Abstract
Background
Several genetic variants of thiopurine metabolic pathway are associated with 6-thiopurine-mediated leucopenia. A population-based evaluation of these variants lays the foundation for Pharmacogenetic-guided thiopurine therapy.
Methods
A total of 2000 subjects were screened for the pharmacogenetic determinants using the infinium global screening array (GSA). The functional relevance of these variants was deduced using SNAP2, SIFT, Provean, Mutalyzer, Mutation Taster, Phyre2, SwissDock, AGGRESCAN, and CUPSAT.
Results
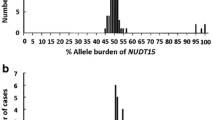
The minor allele frequencies of NUDT15*3, NUDT15*5, TPMT*3C, TPMT*3B variant alleles were 6.78%, 0.11%, 1.98% and 0.69%, respectively. TPMT*3A genotype was observed in 0.35% subjects. No gender-based differences were observed in the incidence of these variants. Data from studies of the Indian population showed that 92.86% subjects heterozygous for NUDT15*3 and 60% subjects heterozygous for TPMT*3C exhibit thiopurine-mediated hematological toxicity. NUDT15 variants have no impact on the binding of ‘dGTP’ to the NUDT protein. NUDT15*3 variant increases aggregation ‘hot spot’ region and induces unfavourable torsion in the protein. NUDT15*5 destabilizes the protein and impairs Mg/Mn binding. TPMT*3A, TPMT*3B and TPMT*3C variants lower binding affinity to 6-mercaptopurine compared to the wild protein. TPMT*3C variant destabilizes the TPMT protein in the thermal experiment. Compared to the data of European and African/African American populations, NUDT15*3 frequency is higher and TPMT*3C frequency is lower in our population.
Conclusions
TPMT variants were less frequent in Indian population, while NUDT15*3 is more frequent compared to European and African/African American populations. NUDT15*3 increases aggregation ‘hot spot’ and induces unfavourable torsion in the protein. NUDT15*5 and TPMT*3C destabilize the respective proteins. TPMT*3A, TPMT*3B and TPMT*3C are associated with a lower binding affinity towards 6-mercaptopurine.
Graphic abstract







Similar content being viewed by others
Abbreviations
- ALFA:
-
Aggregate allele and genotype frequencies computed from non-sensitive dbGaP studies
- ALL:
-
Acute lymphoblastic leukemia
- AZT:
-
Azathioprine
- DNA-TG:
-
DNA-incorporated thioguanine
- GSA:
-
Global Screening array
- IBD:
-
Inflammatory bowel disease
- MAF:
-
Minor allele frequency
- MP:
-
Mercaptopurine
- NUDT15:
-
Nudix hydrolase 15
- PCNA:
-
Proliferating cell nuclear antigen
- SAM:
-
S-Adenosyl methionine
- SIFT:
-
Sorting intolerant from tolerant
- TGN:
-
Thioguanine
- TPMT:
-
Thiopurine S-methyltransferase
References
Carter M, Jemth AS, Hagenkort A, Page BD, Gustafsson R, Griese JJ, et al. Crystal structure, biochemical and cellular activities demonstrate separate functions of MTH1 and MTH2. Nat Commun. 2015;4(6):7871.
Yu Y, Cai JP, Tu B, Wu L, Zhao Y, Liu X, et al. Proliferating cell nuclear antigen is protected from degradation by forming a complex with MutT Homolog2. J Biol Chem. 2009;284(29):19310–20.
Yang SK, Hong M, Baek J, Choi H, Zhao W, Jung Y, et al. A common missense variant in NUDT15 confers susceptibility to thiopurine-induced leukopenia. Nat Genet. 2014;46(9):1017–20.
Evans WE, Hon YY, Bomgaars L, Coutre S, Holdsworth M, Janco R, et al. Preponderance of thiopurine S-methyltransferase deficiency and heterozygosity among patients intolerant to mercaptopurine or azathioprine. J Clin Oncol. 2001;19(8):2293–301.
Sato T, Takagawa T, Kakuta Y, Nishio A, Kawai M, Kamikozuru K, et al. NUDT15, FTO, and RUNX1 genetic variants and thiopurine intolerance among Japanese patients with inflammatory bowel diseases. Intest Res. 2017;15(3):328–37.
Fei X, Shu Q, Zhu H, Hua B, Wang S, Guo L, et al. NUDT15 R139C variants increase the risk of azathioprine-induced leukopenia in Chinese autoimmune patients. Front Pharmacol. 2018;9:460.
Moriyama T, Nishii R, Lin TN, Kihira K, Toyoda H, Jacob N, et al. The effects of inherited NUDT15 polymorphisms on thiopurine active metabolites in Japanese children with acute lymphoblastic leukemia. Pharmacogenet Genomics. 2017;27(6):236–9.
