Abstract
The present research was carried out with the objectives of isolating fungal strains capable of exhibiting inulinase activity from a novel source, then optimizing solid state fermentation for the inulinase production and characterizing crude enzyme. Production of inulinase is first reported in this study with a novel substrate, banana plant shoot powder, using newly isolated fungal strain Nothophoma anigozanthi JAM through solid state fermentation. The fungal strain was isolated from the root soil of a white Jamun (Syzygium cumini) tree. Optimization of the solid state fermentation was carried out by central composite design for determining the interaction and impact of the influential process on inulinase production. A mathematical model was developed based on the statistical analysis and recommended optimal process conditions of 2 g of substrate, 5 mL of moistening media, 5 mL of inoculum volume, and 96 h of fermentation time for enhanced production (2.578 U/gds). Crude inulinase enzyme exhibited an 8.0-fold increase in activity after optimization, in comparison with the activity under unoptimized conditions. The optimum pH (5.0), temperature (60 °C) for the enzyme activity, and Km (0.205 mM) and Vmax (0.333 µM/min) were determined based on the biochemical and inulin hydrolytic properties of the crude inulinase enzyme. The production of inulinase was further validated by using TLC, osazone, and Seliwanoff tests. Hence, the inulinase enzyme produced by Nothophoma anigozanthi JAM can be successfully utilized for the production of edible sugar fructose from non-edible polysaccharide inulin, present in agricultural residues such as banana shoot by solid state fermentation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Inulinase, also known as 1-D-fructan fructanohydrolase, is an enzyme that catalyzes the breakdown of inulin to fructose by hydrolyzing the β-2,1-glycosidic bonds endothermically [1]. Inulin is a homopolysaccharide made up of fructose units termed fructans that are linked together by β-2, 1-glycosidic bonds [2, 3]. Inulin is present in agave, onion, dandelion, banana, and garlic. The human digestive system cannot hydrolyze the β-2,1-glycosidic linkages; hence, inulin must be transformed into digestible forms of carbohydrates such as fructose and fructo-oligosaccharides. Hence, the production of the inulinase enzyme plays a significant role in converting non-digestible inulin to digestible carbohydrates through fermentation processes. The sources of inulin are Jerusalem artichoke, agave, chicory, onion, dandelion, banana, wheat, oats, garlic, etc. [4]. X-ray diffraction has been successfully used for the 3D structure determination of inulinase enzymes from Aspergillus ficuum and Aspergillus awamori [5]. Inulinase can be industrially utilized for the production of high fructose syrup, fructo-oligosaccharides, bioethanol, acetone, butanol, organic acids, single-cell proteins, single-cell oil, and 2,3-butanediol [5,6,7]. In the present study, the banana shoot was chosen as a novel substrate for fermentation and to date, there are no published reports on the application of the material for fermentation studies. The banana shoot finds its application in numerous value-added products such as polymer/fiber composites, packaging materials, textile fabrics, and paper sheets [8]. Alwi et al. [9] have reported that banana shoot contains 38.03% carbon, 0.42% nitrogen, 5.46% hydrogen, and also inulin.
Submerged fermentation (SmF) and solid state fermentation (SSF) methods have been reported to produce inulinase enzymes [3, 10, 11]. Lower moisture requirement, higher yield, ability to utilize natural substrates, low capital investment, reduced energy requirements, improved product recovery, and reduced environmental problems are the key advantages of SSF over submerged fermentation [12,13,14]. Fungi, bacteria, molds, and yeast are the types of microorganisms reported to produce the enzyme inulinase. Among these species, filamentous fungi are the best option for SSF due to their physiological, biochemical, and enzymological features. Furthermore, filamentous fungi have a high capacity to withstand low water activity and flourish in high osmotic stress which is a desired characteristic required for SSF [6]. The substrate for SSF should preferably be a carbon source and act as support for the growth of the microorganism. Smaller-size substrate particles are preferred due to their increased surface area required for higher productivity [14, 15]. Inulinase enzymes are generally produced using rice, millet, sugarcane, garlic waste, wheat bran, and orange rinds as substrates for SSF [16, 17].
