Abstract
The timing of puberty is a highly polygenic childhood trait that is epidemiologically associated with various adult diseases. Using 1000 Genomes Project–imputed genotype data in up to ∼370,000 women, we identify 389 independent signals (P < 5 × 10−8) for age at menarche, a milestone in female pubertal development. In Icelandic data, these signals explain ∼7.4% of the population variance in age at menarche, corresponding to ∼25% of the estimated heritability. We implicate ∼250 genes via coding variation or associated expression, demonstrating significant enrichment in neural tissues. Rare variants near the imprinted genes MKRN3 and DLK1 were identified, exhibiting large effects when paternally inherited. Mendelian randomization analyses suggest causal inverse associations, independent of body mass index (BMI), between puberty timing and risks for breast and endometrial cancers in women and prostate cancer in men. In aggregate, our findings highlight the complexity of the genetic regulation of puberty timing and support causal links with cancer susceptibility.



Similar content being viewed by others
Accession codes
References
Parent, A.S. et al. The timing of normal puberty and the age limits of sexual precocity: variations around the world, secular trends, and changes after migration. Endocr. Rev. 24, 668–693 (2003).
Perry, J.R., Murray, A., Day, F.R. & Ong, K.K. Molecular insights into the aetiology of female reproductive ageing. Nat. Rev. Endocrinol. 11, 725–734 (2015).
Perry, J.R. et al. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature 514, 92–97 (2014).
Lunetta, K.L. et al. Rare coding variants and X-linked loci associated with age at menarche. Nat. Commun. 6, 7756 (2015).
Day, F.R. et al. Shared genetic aetiology of puberty timing between sexes and with health-related outcomes. Nat. Commun. 6, 8842 (2015).
Westra, H.J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Finucane, H.K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Preprint at bioRxiv https://dx.doi.org/10.1101/103069 (2017).
Rao, S.S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin loo**. Cell 159, 1665–1680 (2014).
Smemo, S. et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 507, 371–375 (2014).
Mbarek, H. et al. Identification of common genetic variants influencing spontaneous dizygotic twinning and female fertility. Am. J. Hum. Genet. 98, 898–908 (2016).
Ong, K.K. et al. Genetic variation in LIN28B is associated with the timing of puberty. Nat. Genet. 41, 729–733 (2009).
Perry, J.R. et al. Meta-analysis of genome-wide association data identifies two loci influencing age at menarche. Nat. Genet. 41, 648–650 (2009).
Zhang, J. et al. LIN28 regulates stem cell metabolism and conversion to primed pluripotency. Cell Stem Cell 19, 66–80 (2016).
Zhu, H. et al. Lin28a transgenic mice manifest size and puberty phenotypes identified in human genetic association studies. Nat. Genet. 42, 626–630 (2010).
Zhu, H. et al. The Lin28/let-7 axis regulates glucose metabolism. Cell 147, 81–94 (2011).
Baran, Y. et al. The landscape of genomic imprinting across diverse adult human tissues. Genome Res. 25, 927–936 (2015).
van den Berg, S.M. & Boomsma, D.I. The familial clustering of age at menarche in extended twin families. Behav. Genet. 37, 661–667 (2007).
Ahlgren, M., Melbye, M., Wohlfahrt, J. & Sørensen, T.I. Growth patterns and the risk of breast cancer in women. N. Engl. J. Med. 351, 1619–1626 (2004).
Collaborative Group on Hormonal Factors in Breast Cancer. Menarche, menopause, and breast cancer risk: individual participant meta-analysis, including 118 964 women with breast cancer from 117 epidemiological studies. Lancet Oncol. 13, 1141–1151 (2012).
Bhaskaran, K. et al. Body-mass index and risk of 22 specific cancers: a population-based cohort study of 5·24 million UK adults. Lancet 384, 755–765 (2014).
Locke, A.E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Gao, C. et al. Mendelian randomization study of adiposity-related traits and risk of breast, ovarian, prostate, lung and colorectal cancer. Int. J. Epidemiol. 45, 896–908 (2016).
Guo, Y. et al. Genetically predicted body mass index and breast cancer risk: Mendelian randomization analyses of data from 145,000 women of European descent. PLoS Med. 13, e1002105 (2016).
Abreu, A.P. et al. Central precocious puberty caused by mutations in the imprinted gene MKRN3. N. Engl. J. Med. 368, 2467–2475 (2013).
da Rocha, S.T., Edwards, C.A., Ito, M., Ogata, T. & Ferguson-Smith, A.C. Genomic imprinting at the mammalian Dlk1–Dio3 domain. Trends Genet. 24, 306–316 (2008).
