goldenrod, Gβ1; dark green, Gγ2) complexes. Notable conformational changes observed in the receptor ECD and the ligand (a, b) in the side view and (c) cytoplasmic and extracellular view. d Comparison of the PTH-bound ECD in PTH-PTH1R-Gs EM structure with PTH-bound ECD crystal structure (PDB: 3C4M) and PTHrP-bound ECD in PTHrP-PTH1R-Gs EM structure with PTHrP-bound ECD crystal structure (PDB: 3H3G), yellow, PTHrP; lime green, ECD; sea green, PTH; pale violet red, ECD.
Molecular basis for recognition of PTH and PTHrP by PTH1R
Both PTH-PTH1R-Gs and PTHrP-PTH1R-Gs complexes exhibit a similar and unique peptide-receptor binding interface, which is consistent with the “two-domain” model of the peptide-PTH1R interaction mechanism of hormone recognition common to class B GPCRs (Fig. 3). In these complexes, PTH and PTHrP adopt an amphipathic helix with their N terminus dip** into the receptor TMD, while the C terminus closely interacted with the ECD. The N-terminal fragment of PTH (residues 1–14, PTHN) and PTHrP (residues 1–15, PTHrPN) bind to the TMD and activate the receptor. The PTHN and PTHrPN adopt four helical turns and insert deeply into the TMD pocket. PTHN and PTHrPN are directly surrounded by TM1, TM2, TM3, TM5, TM6 and TM7 and form extensive interactions with the TMD core. PTHN and PTHrPN can also interact with ECL2 and ECL3 of receptor. The C-terminal fragment of PTH (residues 15–34, PTHC) and PTHrP (residues 16–34 PTHrPC) bind to the ECD of PTH1R to confer high affinity and specificity to the receptor (Fig. 3a, d).
a An overall PTH-binding pocket of PTH1R. Cutaway view showing that PTHN penetrates into a pocket formed by TMD, while its C-terminal half is recognized by the ECD. b, c Detailed interactions of PTHC with the PTH1R ECD and interactions of PTHN with the PTH1R TMD pocket, salmon, PTH; cadet blue, PTH1R. The polar contacts are shown as purple dotted lines. d An overall PTHrP-binding pocket of PTH1R. Cutaway view showing that PTHrPN penetrates into a pocket formed by TMD, while its C-terminal half is recognized by the ECD. e, f Detailed interactions of PTHrPC with the PTH1R ECD and interactions of PTHrPN with the PTH1R TMD pocket, steel blue, PTHrP; slate blue, PTH1R. The polar contacts are shown as purple dotted lines. g–j Effect of receptor mutation on PTH and PTHrP-induced cAMP accumulation. g, h Signaling profiles of PTH1R ECD mutants on PTH and PTHrP-induced cAMP accumulation. i, j Signaling profiles of PTH1R TMD, ECL2 and ECL3 mutants on PTH and PTHrP-induced cAMP accumulation. cAMP accumulation was measured in wild-type (WT) and single-point mutated PTH1R expressing in HEK293T cells, respectively. cAMP accumulation was normalized to the maximum response of the WT and dose-response curves were analyzed using a three-parameter logistic equation. Data were generated and graphed as means ± SEM of three independent experiments (n = 3) performed in triplicate.
