Abstract
The Wnt signaling pathways play fundamental roles during both development and adult homeostasis. Aberrant activation of the canonical Wnt signal transduction pathway is involved in many diseases including cancer, and is especially implicated in the development and progression of colorectal cancer. Although extensively studied, new genes, mechanisms and regulatory modulators involved in Wnt signaling activation or silencing are still being discovered. Here we applied a genome-scale CRISPR-Cas9 knockout (KO) screen based on Wnt signaling induced cell survival to reveal new inhibitors of the oncogenic, canonical Wnt pathway. We have identified several potential Wnt signaling inhibitors and have characterized the effects of the initiation factor DExH-box protein 29 (DHX29) on the Wnt cascade. We show that KO of DHX29 activates the Wnt pathway leading to upregulation of the Wnt target gene cyclin-D1, while overexpression of DHX29 inhibits the pathway. Together, our data indicate that DHX29 may function as a new canonical Wnt signaling tumor suppressor and demonstrates that this screening approach can be used as a strategy for rapid identification of novel Wnt signaling modulators.
Similar content being viewed by others
Introduction
The Wnt signal transduction pathways, which are essential for both embryonic development and adult homeostasis, regulate numerous fundamental cell functions, including proliferation, migration, apoptosis, stem cell renewal, and differentiation [1,2,3]. The Wnt pathways can be broadly divided into the canonical-β-catenin dependent, and the more diverse non-canonical-β-catenin independent pathways [4]. Aberrant activation of canonical Wnt signaling is associated with a number of human diseases, including a variety of malignancies such as gastric, breast, liver, and colorectal cancer (CRC) [5].
The canonical Wnt/β-catenin pathway, similarly to other signaling cascades, is initiated at the cell membrane and its primary output involves changes in gene transcriptional programs. These changes occur by regulating the expression levels, post-translational modifications, and subcellular localization, of the Wnt signaling key effector-β-catenin [2, 6, 7]. In unstimulated cells, the Wnt-signalling cascade is silenced due to the activity of a dedicated cytoplasmic destruction complex that phosphorylates β-catenin, marking it for ubiquitination and subsequent degradation. At the core of this complex are the tumour suppressor adenomatous polyposis coli (APC), the scaffold protein axin, two kinases: glycogen synthase kinase-3 (GSK-3) and casein kinase 1 (CK1), and the E3-ubiquitin ligase β-TrCP [8]. Mutations in these components may lead to uncontrolled activation of the pathway and the development of cancer [5].
The Wnt-signalling cascade initiates with the binding of secreted Wnt glycoproteins to a receptor complex composed of frizzled (Fz) and low-density lipoprotein receptor-related protein 5 or 6 (LRP5/6). The binding of a Wnt ligand to the FZD-LRP5/6 complex results in recruitment of the cytoplasmic protein dishevelled, and the subsequent formation of large “signalosomes”. This process leads to disassembly of the destruction complex and stabilization of β-catenin, which translocates into the nucleus, where it associates with T-cell factor/lymphoid enhancer-binding factor (TCF/LEF) transcription factors and other components. The resultant nuclear complex upregulates Wnt target genes and is predicted to be a preferred target for novel Wnt-signalling specific therapeutic approaches [1, 2, 9, 10].
The Wnt cascade is extremely complex and is tightly regulated at different epistatic levels depending on the cellular and environmental context [11]. However, despite decades of research, crucial mechanistic gaps throughout the pathway remain to be filled, and new pathway components are still being identified [12].
To reveal unknown regulators of Wnt/β-catenin signaling, we designed and performed a genetic screen using a Genome-wide Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas9 pooled Knock-out (KO) Library, based on Wnt signaling induced cell survival. Using next-generation sequencing (NGS), we identified several new Wnt signaling inhibitors, of which, we focused on the initiation factor DExH-box protein 29 (DHX29).
