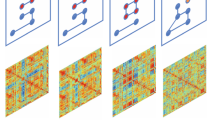
Abstract
Attention deficit hyperactivity disorder (ADHD) is one of the most common psychiatric disorders in school-aged children. Its accurate diagnosis looks after patients’ interests well with effective treatment, which is important to them and their family. Resting-state functional magnetic resonance imaging (rsfMRI) has been widely used to characterize the abnormal brain function by computing the voxel-wise measures and Pearson’s correlation (PC)-based functional connectivity (FC) for ADHD diagnosis. However, exploring the powerful measures of rsfMRI to improve ADHD diagnosis remains a particular challenge. To this end, this paper proposes an automated ADHD classification framework by fusion of multiple measures of rsfMRI in adolescent brain. First, we extract the voxel-wise measures and ROI-wise time series from the brain regions of rsfMRI after preprocessing. Then, to extract the multiple functional connectivities, we compute the PC-derived FCs including the topographical information-based high-order FC (tHOFC) and dynamics-based high-order FC (dHOFC), the sparse representation (SR)-derived FCs including the group SR (GSR), the strength and similarity guided GSR (SSGSR), and sparse low-rank (SLR). Finally, these measures are combined with multiple kernel learning (MKL) model for ADHD classification. The proposed method is applied to the Adolescent Brain and Cognitive Development (ABCD) dataset. The results show that the FCs of dHOFC and SLR perform better than the others. Fusing multiple measures achieves the best classification performance (AUC = 0.740, accuracy = 0.6916), superior to those from the single measure and the previous studies. We have identified the most discriminative FCs and brain regions for ADHD diagnosis, which are consistent with those of published literature.
Similar content being viewed by others
Introduction
Attention deficit hyperactivity disorder (ADHD), typically characterized by the symptoms of inattention, hyperactivity, and impulsivity, has become one of the most common functional disorders in children and adolescents [1]. For example, 8.4% of children from 2 to 17 years of age in the United States were undergoing ADHD, representing 5.4 million children [2]. To date, behavior-based evaluations are the standard clinical approach to diagnosing ADHD, but it is time-consuming and subjective. Moreover, little is known about the association between brain biomarkers (such as brain functional connectivity) and ADHD diagnosis. To tackle this challenge, researchers tend to integrate machine learning (ML) models and brain magnetic resonance imaging (MRI) data for automatical diagnosis and aberrant neuroimaging biomarker identification [3, 4].
Functional magnetic resonance imaging (fMRI), due to its non-invasive and high-resolution properties, has emerged as one of the most frequently used approaches to measuring brain functional connectivities and studying psychiatric diseases [3,4,5,6]. Specifically, fMRI detects the changes of deoxyhemoglobin concentration in local blood vessels of the brain during certain tasks or while remaining still [7]. Accordingly, resting-state fMRI (rsfMRI) can reveal ongoing neural and metabolic activities without an explicit task, facilitating studying of functional regions and networks of the brain and temporal associations among them.
In the past few years, rsfMRI has been extensively utilized to diagnose ADHD and discover the brain’s functional differences between ADHD and healthy controls. Voxel-wise and region of interest (ROI)-wise quantitative features extracted from fMRI can reflect local brain activity and brain region connectivity respectively, therefore they have the potential to serve as biomarkers of ADHD and aid clinical assessment. The voxel-wise measures, such as the regional homogeneity (ReHo), amplitude of low-frequency fluctuations (ALFF), and fractional ALFF (fALFF), are calculated at the voxel level. ReHo [8] measures the similarity or synchronization of the time series in a spatial cluster (usually 27 neighboring voxels), while ALFF [9] focuses on the amplitude of regional activity within the frequency range between 0.01 and 0.1 Hz. FALFF [10] is a normalized ALFF, defined as the original ALFF divided by the total power in the detectable frequency range. On the other hand, in ROI-wise analysis, the whole brain will firstly be parcellated into multiple ROIs according to the brain anatomical or functional atlas [11,12,13,14,15,16,17,18,19]. Then FC matrices that indicate correlative and causal relationships between these predefined ROIs, including the low-order FC (LOFC) and high-order FC (HOFC) networks can be measured. The most popular LOFC construction method is PC analysis, which can capture the pairwise temporal synchronization between two ROIs. However, the limitation of revealing low-order correlation between two brain regions renders the PC-based method notoriously unsuited to capturing high-level relationships among the brain regions. Accordingly, HOFCs, such as tHOFC [20] and dHOFC [21], were proposed to handle this limitation and characterize higher-level brain functional interaction. Besides PC analysis, another category of widely used FC estimation that conducts the partial correlation is the SR method [22], which can characterize the multi-ROI relationship. Moreover, by integrating biological constraints into SR, the generated group SR (GSR) [23], sparse low-rank (SLR) [24], weighted SR (WSR) [25], strength-weighted sparse group representation (WSGR) [25, 26] and strength and similarity guided GSR (SSGSR) [27] are more meaningful for mental disease diagnosis.
In the existing studies on computer-aided ADHD diagnosis, ALFF [9, 28,29,30], ReHo [31,32,33], fALFF [34,35,36], PC-derived FC [37,38,39,40], and SR-derived FC [41, 42] mentioned above were widely adopted as potential predictors. However, the high-order FC and optimized SR, encoding more biologically meaningful clues, were rarely utilized in the automatic diagnosis/classification of ADHD. These measures have shown the potential to improve disease diagnosis performance [43,44,45]. In addition, current studies with high accuracy usually were demonstrated on samples of small size and mostly from a single site [32, 33, 38]. Classification accuracy decreases with the increase of sample size, especially for multi-site heterogeneous datasets [3, 4]. Notably, good performance on small and homogeneous samples does not assure generalizability. Accordingly, multi-site datasets like ADHD-200 [46] and ABCD [47] enable the assessment of the established models’ generalizability on the unseen samples.