In line with a similar conformation observed between the cryo-EM structures and crystal structures of the PTH1R ECD bound PTH and PTHrP, the extensive interactions of the ECD with PTHC and PTHrPC in cryo-EM structure complexes are similar to those observed in the previously reported X-ray structures of peptide-ECD complexes (Fig. 3b, e). In the PTH-PTH1R-Gs complex, the hydrophobic residues V21, W23, L24, L28, V31 and F34 of the PTH, form an extensive hydrophobic interactions with the hydrophobic groove of the three-layer PTH1R-ECD fold (Fig. 3b) [37]. In the PTHrP-PTH1R-Gs complex, the conserved ECD fold also shows the hydrophobic contacted with F23, L24, L27, I28, I31 and I36 of PTHrP (Fig. 3e) [11]. In addition, N16PTH and Q16PTHrP form a hydrogen bond with E35ECD and T33ECD, respectively. The conserved amino acid R20 in the PTH and PTH-related ligands also forms a hydrogen bond with the backbone carbonyls of M32ECD of PTH1R and forms electrostatic interactions with D137ECD. This amino acid R20 is conserved in all three endogenous peptide hormones of PTH1R and PTH2R but distinct from other class B GPCRs, suggesting that R20 is critical for peptide hormones recognition and specificity of PTH and PTH-related ligands [37]. Except for these conserved interaction, these two peptide hormones also interact with PTH1R in a peptide-specific mode. E19PTH forms electrostatic interaction with K34ECD, V21PTH makes hydrophobic contact with D137ECD, and W23PTH makes Pi interaction with K34ECD and hydrophobic interaction with Q37ECD and I38ECD. But different amino acids at cognate positions of PTHrP, R19PTHrP form electrostatic interaction with E35ECD and R21PTHrP also forms electrostatic interaction with D137ECD. F23PTHrP makes hydrophobic interaction with I38ECD and L41ECD of PTH1R (Fig. 3b, e). These interactions receive support from our mutagenesis studies by measuring cAMP responses. Alanine substitution of D137ECD and Y167ECD of PTH1R shows clearly a great reduction in the potency of both PTH and PTHrP-mediated Gs activation (Fig. 3g, h, Supplementary Table S2). In addition, both alanine substitution M32ECD and E35ECD also diminished the potency of PTHrP and slightly decreased the efficacy of PTH. These amino acid residues are important for determining the affinity and specificity of PTH and PTHrP to PTH1R.
The sequences of PTHN, PTHrPN and other PTH-related ligands highly resemble. Although PTHrPN overlapped well with LA-PTHN, while PTHN is different from their positions. Every one of the three N-terminal peptide helix inserts into the receptor TMD core by a analogical angle and orientation, thereby revealing a similar peptide hormones recognition and activation mechanism (Fig. 3c, f and Supplementary Fig. S6). Both PTHN and PTHrPN run parallel to TM2, making extensive contacts with the residues in TM1, TM2, TM3, TM5, TM6, TM7 and ECL2 and ECL3 (Fig. 3c, f). Especially the first six residues of PTHN and PTHrPN play critical roles in the receptor activation. In contrast, PTH (7–34) and PTHrP (7–34) are potent antagonists of PTH receptor [1]. For PTH, S1 makes more polar interactions with F4246.56b, M4256.57b, T4276.59b and Q4407.38b. For PTHrP, A1 forms a hydrogen bond with T4276.59b and makes hydrophobic contacts with Q3645.40b, M4256.57b and Q4407.38b. These observations provide support for our mutagenesis studies by measuring cAMP responses. Alanine substitutions of Q3645.40b, M4256.57b and Q4407.38b notably reduce the potency PTHrP-mediated Gs activation, and have slight effect on the potency of PTH in Gs activation (Fig. 3i-j, Supplementary Table S2). The same amino acid V2 in two peptides makes extensive hydrophobic contacts with L2923.40b, Q3645.40b and I3675.43b. Mutation of L2923.40b remarkably reduces pEC50 and Emax values in the PTH- and PTHrP-induced receptor activity (Fig. 3i-j, Supplementary Table S2). Both S3PTH and S3PTHrP form contacts with E4447.42b and M4457.43b. S3PTH makes additional interaction with N4487.46b, while S3PTHrP makes additional hydrophobic interaction with M4417.39b. E4447.42bA mutant slightly affects the potency of both PTH and PTHrP in Gs activation, while alanine substitutions of M4457.43b indicate a great reduction in the potency of PTHrP-mediated Gs activation, but almost no effect on the receptor activity by PTH in Gs activation (Fig. 3i–j, Supplementary Table S2). E4 in both PTH and PTHrP, forms