The parameters controlling the solid state fermentation process are the type of substrate, the composition of the moistening media, inoculum size, pH, fermentation time, and incubation temperature for the production of the inulinase enzyme [2, 7]. Statistical experimental designs such as central composite design (CCD), Box-Behnken design (BBD), and Plackett–Burman design (PBD) are extensively used for optimization studies [18, 19]. These methods can provide data on the influence of independent parameters, and interactions among parameters to obtain optimum process conditions which can be highly cost-effective when implemented in practice [20]. Different downstream techniques have been considered for the purification of inulinase enzyme based on polarity, size, ligand interactions, solubility such as salt precipitation [21], solvent precipitation, gel filtration chromatography, and ion exchange [22, 23].
The current study was conducted with the objectives of isolating fungal strains capable of exhibiting inulinase activity from a novel source, optimizing process parameters of SSF for the production of inulinase enzyme utilizing the newly isolated fungal strain Nothophoma anigozanthi JAM with a novel substrate, banana shoot powder, and characterizing crude enzyme. For the isolation of fungal strain, the root soil of the Jamun tree was chosen and Czapex Dox Agar medium (CDA) was utilized for the isolation. CCD was utilized for building a second-order model and evaluating the optimal levels and interaction effects of the following parameters: the amount of substrate, volume of moistening media, inoculum volume, and fermentation time. Furthermore, the biochemical characterization of inulinases such as the effect of pH, temperature, metal ions, additives, organic solvents, as well as reaction kinetics studies was performed for evaluating its applicability in various industries. This is the first report on inulinase production using a novel strain of Nothophoma anigozanthi JAM and banana shoot powder, an inulin-containing substrate, through solid state fermentation.
2 Materials and methods
2.1 Isolation and screening of inulin-hydrolyzing fungal strain
Root soil sample from a white Jamun (Syzygium cumini) tree located in the Indrali area of Udupi (74° 44′ 31.7112″ E), India, was utilized for isolation. The collection of the soil samples was carried out in November 2020. The following protocol was utilized for the isolation of the species. Czapex Dox Agar medium (CDA) agar medium was prepared consisting of (g/L) sucrose (30), K2 HPO4 (1), NaNO3 (2), KCl (0.50), MgSO4 (0.50), FeSO4 (0.01), and agar (15). The media was maintained at a pH of 7.4 and was autoclaved at 121 °C, 15–20 psi for 20 min. The collected soil sample was subjected to serial dilution under aseptic conditions. Then, a serially diluted sample (0.1 mL) was plated on the CDA agar medium. The CDA agar plates were subjected to incubation at 30 °C for 3–4 days, thus allowing the growth of the different fungal colonies. For the primary screening, inulin agar media consisting of (g/L) inulin (6.00), Na2HPO4 (34), NH4Cl (5), NaCl (2.50), K2 HPO4 (15), and agar (15) was prepared and maintained at a pH of 6.7 [16]. Individual colonies of the species were picked from the CDA agar media and were aseptically plated on the inulin media. The plates were subjected to incubation at 32 °C for three days. The plate that showed maximum growth upon visual observation was chosen for the study.
2.2 Identification of isolated inulin-hydrolyzing fungal strain
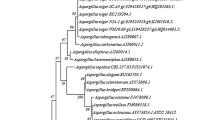
The identification of the isolated fungal strain was carried out at National Centre for Microbial Resource (NCMR), Pune, India, based on internal transcribed spacer (ITS) sequencing. The phylogenetic tree of the isolate was constructed using the neighbor-joining method.
The fungal species were stored at 4 °C in inulin agar plates. The viability of the species was ensured by subculturing periodically once in 30 days [6, 7].