Results
Establishment of a GeCKO screening system based on Wnt-induced cell survival under antibiotic selection
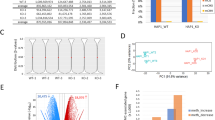
The aim of the study was to use a CRISPR library in order to identify novel regulators of the canonical Wnt signaling pathway. Knocking out potential pathway repressors leads to pathway activation and subsequent hygromycin resistance of the cells mediated by a reporter plasmid TCF/HSV-TK, which we have previously used as a screening tool [13] (Fig. 1A). The reporter was stably transfected into HEK293 cells, in which the β-catenin destruction complex is active and the level of Wnt signaling activity is therefore minimal. We speculated that cells in which a Wnt inhibitor is silenced would become resistant to the hygromycin B antibiotic since the hygromycin resistance gene is regulated by the TCF-binding sites. Preliminary assays were conducted to determine hygromycin concentration and screening duration by comparing a library transduced sample with a nontransduced control sample. Following 10 days of selection with 150µgr/ml hygromycin no live cells were found in the control sample (Fig. S4). Two independent screens were conducted, followed by genomic DNA and deep sequencing analysis. Examining the sgRNA frequency distribution following hygromycin selection revealed that a subset of guide-RNAs was enriched in two independent screen repeats (Fig. 1B). Although a greater enrichment of gRNAs occurring in the cell population treated with hygromycin was expected, similar results were obtained in other screens [14, 15]. As shown in the publication of Shalem et al. longer screening periods further enrich the number of positive guides selected. Our results are comparable to the early timepoints described in these studies. Furthermore, while longer timepoints would increase the magnitude of the fold enrichment, it would probably not change the identity of the selected guides. The pooled library used contains multiple sgRNAs for each gene, and for most enriched genes, more than one sgRNA targeting the same transcript was enriched in the selected cells (Fig. 1C). Ranking the enriched genes using the MAGeCK algorithm [FAP patient Biopsy samples Adenoma and healthy surrounding tissue samples were collected in liquid nitrogen. RNA was extracted using the AllPrep DNA/RNA/protein kit (QIAGEN) following manufacture’s protocol. The samples were obtained for a previous study [19], which was approved by the local IRB committee and registered at the NIH website (NCT02175914). All patients or their legal guardians signed informed consent forms prior to study enrollment. Equal amounts of HEK293-DHX29 or NT1 knockout cells were pelleted and sent for Mass spectrometry analysis at the Smoler proteomics center, Israel Institute of Technology. The samples were digested by trypsin, analyzed by LC-MS/MS on Q-Exactive (Thermo), and identified by Discoverer software against a Human database. The results were semi-quantified by calculating the peak area of each peptide as the average of the three most intense peptides from each protein. The area scores of the protein profile from the HEK293-DHX29 KO sample were compared to that in the NT1 control to identify up and downregulated proteins. Data were analyzed using Graphpad Prism software (version 9.0, GraphPad, La Jolla, CA) and the results are presented as the mean with standard deviation of 3–5 repeats. An unpaired t-test or analysis of variance (ANOVA) to assess the significance of variations; multiple comparisons were conducted according to software recommendations.Mass spectrometry analysis
Statistical methods
References
Flanagan DJ, Vincan E, Phesse TJ. Wnt signaling in cancer: not a binary ON:OFF switch. Cancer Res. 2019;79:5901–6.
Nusse R, Clevers H. Wnt/beta-catenin signaling, disease, and emerging therapeutic modalities. Cell. 2017;169:985–99.
Steinhart Z, Angers S. Wnt signaling in development and tissue homeostasis. Development. 2018;145:1–8.
Gordon MD, Nusse R. Wnt signaling: multiple pathways, multiple receptors, and multiple transcription factors. J Biol Chem. 2006;281:22429–33.
Zhan T, Rindtorff N, Boutros M. Wnt signaling in cancer. Oncogene. 2017;36:1461–73.
Hanley MP, Hahn MA, Li AX, Wu X, Lin J, Wang J, et al. Genome-wide DNA methylation profiling reveals cancer-associated changes within early colonic neoplasia. Oncogene. 2017;36:5035–44.
Stamos JL, Chu ML, Enos MD, Shah N, Weis WI. Structural basis of GSK-3 inhibition by N-terminal phosphorylation and by the Wnt receptor LRP6. Elife. 2014;3:e01998.
Stamos JL, Weis WI. The beta-catenin destruction complex. Cold Spring Harb Perspect Biol. 2013;5:a007898.
Angers S, Moon RT. Proximal events in Wnt signal transduction. Nat Rev Mol Cell Biol. 2009;10:468–77.