the stabilizing hydrogen bond interact and salt bridge with Y1951.47b and R2332.60b, respectively. Both alanine mutations in Y1951.47b and R2332.60b remarkably reduce peptide-mediated PTH1R activation, indicating that E4 of the peptide has a key role for peptide-induced receptor activation (Fig. 3i–j, Supplementary Table S2). I5PTH makes hydrophobic contacts with L2893.37b, L2923.40b and Q3645.40b. While H5PTHrP at the same position of PTHrP not only makes extensive hydrophobic interactions with L2893.37b, K3605.36b and I3635.39b, but also forms an additional electrostatic interaction with Q3645.40b. Q6 of PTH and PTHrP forms highly conserved hydrophobic interactions with Y429ECL3, W4377.35b, Q4407.38b and M4417.39b. Alanine substitutions of Y429ECL3, W4377.35b, Q4407.38b and M4417.39b all decrease activities of both PTH and PTHrP. In addition, N10PTH is hydrogen-bonded with W4377.35b, the same to PTH position, D10PTHrP also forms a hydrogen bond with W4377.35b. W4377.35bA markedly reduces potencies of both PTH and PTHrP in stimulating Gs activation, supporting the fact that N10PTH and D10PTHrP of the peptide are critical for two endogenous hormones-induced receptor activation and signaling (Fig. 3i-j, Supplementary Table S2). H9 in both PTH and PTHrP interacts with ECL2 and ECL3 of receptor, such as the hydrophobic interactions with D353ECL2, S355ECL2, and electrostatic interaction with Y429ECL2. D353ECL2 significantly decreases the PTHrP-induced PTH1R activation (Fig. 3j, Supplementary Table S2), but almost has no effect on the receptor activity by PTH in Gs activation (Fig. 3i, Supplementary Table S2). In addition, L7PTH/PTHrP, M8PTH or L8PTHrP, L11PTH or K11PTHrP, G12PTH/PTHrP, K13PTH/PTHrP, H14PTH or S14PTHrP can form extensive hydrophobic interactions with the TMD core and ECL2. The above interactions of peptideN with the TMD provide the major binding and activation energy for the formation of the signal complex. Therefore, the stability of the interface between peptideN and the TMD is well supported by an extensive network of complementary polar and non-polar interactions between the bound PTH and PTH-related ligands and the TMD. The detailed interactions are listed in Fig. 3c, f and Supplementary Fig. S6. These interactions help to explain the rich and distinct pharmacology of PTH and PTHrP acting through the same receptor.
Activation mechanism of PTH1R
Comparison of the PTH- and PTHrP-bound PTH1R-Gs complex structures with LA-PTH-bound PTH1R-Gs complex structure reveals a remarkably similarity in the G protein binding interface, indicating a common mechanism for G protein-coupling specificity (Fig. 4). Gαs-α5 helix dipped into the cytoplasmic receptor cavity and forms important interactions with TM2, TM3, TM5, TM6, H8 and intracellular loop 3 (ICL3) (Fig. 4c, d). In all three PTH1R-Gs complex structures, L393 in Gαs-α5 helix makes polar interaction with S4096.41b. Both D381 and Q384 of Gαs-α5 helix form an electrostatic interaction with K3885.64b. R385 of Gαs-α5 also makes an additional electrostatic interaction with E391ICL3 of PTH1R in PTH-bound PTH1R-Gs complex and PTHrP-bound PTH1R-Gs complex. E392 forms polar interaction with the backbone amine of N3638.47b and G3648.48b. The bulky residue Y391 at the Gαs-α5 helix C terminus binds to a sub-pocket formed by R2192.46b, H2332.50b, Y3053.53b and L3063.54b in PTH1R. The hydrophobic residues L388, Y391, L393 and L394 of Gαs-α5 helix form extensive hydrophobic interaction with TMD helixes (Fig. 4c, d). F314ICL2 in PTH- PTHrP-bound PTH1R-Gs complexes inserted into a cavity formed by αN-β1 junction and α5 of Gαs and made hydrophobic interactions for stabilizing the interface, but they are different from LA-PTH-bound PTH1R-Gs complex, which F314ICL2 did not directly interact with the hinge region of the Gαs protein (Fig. 4e). In both PTH- and PTHrP-bound PTH1R-Gs complexes, the electrostatic interactions presented between R214ICL1 and D312 of Gβ1, and in PTH- and LA-PTH-bound PTH1R-Gs complex, hydrogen bonds between R4768.60b and the backbone oxygen of G310 of Gβ1, while in PTHrP-bound PTH1R-Gs complex, hydrogen bonds between R4728.56b and the backbone oxygen of G310 of Gβ1. Additional contacts between R213ICL1 and R52 of Gβ1 in PTH- bound PTH1R-Gs complex and in LA-PTH-bound PTH1R-Gs complex, all of these interactions may stabilize the interface of helix 8 and Gβ1 (Fig. 4f).