2.3 Preparation of inoculum
Osmotic shock was given to the spores of the fungal species on the agar plates for 15 min using 11 mL of 0.01% sodium chloride solution to collect the spore suspension [16]. An Improved Neubauer hemocytometer (Rohem, India) was utilized to count the spores in the collected suspension. For SSF, 2 g of the substrate was inoculated with 4 mL of inoculum with a spore concentration of 6.25 × 105 spores/mL.
2.4 Substrate preparation and quantification of inulin content
The solid substrate banana plant shoots were collected from a local market located in Indrali Udupi, India. The plant shoots were thoroughly cleaned, chopped into small-sized cubes, and dried under the sun for three weeks before being blended into a fine powder. The substrate was sifted using a 300-micron sieve and stored under dry conditions using an airtight container.
To quantify the inulin content in the banana plant shoot powder, a modified method of solvent extraction was used as described elsewhere [24]. Briefly, 1 g of banana plant shoot powder was mixed with 5 mL of 0.1 M sodium acetate buffer, and the inulin extraction was carried out in the rotary shaker at 180 rpm for 24 h at 32 ℃. Then, centrifuged (Eppendorf centrifuge 5804 R) at 25 ℃ and 10,000 rpm for 10 min, and the cell-free supernatant was subjected to analysis of inulin content by the oxalic acid method as described by Simonovska [25]. The oxalic acid hydrolyzed the inulin to fructose, and the released fructose was estimated by the Dinitosalicylic acid method. The banana plant shoot extract without oxalic acid hydrolysis was taken as control. The difference in the amount of fructose measured with and without oxalic acid treatment is deemed as the inulin content in the banana plant shoot powder.
2.5 Solid state fermentation
Fermentation was carried out without the addition of the substrate inulin since the banana shoot powder contained inulin required for the growth of the microorganism. This reduces the cost of the fermentation process for the production of the enzyme; 2 g banana plant shoot powder (substrate) was mixed with 4 mL moistening media in 250 mL Erlenmeyer flasks to carry out SSF. The composition of moistening media is as follows (g/L): NH4Cl (5), Na2HPO4, (34), KH2PO4 (15), and NaCl (2.5), with a pH of 7.0. The flasks were autoclaved at 121 °C and 15 psi, for 20 min. Fungal inoculum (4 mL) with pore suspension of 6.25 × 105 spores/mL was introduced into the flasks under aseptic conditions, and the contents were mixed for uniform availability of nutrients. These flasks were subjected to incubation at 32 °C for 72 h [4, 16]. The water activity of this fermentation media was found to be 0.93, as described elsewhere [26]. The fermentation processes were carried out as per the design matrix shown in Table 2. All the experiments were conducted in triplicates.
2.6 Extraction of inulinase enzyme
The fermented matter was mixed with 40 mL of 0.1 M sodium acetate buffer at a pH of 5 and agitated thoroughly for 1 h at 180 rpm to extract the enzyme. Sterile muslin cloth was utilized for the filtration of the enzyme extract. The filtrate was subsequently centrifuged at 10,000 rpm for 10 min and 4 °C to obtain the cell-free extract. Inulinase and invertase activities were determined using the cell-free supernatant [4, 16].
2.7 Analytical methods
Reagents and chemicals of analytical grade used in this research were procured from Merck Limited, Mumbai, India, and Hi-Media Laboratories Pvt. Ltd., Maharastra, India.
2.7.1 Inulinase and invertase assay
Inulinase activity was estimated by measuring the quantum of reducing sugar fructose using the dintrosalicylic acid (DNS) method [27]. Inulin was dissolved in 0.1 M sodium acetate buffer to prepare a 20% inulin solution; 0.1 mL of the cell-free supernatant and 0.9 mL of inulin solution were incubated for 10 min, during which inulinase hydrolyzed inulin to release fructose. Fructose produced was estimated by adding 1 mL of DNS reagent followed by heating for 10 min at 90 °C in a water bath. Blank was prepared by mixing preheated cell-free supernatant and inulin solution in the same proportion. The absorbance was read at 575 nm against the blank using a spectrophotometer (Eppendorf Biospectrometer Kinetic 6136). One unit of inulinase enzyme activity is defined as the quantum of inulinase enzyme which releases 1 μmol of fructose per minute under standard assay conditions. Invertase assay is also defined as similar to inulinase assay, with the reaction mixture consisting of sucrose instead inulin [28].