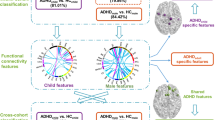
Our study’s experiments have been demonstrated on the multi-site ABCD dataset, including subjects in the 9–10 age range, a narrow one. We extracted ten voxel- and ROI-wise quantitative measures from rsfMRI, and conducted classification using nine basic classifiers combined with Boruta (a Random Forest-based feature extraction method) [48] to diagnose children with ADHD automatically. Nested cross-validation (CV) (10-fold with 5-fold nested) was applied to evaluate the classification of the models. Finally, the model-agnostic and model-based multi-modal fusion approaches were adopted to improve the classification performance by combining the significantly discriminative features. The aims of this study are: (1) to compare the classification performance of different classifiers on different features and (2) to identify biomarkers (brain regions and pairwise connectivity) containing discriminative power of the classification of ADHD. We propose that classification models on HOFCs and optimized SR will achieve better classification performance than traditional measures, and the identified neuroimaging biomarkers will provide new insights into the underlying initial pathogenesis, potentially make ADHD diagnosis and treatment as early as possible. Figure 1 shows the framework of the whole study design. Table 1 shows the abbreviations used in this manuscript.
Materials and methods
Participants
The data in this study were acquired by the ABCD Research Consortium, which recruited 11,875 children aged between 9 and 10 from 21 research sites across the United States. The project was designed to track their biological and behavioral development from adolescence to early adulthood [47]. Therefore, the subjects would have been followed up for at least ten years. Standardized and harmonized assessments have been established of physical and mental health, neurocognition, substance use, culture, environment, multi-modal structural and functional brain imaging, and bioassay protocols. These assessments would be conducted biennially (imaging and bioassays) or annually (non-imaging) [49]. We used the minimally processed baseline neuroimages and the tabulated brain features processed officially from ABCD Fix Release 2.0.1. ADHD patients were diagnosed according to the ABCD Parent Diagnostic Interview for DSM-5 Full (KSADS-5) of the baseline year [50]. Children with the current ADHD diagnoses were labeled as cases and those without any mental disease diagnosis or any ADHD symptom were as healthy controls. The inclusion criteria were as follows: (1) meeting the recommended MRI inclusion criteria according to the ABCD Fix Note 2.0.1; (2) neither hydrocephalus nor herniation; (3) right-handedness; (4) with improbable or possible mild Traumatic brain injury (TBI); (5) with no missing values in the covariables (sex, age, manufacturer, site information, etc.). According to the officially released issue, we excluded the participants scanned on Phillips devices due to incorrect post-processing. Finally, we preprocessed T1w and fMRI data (detailed in 2.2) and removed the participants without passing QC for normalization and head motion (<0.2 mm). The demographic information of the remaining (775 participants) is listed in Table 2. Supplemental Table 1 shows the number of participants remaining after each exclusion step.
fMRI acquisition and preprocessing
The T1w and functional MRI data were collected on the scanners of two vendors (Siemens and GE scanners), across which a harmonized MRI acquisition protocol with comparable acquisition parameters was established. The T1w images were acquired with the following parameters: matrix of 256 × 256, 176 slices (Siemens) and 208 slices (GE), field of view (FOV) size = 256 × 256, voxel size = 1.0 mm3, repetition time (TR) = 2500 ms, echo time (TE) = 2.88 ms (Siemens) and 2 ms (GE), inversion time (TI) = 1060 ms, flip angle (FA) = 8°, and acquisition time 7’12” (Siemens) and 6’09” (GE). The fMRI data were acquired with: matrix of 90 × 90, 60 slices, FOV size = 216 × 216, voxel size = 2.4 × 2.4 × 2.4 mm3, TR = 800 ms, TE = 30 ms, and FA = 52°. All MRI data have been run through standard modality-specific preprocessing stages, including raw file compression, distortion correction, movement correction, alignment to standard space, initial quality control, etc. More details about the MRI acquisition, scanning parameters, and preprocessing pipelines are reported in prior work [51].
Extraction of fMRI measures
The minimally preprocessed fMRI data were further processed using the Data Processing Assistant for Resting-State fMRI (DPARSF v5.1) software (http://rfmri.org/DPARSF), which is based on the toolbox for Data Processing & Analysis of Brain Imaging (DPABI, http://rfmri.org/DPABI) and Statistical Parametric Map** (SPM, http://www.fil.ion.ucl.ac.uk/spm) [52]. The first ten frames were removed to make data reach equilibrium. The time series of images for each subject were realigned and averaged to a mean volume. Then individual structural images (T1w) were co-registered to the mean fMRI and segmented into gray matter (GM), white matter (WM), and cerebrospinal fluid (CSF). After resampling to 3 × 3 × 3 mm3 of voxel size, fMRI images were transformed from individual native space to the Montreal Neurological Institute (MNI) space. Nuisance signals were removed by a linear regression model with head motion parameters, linear trends, and WM and CSF signals included as regressors. Finally, images were spatially smoothed with a 4 mm full width half maximum (FWHM) Gaussian kernel (except for ReHo) and temporally filtered to preserve the signals of 0.01–0.1 Hz (except for fALFF) and remove the high-frequency physiological noise.
We calculated the traditional voxel-wise ALFF, fALFF, and ReHo with DPARSF v5.1. For ROI-wise measures, we first extracted the average time series within each ROI based on the automated anatomical labeling (AAL) template [12] (detailed in Supplementary Table 3). Then the PC-derived LOFC and HOFC (tHOFC and dHOFC), as well as SR-derived FC (SR, GSR, SLR, and SSGSR), were measured with BrainNetClass toolbox v1.1 [45]. After that, the effects of main covariates, including sex, manufacturer, site, and head motion, were regressed, and the residuals were used as inputs for the classification models.