a Comparison of G protein coupling among PTH-, PTHrP- and LA-PTH- bound PTH1R complexes. The receptors and G proteins are colored as the labels. b The Gαs-α5 helix of the Gαs inserts into an intracellular cavity of receptor’s TMD. Comparison of the PTH1R-Gs interface, including (c) the Gαs-α5 C terminus-TMD interface, (d) Gαs-α5-TM3/TM5 interface, (e) ICL2-Gαs-αN and Gαs-α5 helix interface, (f) and helix 8-Gβ1 interface. Cadet blue, PTH1R; medium purple, Gs; light blue, Gβ1; rosy brown, Gγ2 in PTH-PTH1R-Gs complex, slate blue, PTH1R; purple, Gs; dark khaki, Gβ1; yellow green, Gγ2 in PTHrP-PTH1R-Gs complex and medium violet red, PTH1R; dark slate blue, Gs; goldenrod, Gβ1; dark green, Gγ2 in LA-PTH-PTH1R-Gs complex The polar contacts are shown as purple dotted lines.
Conformational changes during PTH1R activation
PTH1R has two different conformation states, the G protein free state R0 and the G protein-coupled state RG [12]. During peptide-bound PTH1R activation, two PTH1R conformations undergo different rearrangement of the network residues to facilitate the ligand binding and signal transduction [38]. Based on structural alignment of PTH1R: agonist complex in the absence of G protein, with three G protein-bound PTH1R (RG): agonist complexes show different conformational changes, involving the ECD and the TMD of the receptor (Fig. 5a–c). The receptor ECD-α1 in the PTH-bound and PTHrP-bound PTH1R RG structures are shifted away TM7 by ~4.4 Å and 3.5 Å (measured at Cα of K50ECD), respectively, relative to their positions in the ePTH-bound PTH1R structure. The main conformational changes happen in TMD (Fig. 5b, c). The class B GPCRs conserved PxxG motif (P4156.48b, L4166.48b, F4176.49b and G4186.50b) in three agonist-G protein-bound PTH1R complexes caused a notable conformational rearrangement and made TM6 kink outward ~90° in the middle (Fig. 5d). The TM6 sharp kink creating an intracellular binding pocket for the C terminus of Gα, thereby enabling intracellular G protein activation and signal transduction. Meanwhile, the kink of TM6 together with other conformational changes caused three classes B GPCRs conserved polar interaction networks rearrangement, including HETY network (H2232.50b, E3023.50b, T4106.42b, and Y4577.57b), the II-VI-VII-VIII network (R2192.46b, K4056.37b, N4637.61b and E4658.49b) and central polar network (R2332.60b, N2953.43b, H4206.52b and Q4517.49b) (Fig. 5e–g). However, this structural hallmark of class B GPCRs activation was not observed in the ePTH-PTH1R complex structure and CGRP-CGRPR-RAMP1 complex structure, which are in the absence of G protein. Their intracellular part of the receptors still displays the hallmarks of an inactive conformation (Fig. 5b and Supplementary Fig. S7), which are called intermediate states on the transition of the receptor toward the G protein-bound state [38,39,40]. The intermediate states of class B GPCRs and the R0 conformation of the PTH1R resemble in agonist binding and in the absence of G protein. The ability of PTH and PTHrP to bind to RG is similar, but PTH has a greater capacity to bind to R0 state [1, 12]. Such conformational selectivity mediates their distinct biological actions [12]. From molecular dynamics (MD) simulations, the lower R0 binding ability of PTHrP was attributed to its higher binding free energy upon specific states (Supplementary Fig. S8). Although the capacities of PTH and PTHrP to bind to R0 are different, these peptides can bind to the G protein-coupled PTH1R complex (RG) with a highly similar G protein binding interface, suggesting a common mechanism for Gs engagement.