Caspi M, Wittenstein A, Kazelnik M, Shor-Nareznoy Y, Rosin-Arbesfeld R. Therapeutic targeting of the oncogenic Wnt signaling pathway for treating colorectal cancer and other colonic disorders. Adv Drug Deliv Rev. 2021;169:118–36.
Wiese KE, Nusse R, van Amerongen R. Wnt signalling: conquering complexity. Development. 2018;145:1–9.
Ji L, Lu B, Zamponi R, Charlat O, Aversa R, Yang Z, et al. USP7 inhibits Wnt/beta-catenin signaling through promoting stabilization of Axin. Nat Commun. 2019;10:4184.
Skalka N, Caspi M, Caspi E, Loh YP, Rosin-Arbesfeld R, Carboxypeptidase E. a negative regulator of the canonical Wnt signaling pathway. Oncogene. 2013;32:2836–47.
Dukhovny A, Lamkiewicz K, Chen Q, Fricke M, Jabrane-Ferrat N, et al. A CRISPR activation screen identifies genes that protect against zika virus infection. J Virol. 2019;93:211–19.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelson T, et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 2014;343:84–87.
Li W, Xu H, **ao T, Cong L, Love MI, Zhang F, et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 2014;15:554.
MacDonald BT, Tamai K, He X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell. 2009;17:9–26.
**e M, Zhao F, Zou X, ** S, **ong S. The association between CCND1 G870A polymorphism and colorectal cancer risk: a meta-analysis. Medicine (Baltimore). 2017;96:e8269.
Kariv R, Caspi M, Fliss-Isakov N, Shorer Y, Shor Y, Rosner G, et al. Resorting the function of the colorectal cancer gatekeeper adenomatous polyposis coli. Int J Cancer. 2020;146:1064–74.
Červenka I, Wolf J, Mašek J, Krejci P, Wilcox WR, Kozubík A, et al. Mitogen-activated protein kinases promote WNT/beta-catenin signaling via phosphorylation of LRP6. Mol Cell Biol. 2011;31:179–89.
Kim W, Kim SY, Kim T, Kim M, Bae DJ, Choi HI, et al. ADP-ribosylation factors 1 and 6 regulate Wnt/beta-catenin signaling via control of LRP6 phosphorylation. Oncogene. 2013;32:3390–6.
Poncet N, Halley PA, Lipina C, Gierliński M, Dady A, Singer GA, et al. Wnt regulates amino acid transporter Slc7a5 and so constrains the integrated stress response in mouse embryos. EMBO Rep. 2020;21:e48469.
Yu Y, Xu P, Cui G, Xu X, Li K, Chen X, et al. UBQLN4 promotes progression of HCC via activating wnt-beta-catenin pathway and is regulated by miR-370. Cancer Cell Int. 2020;20:3.
Zhu Y, Du Y, Zhang Y. DHX33 promotes colon cancer development downstream of Wnt signaling. Gene. 2020;735:144402.
Park J, Son Y, Lee NG, Lee K, Lee DG, Song J, et al. DSG2 is a functional cell surface marker for identification and isolation of human pluripotent stem cells. Stem Cell Rep. 2018;11:115–27.
Chen H, Shen HX, Lin YW, Mao YQ, Liu B, **e LP. Small RNA-induced INTS6 gene up-regulation suppresses castration-resistant prostate cancer cells by regulating beta-catenin signaling. Cell Cycle. 2018;17:1602–13.
Galiatsatos P, Foulkes WD. Familial adenomatous polyposis. Am J Gastroenterol. 2006;101:385–98.
Shalem O, Sanjana NE, Zhang F. High-throughput functional genomics using CRISPR-Cas9. Nat Rev Genet. 2015;16:299–311.
Miles LA, Garippa RJ, Poirier JT. Design, execution, and analysis of pooled in vitro CRISPR/Cas9 screens. FEBS J. 2016;283:3170–80.
Schuster A, Erasimus H, Fritah S, Nazarov PV, van Dyck E, Niclou SP, et al. RNAi/CRISPR screens: from a Pool to a Valid Hit. Trends Biotechnol. 2019;37:38–55.
Sun QY, Ding LW, **ao JF, Chien W, Lim SL, Hattori N, et al. SETDB1 accelerates tumourigenesis by regulating the WNT signalling pathway. J Pathol. 2015;235:559–70.