2.7.2 Inulinase activity to sucrose activity (I/S ratio)
Inulinase enzyme can also hydrolyze sucrose apart from inulin, and hence, the I/S ratio was calculated. An I/S ratio of more than 10−2 indicates higher inulinase activity, while an I/S ratio of less than 10−4 indicates higher invertase production [29].
2.8 Experimental design and statistical model to optimize inulinase production using CCD
Based on preliminary studies, four parameters viz the amount of substrate, volume of moistening media, inoculum volume, and fermentation time were screened to find the optimal values for higher inulinase activity through SSF using banana shoot powder as substrate. The experimental design of CCD was created with the software Minitab 17.0 (trial version) to find the optimal process conditions and also to evaluate the interaction effects among the parameters. This design of the experiment included four parameters with five different levels (− 2, − 1, 0, + 1, + 2), as shown in Table 1. In this study, the experimental design runs comprise 31 experiments, of which 16 cube points, eight axial points, and seven center points, as given in Table 2. The statistical tool Minitab 17.0 (trial version) performs the regression analysis based on the inulinase activity data shown in Table 2 to find the impact of statistically influential parameters on the response.
A second-order polynomial was fitted consisting of the linear, quadratic, and linear interaction terms for each of the parameters (Eq. (1)).
where X is the predicted inulinase activity; pi is the ith endogenous parameter; λ0 is the intercept; λi is the coefficient of linear terms; λii is the coefficient of quadratic terms for the parameter i; and λiλj is the coefficient of interaction terms between parameters i and j. The mathematical model Eq. (2) was verified through analysis of variance (ANOVA) with F-test and R2 (coefficient of determination). Experiments were conducted at the process parameters indicated by the optimal conditions, and subsequently, inulinase activity was determined to validate the model using the strain Nothophoma anigozanthi JAM and the substrate, banana shoot powder. Mean values with standard deviation from the experiments conducted in triplicates were reported [20].
2.9 Characterization of inulinase production
2.9.1 Thin-layer chromatography
The hydrolysis reaction of the inulinase enzyme in optimized media can be analyzed using thin-layer chromatography (TLC). Five mL of cell-free supernatant extract, obtained after centrifugation, and 0.05 g of inulin were mixed thoroughly and incubated at 56 °C for 4 h. Subsequently, boiling the reaction mixture for 10 min arrested the hydrolysis reaction, thereby producing inactivated enzyme. Five mL of sodium acetate buffer was added to each of the following sugars in separate test tubes: 0.02 g fructose, 0.02 g glucose, and 0.02 g sucrose to prepare a solution of fructose (F), glucose (G), and sucrose (S). A few drops of the following were applied on a TLC plate: test samples (T), inactive inulinase (I), glucose (G), fructose (F), and sucrose (S). The TLC plate was later air dried, and to develop the chromatogram, the plate was soaked in a solution of butanol, acetone, and water (4:5:1). Air drying removed the developer solution and diphenylamine reagent consisting of diphenylamine (1%), aniline (1%), and phosphoric acid (10%) was sprayed on the TLC plate. The reaction of sugars with diphenylamine produces colors. Again, the TLC plate was then subjected to oven drying at 90 °C for 10 min [30].