Yang JJ, Landier W, Yang W, Liu C, Hageman L, Cheng C, et al. Inherited NUDT15 variant is a genetic determinant of mercaptopurine intolerance in children with acute lymphoblastic leukemia. J Clin Oncol. 2015;33(11):1235–42.
Moriyama T, Nishii R, Perez-Andreu V, Yang W, Klussmann FA, Zhao X, et al. NUDT15 polymorphisms alter thiopurine metabolism and hematopoietic toxicity. Nat Genet. 2016;48(4):367–73.
Kodidela S, Dorababu P, Thakkar DN, Dubashi B, Sundaram R, Muralidharan N, et al. Association of NUDT15*3 and FPGS 2572C>T variants with the risk of early hematologic toxicity during 6-MP and low-dose methotrexate-based maintenance therapy in Indian Patients with acute lymphoblastic leukemia. Genes (Basel). 2572C;11(6):E594.
Naushad SM, Dorababu P, Rupasree Y, Pavani A, Raghunadharao D, Hussain T, et al. Classification and regression tree-based prediction of 6-mercaptopurine-induced leucopenia grades in children with acute lymphoblastic leukemia. Cancer Chemother Pharmacol. 2019;83(5):875–80.
Shah SAV, Paradkar MU, Desai DC, Ashavaid TF. Preemptive NUDT15 genoty**: redefining the management of patients with thiopurine-induced toxicity. Drug Metab Pers Ther. 2018;33(1):57–60.
Khera S, Trehan A, Bhatia P, Singh M, Bansal D, Varma N. Prevalence of TPMT, ITPA and NUDT 15 genetic polymorphisms and their relation to 6MP toxicity in north Indian children with acute lymphoblastic leukemia. Cancer Chemother Pharmacol. 2019;83(2):341–8.
Valerie NC, Hagenkort A, Page BD, Masuyer G, Rehling D, Carter M, et al. NUDT15 Hydrolyzes 6-thio-deoxyGTP to mediate the anticancer efficacy of 6-thioguanine. Cancer Res. 2016;76(18):5501–11.
Zhang AL, Yang J, Wang H, Lu JL, Tang S, Zhang XJ. Association of NUDT15 c.415C>T allele and thiopurine-induced leukocytopenia in Asians: a systematic review and meta-analysis. Ir J Med Sci. 2018;187(1):145–53.
Cargnin S, Genazzani AA, Canonico PL, Terrazzino S. Diagnostic accuracy of NUDT15 gene variants for thiopurine-induced leukopenia: a systematic review and meta-analysis. Pharmacol Res. 2018;135:102–11.
Liu C, Yang W, Pei D, Cheng C, Smith C, Landier W, et al. Genomewide approach validates thiopurine methyltransferase activity is a monogenic pharmacogenomic trait. Clin Pharmacol Ther. 2017;101(3):373–81.
Rutherford K, Daggett V. Four human thiopurine S-methyltransferase alleles severely affect protein structure and dynamics. J Mol Biol. 2008;379(4):803–14.
Salavaggione OE, Wang L, Wiepert M, Yee VC, Weinshilboum RM. Thiopurine S-methyltransferase pharmacogenetics: variant allele functional and comparative genomics. Pharmacogenet Genom. 2005;15(11):801–15.
Relling MV, Schwab M, Whirl-Carrillo M, Suarez-Kurtz G, Pui CH, Stein CM, et al. Clinical pharmacogenetics implementation consortium guideline for thiopurine dosing based on TPMT and NUDT15 genotypes: 2018 update. Clin Pharmacol Ther. 2019;105(5):1095–105.
Acknowledgements
The study was funded by intramural Grants of Sandor Speciality Diagnostics Pvt Ltd, Hyderabad, India. The authors are grateful to the Deanship of Scientific Research, King Saud University for funding through Vice Deanship of Scientific Research Chairs.
Author information
Authors and Affiliations
Contributions
The study was conceived and designed by SMN and VKK. The recruitment of subjects, extraction of DNA, GSA analysis, and genoty** were performed by SMN, MJR, VKK, TA, and SAA. The data analysis and bioinformatics analysis were performed by SMN, MJR, and TA. The manuscript was drafted by SMN and MJR. The manuscript was revised for the critical content by TA and SAA. The final version of the manuscript was approved by all the authors.
Corresponding author
Ethics declarations
Conflict of interest
All the authors hereby declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Naushad, S.M., Janaki Ramaiah, M., Kutala, V.K. et al. Pharmacogenetic determinants of thiopurines in an Indian cohort. Pharmacol. Rep 73, 278–287 (2021). https://doi.org/10.1007/s43440-020-00158-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43440-020-00158-3