Major MB, Roberts BS, Berndt JD, Marine S, Anastas J, Chung N, et al. New regulators of Wnt/beta-catenin signaling revealed by integrative molecular screening. Sci Signal. 2008;1:ra12.
Wan C, Mahara S, Sun C, Doan A, Chua HK, et al. Genome-scale CRISPR-Cas9 screen of Wnt/β-catenin signaling identifies therapeutic targets for colorectal cancer. Sci Adv. 2021;7. PMID: 34138730.
Callow MG, Tran H, Phu L, Lau T, Lee J, Sandoval WN, et al. Ubiquitin ligase RNF146 regulates tankyrase and Axin to promote Wnt signaling. PLoS ONE. 2011;6:e22595.
DasGupta R, Kaykas A, Moon RT, Perrimon N. Functional genomic analysis of the Wnt-wingless signaling pathway. Science. 2005;308:826–33.
Firestein R, Bass AJ, Kim SY, Dunn IF, Silver SJ, Guney I, et al. CDK8 is a colorectal cancer oncogene that regulates beta-catenin activity. Nature. 2008;455:547–51.
Lebensohn AM, Dubey R, Neitzel LR, Tacchelly-Benites O, Yang E, et al. Comparative genetic screens in human cells reveal new regulatory mechanisms in WNT signaling. Elife. 2016;5:1–40.
Murakami K, Terakado Y, Saito K, Jomen Y, Takeda H, Oshima M, et al. A genome-scale CRISPR screen reveals factors regulating Wnt-dependent renewal of mouse gastric epithelial cells. Proc Natl Acad Sci USA. 2021;118:118.
Steinhart Z, Pavlovic Z, Chandrashekhar M, Hart T, Wang X, Zhang X, et al. Genome-wide CRISPR screens reveal a Wnt-FZD5 signaling circuit as a druggable vulnerability of RNF43-mutant pancreatic tumors. Nat Med. 2017;23:60–68.
Tang W, Dodge M, Gundapaneni D, Michnoff C, Roth M, Lum L. A genome-wide RNAi screen for Wnt/beta-catenin pathway components identifies unexpected roles for TCF transcription factors in cancer. Proc Natl Acad Sci USA. 2008;105:9697–702.
Parsyan A, Shahbazian D, Martineau Y, Petroulakis E, Alain T, Larsson O, et al. The helicase protein DHX29 promotes translation initiation, cell proliferation, and tumorigenesis. Proc Natl Acad Sci USA. 2009;106:22217–22.
Pisareva VP, Pisarev AV. DHX29 reduces leaky scanning through an upstream AUG codon regardless of its nucleotide context. Nucleic Acids Res. 2016;44:4252–65.
Pisareva VP, Pisarev AV, Komar AA, Hellen CU, Pestova TV. Translation initiation on mammalian mRNAs with structured 5’UTRs requires DExH-box protein DHX29. Cell. 2008;135:1237–50.
Attar-Schneider O, Drucker L, Gottfried M. Migration and epithelial-to-mesenchymal transition of lung cancer can be targeted via translation initiation factors eIF4E and eIF4GI. Lab Invest. 2016;96:1004–15.
Zhu Q, Tan P, Li Y, Lin M, Li C, Mao J, et al. DHX29 functions as an RNA co-sensor for MDA5-mediated EMCV-specific antiviral immunity. PLoS Pathog. 2018;14:e1006886.
Cruciat CM, Dolde C, de Groot RE, Ohkawara B, Reinhard C, Korswagen HC, et al. RNA helicase DDX3 is a regulatory subunit of casein kinase 1 in Wnt-beta-catenin signaling. Science. 2013;339:1436–41.
Yang F, Fang E, Mei H, Chen Y, Li H, Li D, et al. Cis-acting circ-CTNNB1 promotes beta-catenin signaling and cancer progression via DDX3-mediated transactivation of YY1. Cancer Res. 2019;79:557–71.
Perfetto M, Xu X, Lu C, Shi Y, Yousaf N, et al. The RNA helicase DDX3 induces neural crest by promoting AKT activity. Development. 2021;148:PMC7847268.