As for PC-derived HOFC, tHOFC can be calculated using the following equation [21, 45]:
where wik is the PC between ith and kth ROI signals, and \(i,j,k \in \{ 1,2, \ldots ,N\} ,k\, \ne\, i,j\), N is the number of ROIs (in our case, N = 116). Note that tHOFC indicates the similarity of the LOFC topographical profiles by measuring the correlation of correlation rather than that of the time series, which reflects the high-level property of the brain network and makes providing supplementary information to the traditional LOFC promising. The calculation of dHOFC is defined as [21, 45]:
where Θ is the number of sliding windows which depends on whole time length (T), step size (s) and window length (L) (\({{{\mathrm{{\Theta}}}}} = \left\lfloor {(T - L)/s} \right\rfloor + 1\), in our case, T = 370, s = 1), wij (θ) means PC between ith and jth ROIs in the θth sliding window. By calculating the correlation between two PC time series, dHOFCij,pq shows how the PC between the ith and the ith ROIs influence the PC between the pth and the qth ROIs. In this way, the total number of elements of dHOFC is proportional to N2 × N2, much larger than that of PC and tHOFC (N × N). Ward’s linkage clustering [53], a widely used hierarchical clustering algorithm, can be applied to group the resulting FCs into K clusters to reduce the dimension of the large-scale network. Each cluster contains several FCs showing a similar pattern of variation along the time. Finally, the averaged correlation time series in each cluster can be calculated to construct a K × K HOFC network. Thus, the final low-scale dHOFC characterizes the temporal synchronization of dynamic FC time series.
As for the SR method, a sparse FC matrix can be generated by minimizing the loss function:
where X is the original fMRI data matrix, W is the FC matrix, λ > 0 controls network sparsity with an l1-norm penalty. Another SR-based approach named SLR is formulated as the following [24, 45].
where λ1 controls sparsity with an l1-norm and λ2 controls modularity with a trace norm. Thus, SLR resulting FC matrices are sparse and low-rank, i.e., modularized and more biologically meaningful [54]. Formula (5) shows the loss function of GSR [23, 45]:
where \({{{\boldsymbol{x}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}\) is the regional mean time series of the ith ROI for the sth subject, and \({{{\boldsymbol{x}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}\) can be regarded as a linear combination of time series of other ROIs: \({{{\boldsymbol{x}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}} = {{{\mathbf{{{{\mathcal{X}}}}}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}{{{\mathcal{w}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}} + {{{\boldsymbol{e}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}\), \({{{\mathbf{{{{\mathcal{X}}}}}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}} = [{{{\boldsymbol{x}}}}_1^{{{\boldsymbol{s}}}}, \ldots ,{{{\boldsymbol{x}}}}_{{{{\boldsymbol{i}}}} - 1}^{{{\boldsymbol{s}}}},{{{\boldsymbol{x}}}}_{{{{\boldsymbol{i}}}} + 1}^{{{\boldsymbol{s}}}}, \ldots ,{{{\boldsymbol{x}}}}_{{{\boldsymbol{N}}}}^{{{\boldsymbol{s}}}}]\) is data matrix of all time series except for the ith ROI, \({{{\mathcal{w}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}\) is the weight vector quantifying the degree of influence of other ROIs on the ith ROI, thus the dimension of \({{{\mathcal{w}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}\) is (N − 1), \({{{\boldsymbol{e}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}}\) is the error, \({{{\mathbf{{{{\mathcal{W}}}}}}}}_{{{\boldsymbol{i}}}} = \left[ {{{{\mathcal{w}}}}_{{{\boldsymbol{i}}}}^1,{{{\mathcal{w}}}}_{{{\boldsymbol{i}}}}^2, \ldots ,{{{\mathcal{w}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{S}}}}} \right]\), S is the number of subjects. Controlled by an l2,1-penalty, GSR resultant FC matrices have less inter-subject variability than SR. SSGSR, as an optimized GSR, is to minimize the following loss function [27]:
where \({{{\boldsymbol{B}}}}_{{{\boldsymbol{i}}}} = \left[ {{{{\boldsymbol{b}}}}_{{{\boldsymbol{i}}}}^1,{{{\boldsymbol{b}}}}_{{{\boldsymbol{i}}}}^2, \ldots ,{{{\boldsymbol{b}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{S}}}}} \right]\), \({{{\boldsymbol{b}}}}_{{{\boldsymbol{i}}}}^{{{\boldsymbol{s}}}} = [b_{i,1}^s, \ldots ,b_{i,i - 1}^s,b_{i,i + 1}^s, \ldots ,b_{i,N}^s]\), and \(b_{i,j} = e^{ - w0_{i,j}^2}\), w0i,j is the LOFC between the ith and the jth ROIs, hence Bi is to penalize the links with weak LOFC. \(l_i^{p,q} = e^{ - \left\| {w0_i^p - w0_i^q} \right\|_2^2}\) defines the similarity between the pth and the qth subjects in terms of their one-to-all LOFC patterns for the ith ROI. Therefore, λ1 and λ2 can control the weighted sparsity and inter-subject variability separately.
The aforementioned HOFCs and SR methods were proposed and applied to various mental diseases, including mild cognitive impairment (MCI) [24, 27, 43] and autism spectrum disorder (ASD) [44], etc. Concerning ADHD, a study [55] built a diagnostic model based on the temporal variability of dynamic functional connectivity, but it did not depend on sliding windows. Another study [56] utilized dynamic functional network connectivity (dFNC) to access FC differences among child, adolescent, and adult ADHD patients rather than between patients and healthy controls. Moreover, the dFNC analysis process is different from dHOFC. To our knowledge, no other study applies tHOFC, dHOFC, and three SR-derived FC (GSR, SSGSR, and SLR) to the classification of ADHD. For the first time, we extracted these features combined with traditional voxel-wise measures and ROI-wise PC and SR to perform a classification task on ADHD and compare the discriminative power of different measures.