2.9.2 Seliwanoff test
Aldose and ketose sugars in a sample can be distinguished by the Seliwanoff test. Ketose sugar produces cherry red or burgundy color, while aldose sugar produces no color when heated for a shorter duration. Prolonged heating is avoided since aldose sugars can also produce a red color. The inulinase and inulin-containing cell-free supernatant as test samples as well as inactivated inulinase with inulin, fructose, and glucose as the three standard controls were subjected to the Seliwanoff test. The Seliwanoff reagent and the test samples were mixed in the ratio of 1:0.5 by volume and boiled for 2 min, while the three standard control samples and Seliwanoff reagent were mixed in the proportion of 1:1 by volume before boiling for 2 min [31].
2.9.3 Osazone test
The osazone formation test is a simple test that assists in the qualitative determination and identification of sugars. Sugars such as fructose and glucose produce characteristic broom or needle-shaped crystals of osazone upon reaction with phenylhydrazine. The solutions of phenylhydrazine hydrochloride were mixed separately with 1 mL of fructose control solution, 2 mL of the inulinase, and inulin-containing extract solution, free of cells, as the test solution. Both solutions were boiled in a water bath for 30 min and cooled. Distinctive osazone crystals formed can be observed under a light microscope (Euromex Iscope, Holland) at a magnification of 40 [32].
2.10 Biochemical properties of inulinase
2.10.1 Effect of pH and stability
For the determination of the optimum pH, the crude enzyme was extracted from the fermented matter using buffers in the pH range of 2.0 to 9.0 utilizing 0.1 M KCl buffer for pH range between 2.0 and 3.0, 0.1 M sodium acetate buffer for pH range between 4.0 and 6.0, and 0.1 M sodium phosphate buffer for pH range between 7.0 and 9.0. The effect on inulinase activity due to variations in pH was determined by incubating the inulinase enzyme in the combinations of the above-stated pH and buffer system under the standard assay conditions. Percentage relative activity is calculated based on test inulinase activity divided by the control activity (highest activity obtained at the optimum pH). For pH stability, the inulinase enzyme was subjected to incubation in a buffer range of pH 2.0 to 9.0 at 4 °C for 24 h, and subsequently, the residual enzyme activity was estimated [23, 33].
2.10.2 Effect of temperature and thermal stability
The optimum temperature was determined for the inulinase activity by incubating the inulin (substrate) – inulinase (crude enzyme) mixtures at temperatures between 10 and 70 ℃ under the standard assay conditions. Similarly, thermal stability was determined in terms of inulinase activity remaining post 1 h incubation period, in the temperature range of 20 to 70 ℃ before the addition of substrate. Initial inulinase activity at the start of the experiment was considered 100% relative activity [34, 35].
2.10.3 Effect of metal ions, additives, and organic solvents
The effects of different cations of metals on the crude enzyme were determined by incubating the inulinase for 30 min in the presence of 1 mM solutions of Fe2+, Mg2+, Ca2+, K+, Zn2+, Mn2+, and Na2+ before the addition of substrate [23]. The effects of 1% (w/v) of SDS, β-mercaptoethanol, Tween-60, EDTA, and Triton x-100 on the activity of inulinase were determined under the same conditions [36, 37]. The effects of 20% (v/v) of toluene, butanol, isoamyl alcohol, methanol, and hexane on the activity of inulinase were similarly determined under the standard assay conditions [23]. Observed activity in the absence of metals, additives, and organic solvents was considered as 100% relative activity.
2.10.4 Study on enzyme reaction kinetics
The Michaelis Menten constants Km and Vmax were determined by Lineweaver–Burk plots [38]; 0.1 mL of crude supernatant was mixed with 0.9 mL of substrate (inulin) solutions at a concentration range of 10 to 70% (w/v). The inulolytic hydrolysis was allowed to proceed in the substrate-enzyme solution at 60 °C for 20 min in a water bath. Inulinase activities were measured, as described in the “Analytical methods” section [23].