Wang Y, He K, Sheng B, Lei X, Tao W, et al. The RNA helicase Dhx15 mediates Wnt-induced antimicrobial protein expression in Paneth cells. Proc Natl Acad Sci USA. 2021;118. https://doi.org/10.1073/pnas.2111936118.
Zhang M, Weng W, Zhang Q, Wu Y, Ni S, Tan C, et al. The lncRNA NEAT1 activates Wnt/beta-catenin signaling and promotes colorectal cancer progression via interacting with DDX5. J Hematol Oncol. 2018;11:113.
Cohen J, Raviv S, Adir O, Padmanabhan K, Soffer A, Luxenburg C. The Wave complex controls epidermal morphogenesis and proliferation by suppressing Wnt-Sox9 signaling. J Cell Biol. 2019;218:1390–406.
Caspi E, Rosin-Arbesfeld R. A novel functional screen in human cells identifies MOCA as a negative regulator of Wnt signaling. Mol Biol Cell. 2008;19:4660–74.
Morin PJ, Sparks AB, Korinek V, Barker N, Clevers H, Vogelstein B, et al. Activation of beta-catenin-Tcf signaling in colon cancer by mutations in beta-catenin or APC. Science. 1997;275:1787–90.
Joung J, Konermann S, Gootenberg JS, Abudayyeh OO, Platt RJ, Brigham MD, et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat Protoc. 2017;12:828–63.
Golan T, Yaniv A, Bafico A, Liu G, Gazit A. The human Frizzled 6 (HFz6) acts as a negative regulator of the canonical Wnt. beta-catenin signaling cascade. J Biol Chem. 2004;279:14879–88.
Chen S, Sanjana NE, Zheng K, Shalem O, Lee K, Shi X, et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell. 2015;160:1246–60.
Wang B, Wang M, Zhang W, **ao T, Chen CH, Wu A, et al. Integrative analysis of pooled CRISPR genetic screens using MAGeCKFlute. Nat Protoc. 2019;14:756–80.
Chavez A, Scheiman J, Vora S, Pruitt BW, Tuttle M, P R Iyer E, et al. Highly efficient Cas9-mediated transcriptional programming. Nat Methods. 2015;12:326–8.
Gilbert LA, Horlbeck MA, Adamson B, Villalta JE, Chen Y, Whitehead EH, et al. Genome-scale CRISPR-Mediated control of gene repression and activation. Cell. 2014;159:647–61.
Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C, et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature. 2015;517:583–8.
Maeder ML, Linder SJ, Cascio VM, Fu Y, Ho QH, Joung JK. CRISPR RNA-guided activation of endogenous human genes. Nat Methods. 2013;10:977–9.
Mali P, Aach J, Stranges PB, Esvelt KM, Moosburner M, Kosuri S, et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat Biotechnol. 2013;31:833–8.
Perez-Pinera P, Kocak DD, Vockley CM, Adler AF, Kabadi AM, Polstein LR, et al. RNA-guided gene activation by CRISPR-Cas9-based transcription factors. Nat Methods. 2013;10:973–6.
Tanenbaum ME, Gilbert LA, Qi LS, Weissman JS, Vale RD. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell. 2014;159:635–46.
Author information
Authors and Affiliations
Contributions
TE performed the vast majority of the experiments and analysis and edited the manuscript; MC helped in experimental design and manuscript preparation; MK, YS, and SAE conducted experimental work; RK performed the colonoscopies and provided the samples; ZM and RE performed bioinformatics analysis; EHS contributed the to the screen methodology, analyzed the data, and helped in experimental design; RRA contributed the idea, oversaw the project, analyzed the data, and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Evron, T., Caspi, M., Kazelnik, M. et al. A CRISPR knockout screen reveals new regulators of canonical Wnt signaling. Oncogenesis 10, 63 (2021). https://doi.org/10.1038/s41389-021-00354-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-021-00354-7
- Springer Nature Limited
This article is cited by
-
Wnt signaling regulates chemokine production and cell migration of circulating human monocytes
Cell Communication and Signaling (2024)
-
CRISPR/Cas9: a powerful tool in colorectal cancer research
Journal of Experimental & Clinical Cancer Research (2023)
-
Wnt Signaling in Heart Development and Regeneration
Current Cardiology Reports (2022)