a–c The structural alignment of ePTH-bound PTH1R (PDB: 6FJ3, lime, ePTH; dark goldenrod, PTH1R) with PTH-bound PTH1R (salmon, PTH; cadet blue, PTH1R), PTHrP-bound PTH1R (steel blue, PTHrP; slate blue, PTH1R) and LA-PTH-bound PTH1R (sandy brown, LA-PTH; medium violet red, PTH1R), which are in Gs bound state, showing different conformational changes of the receptor between PTH1R in Gs bound and no Gs bound states (a) in the side view, (b) extracellular view and (c) intracellular view, the red arrows indicate conformational changes. d The structural alignment of PTH1R in the absence of G protein and in G protein-bound state, showing the outward bending of the intracellular portion of TM6 of activated PTH1R in Gs-bound state, which results in a kink at the PxxG motif in TM6 and a ~90° angle in the middle of TM6 of the activated PTH1R in G protein-bound state. The TM6 kink in the active PTH1R structure is indicated by a black line. The conserved PxxG motifs in TM6 are shown in stick representation. e Conformational changes during receptor activation in the absence of G protein and in G protein-bound state in HETY network, f in the II-VI-VII-VIII network, g and in central polar network. The residues are shown in stick representation and conformational changes are indicated by red arrows.
Discussion
The cryo-EM structures of PTH-PTH1R-Gs and PTHrP-PTH1R-Gs complexes further reveal the structural basis for recognition of PTH and PTHrP by PTH1R and the receptor activation and G protein-coupling mechanisms, providing an opportunity to use these structures as modules for designing new PTH replacement drugs in the future. Two synthetic analogs of PTH and PTHrP, teriparatide and abaloparatide were approved drugs by FDA and used in clinic, but as conventional therapy for osteoporosis, peptide drugs with daily injections and high costs, are probably not suitable for treating this widespread disease. Therefore, PTH1R-specific targeted orally available agonist drugs for osteoporosis are urgently needed to be developed. For many years, pursuing to discover such orally available agonists has been hampered owing to lacking of understanding the basic molecular mechanism and modes of action endogenous hormones. In this work, to understand the molecular mechanisms of PTH1R-ligand recognition and activation, we compared two endogenous hormones respectively bound PTH1R-Gs structures with LA-PTH-bound PTH1R-Gs structure and ePTH-bound PTH1R structure in the absence of G protein. All structures uncover the distinct peptide-receptor interactions underlying ligand selectivity, the conformational changes in receptor activation, and key hallmarks of peptide-bound, activated PTH1R, also provide precise templates for PTH1R drug discovery.