3 Results and discussion
3.1 Isolation, screening, and identification of inulin-degrading fungal strain
In this study, a novel fungal strain was isolated from the root soil of the white Jamun tree (Syzygium cumini) (Udupi, India) to produce the inulinase enzyme by adopting serial dilution and spread plate method. This isolate showed its growth on an inulin agar media, as depicted in Fig. 1, which indicates that the isolate has the capability to hydrolyze substrate inulin into fructose or fructo-oligosaccharides with its inulinase activity [39,40,41]. Henceforth, this newly isolated fungal strain was selected for further study. Alongside, the isolate was identified based on internal transcribed spacer (ITS) sequencing at the National Centre for Microbial Resource (NCMR), Pune, India. The fungal isolate showed 99.55% sequence similarity with Nothophoma anigozanthi NR_135992.1 through the homology of ~ 550 bp sequence with its closest neighbor in the NCBI database (Fig. 2). The internal transcribed space (ITS) rRNA sequence data of the newly isolated species Nothophoma anigozanthi JAM has been submitted to Gen Bank (Accession no. OM103046).
This study is in good agreement with Surti and Mhatre [38], where a fungal strain was isolated from rotten garlic samples for the production of inulinase. Similarly, P. lanosocoeruleum was isolated from rhizosphere and soil environment for the production of inulinase [42].
3.2 Quantification of inulin content
The presence of inulin content in the substrate banana plant shoot powder was estimated by hydrolyzing with oxalic acid, upon which it releases fructose units [25, 27]. In this study, the free fructose content in the banana plant shoot powder was found to be 0.406 mg/gds (gds: gram dry substrate), whereas the hydrolyzed banana plant shoot powder exhibited a fructose content of 14.12 mg/gds; this indicates the presence of inulin content in the substrate (banana plant shoot powder) which is chosen for the production of inulinase through SSF with Nothophoma anigozanthi JAM. Similarly, Alwi et al. [9] reported that inulin concentration in the banana stem was 1.22 mg/mL.
3.3 Experimental design and statistical model to optimize inulinase production using CCD
The quadratic and interaction effects between the four prominent parameters (amount of substrate, volume of moistening media, inoculum volume, and fermentation time) were considered to optimize the inulinase production with banana shoot powder as substrate and strain Nothophoma anigozanthi JAM using CCD. The experimental design matrix of CCD with inulinase activity in real and coded values of parameters is shown in Table 2. The highest and lowest activity was exhibited in trial 9 (0.733 U/gds) and trial 12 (0.068 U/gds), respectively. Higher reproducibility of the experimental data is indicated by the low capriciousness of the central points (run 25, 26, 27, 28, 29, 30, and 31) (Table 2). A multiple regression mathematical model was obtained for the inulinase activity using the experimental design matrix data (Eq. (2)).
where X is the inulinase activity, U/gds; A is the amount of substrate, g; B is the volume of moistening media, mL; C is the inoculum volume, mL; and D is the fermentation time, h.
ANOVA (analysis of variance) was used for evaluating the competence of the model and its fitness (Table 3). Table 3 depicts the significant linear and quadratic terms of the amount of substrate, the volume of moistening media, inoculum volume, and fermentation time at 95% confidence (p < 0.05). Based on ANOVA, the proposed mathematical model was highly significant for the SSF of Nothophoma anigozanthi JAM with banana shoot powder as a substrate to produce inulinase, with a high F-value of 200.53 and a very low P-value of 0.000. In this study, the F-values of the model (200.53) are much higher than “lack of fit” (5.31), which indicates that a better fit of data to the proposed mathematical model (Table 3), and can be ascertained by the determination coefficient (R2). The value of R2 was 0.9943, indicating that 99.43% of the total variation in the inulinase production was ascribed to the parameters. Furthermore, the rational agreement between the predicted R2 (0.97) and adjusted R2 (0.9894) of the mathematical model (Eq. (2)) was observed.