ADHD classification
We tested the performance of nine classification algorithms, including four basic classifiers (Logistic Regression (LR), K-Nearest Neighbors (KNN), Ridge Classifier, and Gaussian Naïve Bayes (GaussianNB)), two SVM-based classifiers (Linear SVM and non-linear SVM), and three tree-based ensemble classifiers (Random Forest (RF), Light Gradient Boosting Model (LGBM) and Adaptive Boosting (AdaBoost)). Owing to the high dimension and redundancy of the candidate features, we used Boruta [48] to identify the features with discriminative power on the training set. This RF-based feature selection method has been proven effective in our previous study [40]. Noting that indices including dHOFC and SR-derived FCs are based on parameter-required methods, we tested the parameter sensitivity for these indices and chose the suggested parameters for the subsequent nested CV. Specifically, for each parameter (or parameter combination) 10-fold was evaluated, which created a plot of parameter sensitivity showing the changes in the average AUCs corresponding to various parameters. Then the parameter with the highest average AUC was selected. In each CV loop, a non-linear SVM with the default configuration (kernel = ’rbf’ and C = 1.0) was used as the classifier. The details of parameters associated with different indices are shown in Table 3. Finally, we applied two multi-modal classification methods (model-agnostic early fusion strategy and model-based Multiple Kernel Learning (MKL) algorithm [57, 58]) to fusing the discriminative features from different modalities and improving the classification performance.
Nested cross-validation
We applied a nested CV, including an outer 10-fold CV and an inner 5-fold CV, to evaluating the performance of the classification models (shown in Fig. 2). Compared to single CV, the nested CV will get a more accurate estimate of the models’ generalization performance. In the outer 10-fold CV, we applied feature selection on the training set. Then parameter optimization through grid search was conducted on the inner 5-fold CV within the training set. The final model that leads to the highest AUC was re-trained on the training set and evaluated on the testing set of outer CV. This can lead to a more realistic estimate of the models’ generalization performance, since the models have not been overfitted to the test set. Finally, ten folds metrics, including AUC, ACC, F1-score, precision and recall, were summarized as the performance of the models.
The classification metrics can be calculated as follows:
where TP and TN are the counts of correctly classified ADHD patients and healthy subjects respectively, while FN and FP are counts of falsely classified ADHD patients and healthy subjects respectively. In addition, classification AUC represents the area under the receiver operating characteristic (ROC) curve drawn when the discrimination cutoff varies.
Biomarker identification
We identified fMRI biomarkers with significant discriminative power in classifying ADHD and the control. For ROI-wise measures, the discriminative features always selected in each outer CV fold were firstly regarded as feature candidates. Then, for each candidate, a two-sample t-test with false discovery rate (FDR) correction was performed between the groups of subjects, and the features with a statistically significant difference were selected as the final biomarkers. However, for voxel-wise measures, the voxels selected across all folds were spatially discontinuous. In other words, they were unable to form clusters. Accordingly, we applied a two-sample t-test with Gaussian random field (GRF) correction to the whole subjects using DPABI. Voxel level value were set P < 0.001 and cluster level P < 0.05 (two-tailed). The features passing GRF correction were regarded as the final biomarkers.
Results
Parameter sensitivity test
We first conducted a parameter sensitivity test on the parameter-dependent measures (PC-derived dHOFC, SR-derived SR, GSR, SSGSR, and SLR) to determine the best parameter (or parameter combination) for the subsequent classification. As shown in Fig. 3, classification AUC is sensitive to parameter combinations for dHOFC, while, for SR-derived measures, AUC changes only slightly with a different parameter (or parameter combination). We chose the parameters with the highest average AUC on the 10-fold CV as the optimal, whose details are shown in Table 3.
Classification performance
The classification performance of nine unimodal classifiers and two multi-modal fusion methods based on four measures of ALFF, dHOFC, SSGSR, and SLR is summarized in Table 4 and Fig. 4. In voxel-wise measures, ALFF outperformed the others with the best AUC (0.624) achieved on non-linear SVM and the best ACC (0.5921) on Gaussian Naïve Bayes. Likewise, dHOFC has the best AUC (0.7315) and ACC (0.675) achieved by linear SVM, performing better than the other ROI-wise PC-derived measures. In SR-derived measures, SSGSR and SLR had better performance than the others: SLR achieved the best AUC (0.6616) and ACC (0.6219) by LGBM, SSGSR the best precision (0.7157) by Ridge Classifier. Furthermore, MKL reached the highest AUC (0.7408) and ACC (0.6916) and outperformed all unimodal methods, which can be explained by the combination of complementary neuroimaging information reflected by different measures. The full results based on the ten measures can be found in Supplemental Table 2.
Biomarker identification
We selected the voxels passing the two-sample t-test with GRF correction for voxel-wise ALFF as abnormal voxels, which formed 4 clusters shown in Fig. 5A and Supplemental Table 4. For the ROI-wise measures, we selected biomarkers (dHOFC: pairs of clusters, SSGSR, and SLR: FCs) that were always chosen in each CV fold by Boruta and passed t-test and FDR correction. Then, for dHOFC, we calculated the counts of each cluster that appeared in all pairs and chose the top 10 clusters of FCs as the final biomarkers (shown in Fig. 5D and Supplemental Table 5). For SLR and SSGSR, we sorted FCs according to their p-values and then selected the top-ranked 20 connections of SLR and the whole 15 discriminative connections of SSGSR (shown in Fig. 5B, C and Supplemental Table 6).