Giles, G.G. et al. Early growth, adult body size and prostate cancer risk. Int. J. Cancer 103, 241–245 (2003).
de Vries, L., Kauschansky, A., Shohat, M. & Phillip, M. Familial central precocious puberty suggests autosomal dominant inheritance. J. Clin. Endocrinol. Metab. 89, 1794–1800 (2004).
Wehkalampi, K., Widén, E., Laine, T., Palotie, A. & Dunkel, L. Patterns of inheritance of constitutional delay of growth and puberty in families of adolescent girls and boys referred to specialist pediatric care. J. Clin. Endocrinol. Metab. 93, 723–728 (2008).
Winkler, T.W. et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 9, 1192–1212 (2014).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Day, F.R., Elks, C.E., Murray, A., Ong, K.K. & Perry, J.R. Puberty timing associated with diabetes, cardiovascular disease and also diverse health outcomes in men and women: the UK Biobank study. Sci. Rep. 5, 11208 (2015).
Bulik-Sullivan, B.K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Loh, P.R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Segrè, A.V., Groop, L., Mootha, V.K., Daly, M.J. & Altshuler, D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, e1001058 (2010).
Finucane, H.K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Gamazon, E.R. et al. A gene-based association method for map** traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Sarkar, A., Ward, L.D. & Kellis, M. Functional enrichments of disease variants across thousands of independent loci in eight diseases. Preprint at bioRxiv http://dx.doi.org/10.1101/048066 (2016).
Ernst, J. et al. Map** and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Acknowledgements
This research has been conducted using the UK Biobank Resource under applications 5122 and 9797. Full study-specific acknowledgments can be found in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
All authors reviewed the original and revised manuscripts. Statistical analysis: F.R.D., D.J.T., H.H., D.I.C., H.F., P.S., K.S.R., S.W., A.K.S., E. Albrecht, E. Altmaier, M.A., C.M.B., T. Boutin, A. Campbell, E.D., A.G., C. He, J.J.H., R.K., I.K., P.-R.L., K.L.L., M.M., B.M., G.M., S.E.M., I.M.N., R.N., T.N., L.P., N. Perjakova, E.P., L.M.R., K.E.S., A.V. Segrè, A.V. Smith, L.S., A.T., J.R.B.P. Sample collection, genoty** and phenoty**: I.L.A., S. Bandinelli, M.W.B., J.B., S. Bergmann, M.B., E.B., S.E.B., M.K.B., J.S.B., H. Brauch, H. Brenner, L.B., T. Brüning, J.E.B., H.C., E.C., S.C., G.C.-T., T.C., F.J.C., D.L.C., A. Cox, L.C., K.C., G.D.S., E.J.C.N.d., R.d., I.D.V., J.D., P.D., I.d.-S.-S., A.M.D., J.G.E., P.A.F., L.F.-R., L. Ferrucci, D.F.-J., L. Franke, M.G., I.G., G.G.G., H.G., D.F.G., P.G., P.H., E.H., U.H., T.B.H., C.A.H., G.H., M.J.H., J.L.H., F.H., D.J.H., H.K.I., M.-R.J., P.K.J., D.K., Z.K., G.L., D.L., C.L., L.J.L., J.S.E.L., S. Lenarduzzi, J. Li, P.A.L., S. Lindstrom, Y.L., J. Luan, R.M., A. Mannermaa, H.M., M.I.M., C. Meisinger, T.M., C. Menni, A. Metspalu, K.M., L.M., R.L.M., G.W.M., A.M.M., M.A.N., P.N., H.N., D.R.N., A.J.O., T.A.O., S.P., A. Palotie, N. Pedersen, A. Peters, J.P., P.D.P.P., A. Pouta, P.R., I. Rahman, S.M.R., A.R., F.R.R., I. Rudan, R.R., D.R., C.F.S., M.K.S., R.A.S., M. Shah, R.S., M.C.S., U.S., M. Stampfer, M. Steri, K. Strauch, T. Tanaka, E.T., N.J.T., M.T., T. Truong, J.P.T., A.G.U., D.R.V.E., V.V., U.V., P.V., Q.W., E.W., K.W.v.D., G.W., R.W., B.H.R.W., J.H.Z., M. Zoledziewska, M. Zygmunt. Individual study principal investigators: B.Z.A., D.I.B., M.C., F.C., T.E., N.F., C.G., V.G., C. Hayward, P.K., D.A.L., P.K.E.M., N.G.M., D.O.M.-K., E.A.N., O.P., D.P., A.L.P., P.M.R., H.S., T.D.S., D.S., D.T., S.U., J.A.V., H.V., N.J.W., J.F.W., A.B.S., U.T., K.S.P., D.F.E., J.Y.T., J.C.-C., D.H., A. Murray, J.M.M., K. Stefansson, K.K.O., J.R.B.P. Working group: F.R.D., D.J.T., H.H., D.I.C., H.F., P.S., K.S.R., S.W., A.K.S., A.B.S., U.T., K.S.P., D.F.E., J.Y.T., J.C., D.H., A. Murray, J.M.M., K. Stefansson, K.K.O., J.R.B.P.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
A full list of members and affiliations appears in the Supplementary Note.