Figure 3 depicts the three-dimensional surface plots, which explain the interaction effects among the parameters on inulinase production. The degree of interactions among the parameters can be evaluated based on the outline of the surface plot. A convex type of outline between the parameter indicates the presence of interaction between the parameters, whereas a concave shape indicates less or no interaction [43]. The effect of interaction between two parameters is shown with other parameters held constant at their central level (0). A convex type of outline between the parameters, A × B, A × D, B × C, B × D, and C × D, specified the substantial interaction effect at its optimal conditions for improved inulinase activity except for A × C. The equivalent interpretations are statistically made known in Table 3.
Response plots presenting the interaction effects between two parameters on the production of inulinase with SSF and Nothophoma anigozanthi JAM when other parameters are held at a constant level. (a) amount of substrate, A, and volume of moistening media, B; (b) amount of substrate, A, and inoculum volume, C; (c) amount of substrate, A, and fermentation time, D; (d) volume of moistening media, B, and inoculum volume, C; (e) volume of moistening media, B, and fermentation time, D; and (f) inoculum volume, C, and fermentation time, D
The response optimizer tool of the MINITAB 17 (free trial version) was utilized for predicting the optimal level of parameters for enhanced production of inulinase, and the values of their optimum levels are as follows: the amount of substrate (banana shoot powder) = 2.0 g, the volume of moistening media = 5.0 mL, inoculum volume = 5.0 mL, and fermentation time = 96 h, and an inulinase activity of 2.047 U/gds was predicted at the above-mentioned optimum condition (Table 4).
Das et al. [16] optimized inulinase production using a similar approach and reported the highest inulinase activity of 2.45 U/gds from Penicillium amphipolaria strain at 96 h of fermentation time under optimal conditions using hardy sugarcane stem as substrate. Similarly, Sheng et al. [44] reported a maximum inulinase activity from marine yeast strain Cryptococcus aureus G7a and substrates wheat bran and rice husk at a fermentation time of 120 h. Yazici et al. [45] optimized the process parameters for the production of inulinase from onion peel and stevia roots and reported the highest activity in 120 h. In this study, the newly isolated strain Nothophoma anigozanthi JAM produced inulinase in a reduced fermentation period of 96 h by degrading inulin present in the crude substrate banana plant shoot powder, whereas other strains reported higher fermentation periods than 96 h described elsewhere [35, 44].
To validate the proposed mathematical model (Eq. (2)), experimental trials were carried out at the optimized levels in triplicates, as shown in Table 4. The predicted and experimental inulinase activities were 2.109 U/gds and 2.578 ± 0.166 U/gds, respectively (Table 4); this low variability signifies the adequacy of the model. The statistical approach CCD led to an increase in inulinase activity from 0.322 ± 0.025 U/gds (unoptimized medium) to 2.578 ± 0.166 U/gds (optimized medium), with substrate banana shoot powder in SSF by Nothophoma anigozanthi JAM.
3.4 Characterization of inulinase production
3.4.1 Thin-layer chromatography
Standards used in the TLC viz. fructose (F), glucose (G), and sucrose (S) developed brown-colored spots, as observed in Fig. 4, while the control (I) containing both substrate inulin and deactivated enzyme inulinase produced very weak spots. The three test samples (T1 to T3) produced single-colored spots confirming the presence of fructose from the hydrolysis reaction. The exoenzyme inulinase hydrolyzed inulin to generate fructose which in turn produced brown spots in the developed TLC plate. Das et al. [16] could observe reddish brown spots for the test samples resembling the spots for fructose and glucose standards in the TLC plate and proved P. amphipolaria inulinase to have an Exo-inulolytic nature. Garuba et al. [34] also obtained similar findings while analyzing the hydrolysis products of Aspergillus tamarii inulinase using TLC with their test samples and the standards showing similar spots.
3.4.2 Seliwanoff test
Aldose and ketose sugars in a cell-free extract can be distinguished by Seliwanoff’s test. Fructose solution (F) and the cell-free extract (E) produced red color on heating with Seliwanoff solution, while glucose solution (G) and the control (D) containing thermally deactivated enzyme did not generate a color (Fig. 5). Hence, this study validates the production of inulinase upon hydrolysis of inulin present in banana plant shoot and releasing the product fructose [31].