A The cluster map shows the results of the two-sample t-test with GRF correction on ALFF. The clusters with the peak activation strength (t-value) signed ‘+’ are colored in orange, and those signed ‘-’ are colored in blue; B, C The circos maps shows the abnormal brain region connectivity of SLR (B) and SSGSR (C). The ROIs of the AAL atlas are denoted on the inner circle, the corresponding brain networks [16] on the outer one. The width and color of links encode the t-value. The wider links correspond to the larger absolute t-values; the warm colors (red and orange) and cool colors (blue and cyan) correspond to the positive and negative t-value, respectively. D The selected top 10 dHOFC clusters of functional connectivity were drawn within the brain surface. The node color represents the functional network to which it belongs. VN visual network; SMN sensory-motor network; DAN dorsal attention network; VAN ventral attention network; LN limbic network; FPN frontoparietal network; DMN default mode network; CN cerebellum network.
Discussion
This study investigated the performances of different machine learning algorithms in classifying ADHD patients vs. healthy controls. To our knowledge, this is the first study that applied PC-derived tHOFC and dHOFC and SR-derived GSR, SSGSR, and SLR to diagnose ADHD based on the ABCD dataset. We also extracted the traditional fMRI-based measures (voxel-wise measures, PC, and SR) and compared their discriminative power. Our results showed that dHOFC achieved the best classification AUC of 0.7315, whose superior ability to identify ADHD patients suggests that the dynamic FC might underlie spontaneous fluctuations in attention [55, 59]. Notably, Wang et al. [55] achieved a higher AUC of 0.84 based on the measure of the temporal variability of dynamic functional connectivity, outperforming other traditional measures. However, the total sample size of 240 in their study is much smaller than ours (775). For ADHD, the reported classification model’s performance deteriorates with sample size [3, 4]. SLR resulted in sparsity but it preserved modularity structure in FC networks [45] and also achieved a high classification AUC of 0.6616 compared with other measures except for dHOFC. These two biologically meaningful measures provide complementary neuroimaging information for traditional indices.
The existing research on ADHD with rsfMRI is mainly based on the ADHD-200 dataset [46], collected from the subjects aged 7–27. However, only two published ADHD neuroimaging studies are based on the ABCD dataset. Owens et al. [60] considered the measures from sMRI and three task fMRI as predictors respectively, and used Elastic Net Regression to predict a continuous ADHD symptomatology coefficient. The best model was achieved on EN-Back features and explained 2.0% of the variance (R2 = 2.0%) in ADHD symptomatology regardless of covariates, while after all covariates are taken into account, R2 drops to 0.6%. Regarding the categorical analyses, they did not robustly predict the ADHD diagnosis from KSADS. Another study [40] comprehensively considered features from three modalities (sMRI, DTI, and resting-state fMRI) and finally reached an AUC of 0.698 using the Multiple Kernel Learning framework. Our study achieved an AUC of 0.7408, considering only resting-state fMRI, superior to the studies above. In addition, our results are better than most studies with similar sample sizes, only inferior to these two [61, 62] (detailed in Table 5).
Our study found the aberrant neuroimaging biomarkers with significant discriminative power in identifying ADHD patients based on fMRI-derived measures. For voxel-based ALFF, the enhanced brain activities lay in the cerebellum and caudate nucleus of ADHD patients, whereas the decreased ones were found in the medial superior frontal gyrus and pre-motor and supplementary motor cortex. These regional brain aberrances are in line with previous findings [32, 33, 35, 63, 64]. The caudate nucleus and supplementary motor cortex are part of the executive control network, and these regions are involved in attention controls [65, 66], which also verifies our findings. For ROI-based dHOFC, SSGSR, and SLR, the number of increased FCs in ADHD was much more than that of the decreased ones. These abnormal FCs of brain regions were within and between cerebellum network (CN), limbic network (LN), ventral attention network (VAN), frontoparietal network (FPN), and default mode network (DMN). The connections from DMN to LN, CN, FPN, and VAN are much stronger in ADHD patients than in healthy controls, and similar tendencies were reported in the previous studies [67, 68]. Moreover, the connections from LN (left thalamus, left parahippocampal gyrus, left/right gyrus rectus, left pallidum, and right middle/superior temporal gyrus) to DMN, CN, and FPN are abnormal in ADHD. Earlier, the thalamus was reported as a mediator in frontostriatal circuitry during attention tasks [71, 72]) that can automatically extract high-level and compact feature representations will be introduced in the future. Another is that we performed binary classification of patients/controls but neglected the ADHD subtypes.
Conclusion
This study proposed an automated ADHD classification framework on a multi-site ABCD dataset containing children aged between 9 and 10. We extracted voxel-wise and ROI-wise quantitative measures from resting-state fMRI and used several machine learning methods to predict the ADHD diagnosis. Classification models on ROI-wise dHOFC and SLR outperformed other measures owing to their more biologically meaningful and complementary neuroimaging information. The highest classification AUC was achieved using MKL by fusing different features. The identified aberrant regions (cerebellum, caudate nucleus, medial superior frontal gyrus, pre-motor and supplementary motor cortex) and FCs (within and between CN, LN, VAN, FPN, and DMN) are widespread in the whole brain and conform with previous literature generally.
Code availability
We used publicly available MATLAB-based tools (DPARSF v5.1 and BrainNetClass toolbox v1.1) to implement the calculation of measures. All codes used to extract measures and generate results that are reported in this paper are available upon request.
References
Wolraich ML, Hagan JF, Allan C, Chan E, Davison D, Earls M, et al. Clinical practice guideline for the diagnosis, evaluation, and treatment of attention-deficit/hyperactivity disorder in children and adolescents. Pediatrics. 2019;144:e20192528.
Danielson ML, Bitsko RH, Ghandour RM, Holbrook JR, Kogan MD, Blumberg SJ. Prevalence of parent-reported ADHD diagnosis and associated treatment among US children and adolescents. 2016 J Clin Child Adolesc. 2018;47:199–212.
Arbabshirani MR, Plis S, Sui J, Calhoun VD. Single subject prediction of brain disorders in neuroimaging: promises and pitfalls. Neuroimage. 2017;145:137–65.