Integrated supplementary information
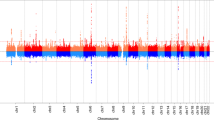
Supplementary Figure 1 Manhattan plot displaying the genomic locations of the 389 genome-wide significant loci.
Previously identified genome-wide significant loci are shown in gold, and new loci are shown in purple. SNPs within 300 kb of the lead SNP at each locus are highlighted. The y axis has been truncated above 30.
Supplementary Figure 2 Number of genome-wide significant menarche loci per chromosome by chromosome size.
The X chromosome is highlighted in red.
Supplementary Figure 4 LocusZoom plots of menarche-associated variants at the MKRN3 locus (hg38) in the deCODE study, Iceland.
The 5′ UTR variant rs530324840 at position chr15:23565461 is labeled as a diamond and shown in purple; other variants are colored according to correlation (r2) with rs530324840 (see legends). –log10 P values are shown along the left y axis, and the right y axis corresponds to recombination rate, plotted as a solid gray line. (a) Associations under the paternal model, where the signal near 24 Mb corresponds to the common reported variant rs12148769. (b) Associations under the maternal model. (c) Associations under the additive model.
Supplementary Figure 5 LocusZoom plots of menarche-associated variants at the DLK1 locus (hg38) in the deCODE study, Iceland.
The variant rs138827001 at position chr14:100771634 is labeled as a diamond and shown in purple; other variants are colored according to correlation (r2) with rs138827001 (see legends). –log10 P values are shown along the left y axis, and the right y axis corresponds to recombination rate, plotted as a solid gray line. (a) Associations under the paternal model. (b) Associations under the maternal model. (c) Associations under the additive model.
Supplementary Figure 6 Association of menarche-associated variants with adult body mass index.
The blue bars (and left y axis) indicate the collective variance explained in adult BMI (with bootstrap-generated 95% CIs) by index menarche-associated SNPs grouped by their individual associations with BMI (in UK Biobank using an additive model controlling for chip and principal components). The red line (and right y axis) indicates the –log10 P value for the collective association with BMI of each group of SNPs. Purple squares correspond to collective associations with BMI at P < 0.05.
Supplementary Figure 7 Dose-response plots for Mendelian randomization analyses.
(a–d) The individual effects on AAM of the 314 ‘BMI-unrelated’ autosomal AAM variants are plotted against risks for breast (a), prostate (b), endometrial (c) and ovarian (d) cancer. Red line, IVW regression; blue line, MR-Egger regression. MR-Egger supports the protective effect of later age at puberty on breast, prostate and endometrial cancers, but indicates pleiotropy in the association with ovarian cancer.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note (PDF 2390 kb)
Supplementary Tables
Supplementary Tables 1–16 (XLSX 5253 kb)
Rights and permissions
About this article
Cite this article
Day, F., Thompson, D., Helgason, H. et al. Genomic analyses identify hundreds of variants associated with age at menarche and support a role for puberty timing in cancer risk. Nat Genet 49, 834–841 (2017). https://doi.org/10.1038/ng.3841
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3841
- Springer Nature America, Inc.
This article is cited by
-
Metabolome-wide Mendelian randomization for age at menarche and age at natural menopause
Genome Medicine (2024)
-
Assessing causal links between age at menarche and adolescent mental health: a Mendelian randomisation study
BMC Medicine (2024)
-
Persistent thinness and anorexia nervosa differ on a genomic level
European Journal of Human Genetics (2024)
-
Prenatal, newborn and childhood factors and the timing of puberty in boys and girls
Pediatric Research (2024)
-
Genome-wide association and expression quantitative trait loci in cattle reveals common genes regulating mammalian fertility
Communications Biology (2024)