To our knowledge, PTH and PTHrP differ in selectivity for two pharmacologically distinguishable high-affinity PTH1R conformations, R0 and RG [1]. The ability of PTH and PTHrP to bind to RG is similar, but PTH has a greater capacity to bind to R0 state. Such R0-selective analogs, including PTH, M-PTH1–28 and M-PTH1–34 display high affinity binding to R0 as well as notably mediate prolonged cAMP signaling responses and are now being investigated as potential new drugs for treating hypoparathyroidism [41]. While another PTH analogs, such as PTHrP1–36 and M-PTH1–14 exhibit a high- affinity to bind to RG conformation, and produces a shorter-lived and more pulsatile action at the receptor, might prove to be more efficacious in this class of patients [1, 14]. Therefore, peptides with different conformational selectivity mediate their distinct biological actions. The structures of PTH-PTH1R-Gs and PTHrP-PTH1R-Gs complexes, combined with mutational studies and MD analyses of the ligand-receptor binding sites indicate the distinctive molecular mechanisms of ligand recognition and receptor activation. The detailed patterns of two endogenous hormones-receptor interactions may aid in the design of new drugs with high affinity to the R0 or RG state for osteoporosis and hypoparathyroidism.
While this manuscript was being completed, Kobayashi et al., Zhai et al. and Cary et al. reported cryo-EM structures of PTH1R bound to PTH and PTHrP [16], PTH and ABL [42], and PTH, PTHrP, ABL, M-PTH and LA-PTH respectively [43]. Our structures of PTH1R bound to PTH and PTHrP agree with those reported by the other three groups, these works provide molecular basis of hormone recognition and PTH1R activation and signaling.
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank under accession codes: EMD-34585 (PTH-bound PTH1R-Gs complex), EMD-34587 (PTHrP-bound PTH1R-Gs complex) and EMD-34598 (dimer of PTH-bound PTH1R-Gs complex). The atomic coordinates have been deposited in the Protein Data Bank under accession codes: 8HA0 (PTH-bound PTH1R-Gs complex), 8HAF (PTHrP-bound PTH1R-Gs complex) and 8HAO (dimer of PTH-bound PTH1R-Gs complex).
References
Gardella TJ, Vilardaga JP. International Union of Basic and Clinical Pharmacology. XCIII. The parathyroid hormone receptors–family B G protein-coupled receptors. Pharmacol Rev. 2015;67:310–37.
Zhao LH, Ma S, Sutkeviciute I, Shen DD, Zhou XE, de Waal PW, et al. Structure and dynamics of the active human parathyroid hormone receptor-1. Science. 2019;364:148–53.
Nishimura Y, Esaki T, Isshiki Y, Furuta Y, Mizutani A, Kotake T, et al. Lead optimization and avoidance of reactive metabolite leading to PCO371, a potent, selective, and orally available human parathyroid hormone receptor 1 (hPTHR1) agonist. J Med Chem. 2020;63:5089–99.
Kir S, White JP, Kleiner S, Kazak L, Cohen P, Baracos VE, et al. Tumour-derived PTH-related protein triggers adipose tissue browning and cancer cachexia. Nature. 2014;513:100–4.
Wider J, Undyala VVR, Lanske B, Datta NS, Przyklenk K. Parathyroid hormone-related peptide and its analog, abaloparatide, attenuate lethal myocardial ischemia-reperfusion injury. J Clin Med. 2022;11:2273; https://doi.org/10.3390/jcm11092273.
Chew CK, Clarke BL. Abaloparatide: recombinant human PTHrP (1-34) anabolic therapy for osteoporosis. Maturitas. 2017;97:53–60.
Srivastava M, Deal C. Osteoporosis in elderly: prevention and treatment. Clin Geriatr Med. 2002;18:529–55.
Bilezikian JP, Rubin MR, Finkelstein JS. Parathyroid hormone as an anabolic therapy for women and men. J Endocrinol Invest. 2005;28:41–9.
Quattrocchi E, Kourlas H. Teriparatide: a review. Clin Ther. 2004;26:841–54.
Gonnelli S, Caffarelli C. Abaloparatide. Clin Cases Min Bone Metab. 2016;13:106–9.
Pioszak AA, Parker NR, Gardella TJ, Xu HE. Structural basis for parathyroid hormone-related protein binding to the parathyroid hormone receptor and design of conformation-selective peptides. J Biol Chem. 2009;284:28382–91.