Sakai K, Yamada K. Machine learning studies on major brain diseases: 5-year trends of 2014–2018. Jpn J Radio. 2019;37:34–72.
Cortese S, Aoki YY, Itahashi T, Castellanos FX, Eickhoff SB. Systematic review and meta-analysis: resting-state functional magnetic resonance imaging studies of attention-deficit/hyperactivity disorder. J Am Acad Child Adolesc Psychiatry. 2021;60:61–75.
Gui Y, Zhou X, Wang Z, Zhang Y, Wang Z, Zhou G, et al. Sex-specific genetic association between psychiatric disorders and cognition, behavior and brain imaging in children and adults. Transl Psychiatry. 2022;12:1–8.
Ogawa S, Lee TM, Kay AR, Tank DW. Brain magnetic resonance imaging with contrast dependent on blood oxygenation. Proc Natl Acad Sci USA. 1990;87:9868–72.
Zang YF, Jiang TZ, Lu YL, He Y, Tian LX. Regional homogeneity approach to fMRI data analysis. Neuroimage. 2004;22:394–400.
Zang YF, He Y, Zhu CZ, Cao QJ, Sui MQ, Liang M, et al. Altered baseline brain activity in children with ADHD revealed by resting-state functional MRI. Brain Dev-Jpn. 2007;29:83–91.
Zou QH, Zhu CZ, Yang YH, Zuo XN, Long XY, Cao QJ, et al. An improved approach to detection of amplitude of low-frequency fluctuation (ALFF) for resting-state fMRI: Fractional ALFF. J Neurosci Meth. 2008;172:137–41.
Eickhoff SB, Yeo BTT, Genon S. Imaging-based parcellations of the human brain. Nat Rev Neurosci. 2018;19:672–86.
Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage. 2002;15:273–89.
Desikan RS, Segonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D, et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage. 2006;31:968–80.
Destrieux C, Fischl B, Dale A, Halgren E. Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. Neuroimage. 2010;53:1–15.
Auzias G, Coulon O, Brovelli A. MarsAtlas: a cortical parcellation atlas for functional map**. Hum Brain Mapp. 2016;37:1573–92.
Yeo BTT, Krienen FM, Sepulcre J, Sabuncu MR, Lashkari D, Hollinshead M, et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J Neurophysiol. 2011;106:1125–65.
Glasser MF, Coalson TS, Robinson EC, Hacker CD, Harwell J, Yacoub E, et al. A multi-modal parcellation of human cerebral cortex. Nature. 2016;536:171-+.
Gordon EM, Laumann TO, Adeyemo B, Huckins JF, Kelley WM, Petersen SE. Generation and evaluation of a cortical area parcellation from resting-state correlations. Cereb Cortex. 2016;26:288–303.
Kong R, Li JW, Orban C, Sabuncu MR, Liu HS, Schaefer A, et al. Spatial topography of individual-specific cortical networks predicts human cognition, personality and emotion (vol 29, pg 2533, 2019). Cereb Cortex. 2021;31:3974–3974.
Zhang H, Chen XB, Shi F, Li G, Kim M, Giannakopoulos P, et al. Topographical information-based high-order functional connectivity and its application in abnormality detection for mild cognitive impairment. J Alzheimers Dis. 2016;54:1095–112.
Chen XB, Zhang H, Gao Y, Wee CY, Li G, Shen DG, et al. High-order resting-state functional connectivity network for MCI classification. Hum Brain Mapp. 2016;37:3282–96.
Lee H, Lee DS, Kang H, Kim BN, Chung MK. Sparse brain network recovery under compressed sensing. IEEE T Med Imaging. 2011;30:1154–65.
Wee CY, Yap PT, Zhang D, Wang L, Shen D. Group-constrained sparse fMRI connectivity modeling for mild cognitive impairment identification. Brain Struct Funct. 2014;219:641–56.
Qiao L, Zhang H, Kim M, Teng S, Zhang L, Shen D. Estimating functional brain networks by incorporating a modularity prior. Neuroimage. 2016;141:399–407.
Yu R, Zhang H, An L, Chen X, Wei Z, Shen D. Connectivity strength-weighted sparse group representation-based brain network construction for MCI classification. Hum Brain Mapp. 2017;38:2370–83.
Simon N, Friedman J, Hastie T, Tibshirani R. A sparse-group lasso. J Comput Graph Stat. 2013;22:231–45.
Zhang Y, Zhang H, Chen X, Liu M, Zhu X, Lee SW, et al. Strength and similarity guided group-level brain functional network construction for MCI diagnosis. Pattern Recognit. 2019;88:421–30.
Li F, He N, Li YY, Chen LH, Huang XQ, Lui S, et al. Intrinsic brain abnormalities in attention deficit hyperactivity disorder: a resting-state functional MR imaging study. Radiology. 2014;272:514–23.
Yang H, Wu QZ, Guo LT, Li QQ, Long XY, Huang XQ, et al. Abnormal spontaneous brain activity in medication-naive ADHD children: a resting state fMRI study. Neurosci Lett. 2011;502:89–93.
Sato JR, Hoexter MQ, Fujita A, Rohde LA. Evaluation of pattern recognition and feature extraction methods in ADHD prediction. Front Syst Neurosci. 2012;6:68.
Alonso BD, Tobon SH, Suarez PD, Flores JG, Carrillo BD, Perez EB. A multi-methodological MR resting state network analysis to assess the changes in brain physiology of children with ADHD. PLoS One. 2014;9:e99119.
Zhu CZ, Zang YF, Cao QJ, Yan CG, He Y, Jiang TZ, et al. Fisher discriminative analysis of resting-state brain function for attention-deficit/hyperactivity disorder. Neuroimage. 2008;40:110–20.