Dean T, Vilardaga JP, Potts JT Jr., Gardella TJ. Altered selectivity of parathyroid hormone (PTH) and PTH-related protein (PTHrP) for distinct conformations of the PTH/PTHrP receptor. Mol Endocrinol. 2008;22:156–66.
Hattersley G, Dean T, Corbin BA, Bahar H, Gardella TJ. Binding selectivity of abaloparatide for PTH-Type-1-receptor conformations and effects on downstream signaling. Endocrinology. 2016;157:141–9.
Okazaki M, Ferrandon S, Vilardaga JP, Bouxsein ML, Potts JT Jr., Gardella TJ. Prolonged signaling at the parathyroid hormone receptor by peptide ligands targeted to a specific receptor conformation. Proc Natl Acad Sci USA. 2008;105:16525–30.
Zhao LH, Lin J, Ji SY, Zhou XE, Mao C, Shen DD, et al. Structure insights into selective coupling of G protein subtypes by a class B G protein-coupled receptor. Nat Commun. 2022;13:6670.
Kobayashi K, Kawakami K, Kusakizako T, Miyauchi H, Tomita A, Kobayashi K, et al. Endogenous ligand recognition and structural transition of a human PTH receptor. Mol Cell. 2022;82:3468–83. e5
Maeda S, Koehl A, Matile H, Hu H, Hilger D, Schertler GFX, et al. Development of an antibody fragment that stabilizes GPCR/G-protein complexes. Nat Commun. 2018;9:3712.
Chan P, Gabay M, Wright FA, Kan W, Oner SS, Lanier SM, et al. Purification of heterotrimeric G protein alpha subunits by GST-Ric-8 association: primary characterization of purified G alpha(olf). J Biol Chem. 2011;286:2625–35.
Dixon AS, Schwinn MK, Hall MP, Zimmerman K, Otto P, Lubben TH, et al. NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem Biol. 2016;11:400–8.
Ma S, Shen Q, Zhao LH, Mao C, Zhou XE, Shen DD, et al. Molecular basis for hormone recognition and activation of corticotropin-releasing factor receptors. Mol Cell. 2020;77:669–80 e4.
Zivanov J, Nakane T, Scheres SHW. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. Iucrj. 2020;7:253–67.
Rohou A, Grigorieff N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J Struct Biol. 2015;192:216–21.
Punjani A, Rubinstein JL, Fleet DJ, Brubaker MA. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat Methods. 2017;14:290–6.
Sanchez-Garcia R, Gomez-Blanco J, Cuervo A, Carazo JM, Sorzano COS, Vargas J. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun Biol. 2021;4:874.
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, et al. UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605–12.
Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr. 2004;60:2126–32.
Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr. 2010;66:213–21.
Chen VB, Arendall WB, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D. 2010;66:12–21.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Jo S, Cheng X, Lee J, Kim S, Park S-J, Patel DS, et al. CHARMM-GUI 10 years for biomolecular modeling and simulation. J Comput Chem. 2017;38:1114–24.
Lee J, Hitzenberger M, Rieger M, Kern NR, Zacharias M, Im W. CHARMM-GUI supports the Amber force fields. J Chem Phys. 2020;153:35103.
Tian C, Kasavajhala K, Belfon KAA, Raguette L, Huang H, Migues AN, et al. ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J Chem Theory Comput. 2020;16:528–52.
He X, Man VH, Yang W, Lee TS, Wang J. A fast and high-quality charge model for the next generation general AMBER force field. J Chem Phys. 2020;153:114502.
Roe DR, Cheatham TE III. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J Chem theory Comput. 2013;9:3084–95.
Harrigan MP, Sultan MM, Hernández CX, Husic BE, Eastman P, Schwantes CR, et al. MSMBuilder: statistical models for biomolecular dynamics. Biophys J. 2017;112:10–5.