Wang XH, Jiao Y, Tang TY, Wang H, Lu ZH. Altered regional homogeneity patterns in adults with attention-deficit hyperactivity disorder. Eur J Radio. 2013;82:1552–7.
Tang C, Wei YQ, Zhao JJ, Nie JX. The dynamic measurements of regional brain activity for resting-state fMRI: d-ALFF, d-fALFF and d-ReHo. Lect Notes Comput Sci. 2018;11072:190–7.
Tan LR, Guo XY, Ren S, Epstein JN, Lu LJ. A computational model for the automatic diagnosis of attention deficit hyperactivity disorder based on functional brain volume. Front Comput Neurosci. 2017;11:75.
Zou L, Zheng JN, Mia CY, Mckeown MJ, Wang ZJ. 3D CNN based automatic diagnosis of attention deficit hyperactivity disorder using functional and structural MRI. IEEE Access. 2017;5:23626–36.
Siqueira AD, Biazoli CE, Comfort WE, Rohde LA, Sato JR. Abnormal functional resting-state networks in ADHD: graph theory and pattern recognition analysis of fMRI data. Biomed Res Int. 2014;2014:380531.
Qureshi MNI, Oh JY, Min B, Jo HJ, Lee B. Multi-modal, multi-measure, and multi-class discrimination of ADHD with hierarchical feature extraction and extreme learning machine using structural and functional brain MRI (vol 11, 157, 2017). Front Hum Neurosci. 2017;11:157.
Riaz A, Asad M, Alonso E, Slabaugh G. Fusion of fMRI and non-imaging data for ADHD classification. Comput Med Imaging Graph. 2018;65:115–28.
Zhou XC, Lin QM, Gui YY, Wang ZX, Liu MH, Lu H. Multimodal MR images-based diagnosis of early adolescent attention-deficit/hyperactivity disorder using multiple kernel learning. Front Neurosci-Switz. 2021;15:710133.
Zhang Y, Tang YB, Chen Y, Zhou L, Wang C. ADHD classification by feature space separation with sparse representation. Int Conf Digit Sig. 2018. https://doi.org/10.1109/ICDSP.2018.8631658.
Strength and similarity guided GSR based network to diagnose ADHD. Proceedings of the 2020 IEEE International Conference on Progress in Informatics and Computing (PIC). IEEE; 2020.
Zhou YY, Qiao LS, Li WK, Zhang LM, Shen DG. Simultaneous estimation of low- and high-order functional connectivity for identifying mild cognitive impairment. Front Neuroinform. 2018;12:3.
Zhou YY, Zhang LM, Teng SH, Qiao LS, Shen DG. Improving sparsity and modularity of high-order functional connectivity networks for MCI and ASD identification. Front Neurosci-Switz. 2018;12:959.
Zhou Z, Chen XB, Zhang Y, Hu D, Qiao LS, Yu RP, et al. A toolbox for brain network construction and classification (BrainNetClass). Hum Brain Mapp. 2020;41:2808–26.
Consortium HD. The ADHD-200 Consortium: a model to advance the translational potential of neuroimaging in clinical neuroscience. Front Syst Neurosci. 2012;6:62.
Jernigan TL, Brown SA, Dowling GJ. The adolescent brain cognitive development study. J Res Adolescence: Off J Soc Res Adolescence. 2018;28:154.
Kursa MB, Rudnicki WR. Feature selection with the Boruta Package. J Stat Softw. 2010;36:1–13.
Alcohol Research: Current Reviews Editorial S. NIH’s Adolescent Brain Cognitive Development (ABCD) Study. Alcohol Res. 2018;39:97.
Barch DM, Albaugh MD, Avenevoli S, Chang L, Clark DB, Glantz MD, et al. Demographic, physical and mental health assessments in the adolescent brain and cognitive development study: rationale and description. Dev Cogn Neurosci. 2018;32:55–66.
Hagler DJ, Hatton S, Cornejo MD, Makowski C, Fair DA, Dick AS, et al. Image processing and analysis methods for the Adolescent Brain Cognitive Development Study. Neuroimage. 2019;202:116091.
Yan CG, Wang XD, Zuo XN, Zang YF. DPABI: Data Processing & Analysis for (Resting-State) Brain Imaging. Neuroinformatics. 2016;14:339–51.
Ward JH Jr. Hierarchical grou** to optimize an objective function. J Am Stat Assoc. 1963;58:236–44.
Bullmore ET, Sporns O. The economy of brain network organization. Nat Rev Neurosci. 2012;13:336–49.
Wang XH, Jiao Y, Li LH. Identifying individuals with attention deficit hyperactivity disorder based on temporal variability of dynamic functional connectivity. Sci Rep. 2018;8:11789.
Agoalikum E, Klugah-Brown B, Yang H, Wang P, Varshney S, Niu BC, et al. Differences in disrupted dynamic functional network connectivity among children, adolescents, and adults with attention deficit/hyperactivity disorder: a resting-state fMRI study. Front Hum Neurosci. 2021;15:697696.
Gonen M, Alpaydin E. Multiple kernel learning algorithms. J Mach Learn Res. 2011;12:2211–68.
Lauriola I, Aiolli F. MKLpy: a python-based framework for Multiple Kernel Learning. Preprint at https://arxiv.org/abs/2007.09982. 2020.
Kucyi A, Hove MJ, Esterman M, Hutchison RM, Valera EM. Dynamic brain network correlates of spontaneous fluctuations in attention. Cereb Cortex. 2017;27:1831–40.
Owens MM, Allgaier N, Hahn S, Yuan D, Albaugh M, Adise S, et al. Multimethod investigation of the neurobiological basis of ADHD symptomatology in children aged 9–10: baseline data from the ABCD study. Transl Psychiatry. 2021;11:64.
Dey S, Rao AR, Shah M. Exploiting the brain’s network structure in identifying ADHD subjects. Front Syst Neurosci. 2012;6:75.