Miller BR 3rd, McGee TD Jr., Swails JM, Homeyer N, Gohlke H, Roitberg AE. MMPBSA.py: an efficient program for end-state free energy calculations. J Chem Theory Comput. 2012;8:3314–21.
Pioszak AA, Xu HE. Molecular recognition of parathyroid hormone by its G protein-coupled receptor. Proc Natl Acad Sci USA. 2008;105:5034–9.
Ehrenmann J, Schoppe J, Klenk C, Pluckthun A. New views into class B GPCRs from the crystal structure of PTH1R. FEBS J. 2019;286:4852–60.
Ehrenmann J, Schoppe J, Klenk C, Rappas M, Kummer L, Dore AS, et al. High-resolution crystal structure of parathyroid hormone 1 receptor in complex with a peptide agonist. Nat Struct Mol Biol. 2018;25:1086–92.
Josephs TM, Belousoff MJ, Liang YL, Piper SJ, Cao J, Garama DJ, et al. Structure and dynamics of the CGRP receptor in apo and peptide-bound forms. Science. 2021;372:eabf7258. https://doi.org/10.1126/science.abf7258.
Mannstadt M, Clarke BL, Vokes T, Brandi ML, Ranganath L, Fraser WD, et al. Efficacy and safety of recombinant human parathyroid hormone (1-84) in hypoparathyroidism (REPLACE): a double-blind, placebo-controlled, randomised, phase 3 study. Lancet Diabetes Endocrinol. 2013;1:275–83.
Zhai X, Mao C, Shen Q, Zang S, Shen DD, Zhang H, et al. Molecular insights into the distinct signaling duration for the peptide-induced PTH1R activation. Nat Commun. 2022;13:6276.
Cary BP, Gerrard EJ, Belousoff MJ, Fletcher MM, Jiang Y, Russell IC, et al. Molecular insights into peptide agonist engagement with the PTH1 receptor. Biorxiv. 2022. https://doi.org/10.1101/2022.09.04.506565.
Acknowledgements
The cryo-EM data were collected by Wen Hu and Kai Wu at Advanced Center for Electron Microscopy at Shanghai Institute of Materia Medica, Chinese Academy of Sciences. We are grateful to them for collecting the cryo-EM data. This work was supported by National Natural Science Foundation of China (32071203 to LHZ; 82073904 to MWW and 81973373 to DHY), the National Key R&D Program of China (2019YFA0904200), the Young Innovator Association of CAS (2018325 to LHZ) and SA-SIBS Scholarship Program to LHZ and DHY; Ministry of Science and Technology (China) grants (2018YFA0507002 to HEX and 2018YFA0507000 to MWW), the Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 to HEX; 18ZR1447800 to LHZ and 21JC1401600 to DHY), the CAS Strategic Priority Research Program (XDB08020303 to HEX).
Author information
Authors and Affiliations
Contributions
LHZ designed the expression constructs, purified the complexes, prepared the final samples for negative stain and data collection toward the structures, participated in model building and performed structure and function data analysis, prepared figures and wrote the manuscript; LHZ prepared the cryo-EM grids and QNY collected cryo-EM images and performed map calculations, built and refined the structure models; YWX participated in model refinement; XHH conducted MD simulations; ATD, CWC, CZ and YZ performed signaling experiments under the supervision of DHY and MWW; LHZ and HEX conceived the project, wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, Lh., Yuan, Qn., Dai, At. et al. Molecular recognition of two endogenous hormones by the human parathyroid hormone receptor-1. Acta Pharmacol Sin 44, 1227–1237 (2023). https://doi.org/10.1038/s41401-022-01032-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-022-01032-z
- Springer Nature Singapore Pte Ltd.
Keywords
This article is cited by
-
Structural basis of tolvaptan binding to the vasopressin V2 receptor
Acta Pharmacologica Sinica (2024)
-
Mechanisms of ligand recognition and activation of melanin-concentrating hormone receptors
Cell Discovery (2024)
-
Conserved class B GPCR activation by a biased intracellular agonist
Nature (2023)