Shao LZ, You Y, Du HP, Fu DM. Classification of ADHD with fMRI data and multi-objective optimization. Comput Meth Prog Biol. 2020;196:105676.
Hart H, Chantiluke K, Cubillo AI, Smith AB, Simmons A, Brammer MJ, et al. Pattern classification of response inhibition in ADHD: toward the development of neurobiological markers for ADHD. Hum Brain Mapp. 2014;35:3083–94.
Soros P, Hoxhaj E, Borel P, Sadohara C, Feige B, Matthies S, et al. Hyperactivity/restlessness is associated with increased functional connectivity in adults with ADHD: a dimensional analysis of resting state fMRI. Bmc Psychiatry. 2019;19:43.
Lanka P, Rangaprakash D, Dretsch MN, Katz JS, Denney TS Jr, Deshpande G. Supervised machine learning for diagnostic classification from large-scale neuroimaging datasets. Brain Imaging Behav. 2020;14:2378–416.
Elton A, Alcauter S, Gao W. Network connectivity abnormality profile supports a categorical-dimensional hybrid model of ADHD. Hum Brain Mapp. 2014;35:4531–43.
Guo X, Yao D, Cao Q, Liu L, Zhao Q, Li H, et al. Shared and distinct resting functional connectivity in children and adults with attention-deficit/hyperactivity disorder. Transl Psychiatry. 2020;10:65.
Rubia K, Criaud M, Wulff M, Alegria A, Brinson H, Barker G, et al. Functional connectivity changes associated with fMRI neurofeedback of right inferior frontal cortex in adolescents with ADHD. Neuroimage. 2019;188:43–58.
**a S, Li X, Kimball AE, Kelly MS, Lesser I, Branch C. Thalamic shape and connectivity abnormalities in children with attention-deficit/hyperactivity disorder. Psychiatry Res. 2012;204:161–7.
Lopez-Larson MP, King JB, Terry J, McGlade EC, Yurgelun-Todd D. Reduced insular volume in attention deficit hyperactivity disorder. Psychiatry Res. 2012;204:32–39.
Kong Y, Genchev GZ, Wang XL, Zhao HY, Lu H. Nuclear segmentation in histopathological images using two-stage stacked U-nets with attention mechanism. Front Bioeng Biotech. 2020;8:573866.
Yao SQ, Yan JC, Wu MY, Yang X, Zhang WT, Lu H, et al. Texture synthesis based thyroid nodule detection from medical ultrasound images: interpreting and suppressing the adversarial effect of in-place manual annotation. Front Bioeng Biotech. 2020;8:599.
Zhan Y, Wei J, Liang J, Xu X, He R, Robbins TW, et al. Diagnostic classification for human autism and obsessive-compulsive disorder based on machine learning from a primate genetic model. Am J Psychiatry. 2021;178:65–76.
Sen B, Borle NC, Greiner R, Brown MRG. A general prediction model for the detection of ADHD and Autism using structural and functional MRI. PLoS One. 2018;13:e0194856.
Ghiassian S, Greiner R, ** P, Brown MR. Using functional or structural magnetic resonance images and personal characteristic data to identify ADHD and autism. PLoS One. 2016;11:e0166934.
Sidhu GS, Asgarian N, Greiner R, Brown MR. Kernel principal component analysis for dimensionality reduction in fMRI-based diagnosis of ADHD. Front Syst Neurosci. 2012;6:74.
Acknowledgements
This work is partly supported by Innovative Research Team of High-Level Local Universities in Shanghai (SHSMUZDCX20212200), the Neil Shen’s SJTU Medical Research Fund, SJTU Trans-med Awards Research 20210106, Clinical Research Plan of SHDC (SHDC2020CR6028), National Natural Science Foundation of China (No.62171283), Natural Science Foundation of Shanghai (20ZR1426300), and Shanghai Municipal Science and Technology Major Project (2021SHZDZX0102). It is appreciated that Zixin Wang and **gyu Du from Shanghai Jiao Tong University helped provide valuable advice to this work. Data used in the preparation of this article were obtained from the Adolescent Brain Cognitive Development (ABCD) Study (https://abcdstudy.org), held in the NIMH Data Archive (NDA). This is a multisite, longitudinal study designed to recruit more than 10,000 children age 9–10 and follow them over 10 years into early adulthood. The ABCD Study® is supported by the National Institutes of Health and additional federal partners under award numbers U01DA041048, U01DA050989, U01DA051016, U01DA041022, U01DA051018, U01DA051037, U01DA050987, U01DA041174, U01DA041106, U01DA041117, U01DA041028, U01DA041134, U01DA050988, U01DA051039, U01DA041156, U01DA041025, U01DA041120, U01DA051038, U01DA041148, U01DA041093, U01DA041089, U24DA041123, U24DA041147. A full list of supporters is available at https://abcdstudy.org/federal-partners.html. A listing of participating sites and a complete listing of the study investigators can be found at https://abcdstudy.org/consortium_members/. ABCD consortium investigators designed and implemented the study and/or provided data but did not necessarily participate in the analysis or writing of this report. This manuscript reflects the views of the authors and may not reflect the opinions or views of the NIH or ABCD consortium investigators. The ABCD data repository grows and changes over time. The ABCD data used in this report came from version 2.0.1.
Author information
Authors and Affiliations
Contributions
ZBW: design and implementation of study, figure plotting, drafting and revising the manuscript. XCZ: data analysis and revising the manuscript. YYG: acquisition of data and interpretation of the literature about ADHD. HL and MHL: supervising the project, design of study, and revising the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Z., Zhou, X., Gui, Y. et al. Multiple measurement analysis of resting-state fMRI for ADHD classification in adolescent brain from the ABCD study. Transl Psychiatry 13, 45 (2023). https://doi.org/10.1038/s41398-023-02309-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02309-5
- Springer Nature Limited