3.4.3 Osazone formation test
The hydrolysis of inulin by inulinase release fructose, which was further validated through the osazone formation test. Needle and broom-shaped crystals were observed under a light microscope with both fructose standard and synthesized fructose (Fig. 6). Kamble et al. [46] and Das et al. [16] reported the formation of needle and broom-shaped osazone crystals like that of fructose standards while performing osazone test with their crude inulinase enzymes obtained from Arthrobacter mysorens and P. amphipolaria, respectively.
3.5 Biochemical properties of inulinase
3.5.1 Effect of pH and stability
Figure 7a depicts that the crude inulinase enzyme was very active in the pH range of 4–6 and the peak activity and stability were observed at a pH of 5.0. The reaction temperature maintained during this study was 30 °C, i.e., an assay condition. An optimum pH of 7.0 was reported for inulinase from Chrysosporium pannorum [47]. Inulinase from Fusarium oxysporum had an optimum pH of 6.2 [2]. An optimum pH of 6.0 was reported for inulinase from Aspergillus fumigatus [48]. Pessoni et al. [49] obtained an optimum pH of 6.0 from Penicillium janczewskii inulinase. Inulinase from Pichia guilliermondii had an optimum pH of 6.0 [50]. de Oliveira et al. [17] reported that crude inulinase from A. terreus showed maximum activity at pH of 4.0. The optimum pH of enzymes isolated from different sources generally varies, attributed probably due to the composition variations in the molecular structure. This affects the enzyme ionization in a solution.
3.5.2 Effect of temperature and thermal stability
Inulinase activity gradually increased from 10 °C, reaching a peak at 60 °C, and subsequently, a drop in the activity was observed at 70 °C, as depicted in Fig. 7b. An optimized pH of 5.0 was maintained during this study. Most of the fungal inulinases have been reported to have the best operational temperature range of 45–55 °C. However, some studies have reported 60 ℃ as the optimum temperature, such as inulinase from A.terreus [17], Aspergillus fumigatus [48], and Pichia guilliermondii [50].
The maximum stability of the inulinase enzyme was observed at a storage temperature of 30 °C (Fig. 7b). The thermal stability of the enzyme steadily decreased from 30 to 60 °C, and a drop in the activity was observed at 70 °C, as was observed previously in the case of the effect of temperature on inulinase activity. Inulinases from fungal species such as Aspergillus ochraceus have thermal stability of 60 ℃ [51]. Inulinase from Cryptococcus aureus is thermally stable until 65 ℃ [44]. Inulinase from Aspergillus ficuum has thermal stability in the range of 50–60 ℃ [52]. Yuan et al. [53] reported that inulinase from Candida kutaonensis has a thermal stability range of 45–60 ℃.
3.5.3 Influence of metal ions, additives, and organic solvents on inulinase activity
Different metal ions were tested at various concentrations for effect on crude inulinase activity. It was observed that the enzyme activity was completely inhibited and no activity was observed at metal ion concentrations of 2 mM and above. Results of the metal ion on enzyme activity are depicted in Fig. 8a. Metal ions Zn++ and K+ completely inhibited the enzyme activity, while Ca++ displayed no effect on the activity. Also, it must be understood that several factors such as enzyme protein conformation and source of isolation can control how different metal ions affect the activity of inulinase or any enzyme. For example, Singh et al. [6] reported that K+ has no effect and Ca++ has a positive effect on the activity of Penicillium oxalicum BGPUP-4 inulinase enzyme. Rawat et al. [23] observed the presence of Ca++ and Fe+++ to elicit a rather positive effect on the activity of Aspergillus fumigatus inulinase and Hg++ to strongly inhibit the activity.
Figure 8b depicts that Triton × 100 had a beneficial effect and improved the activity of the inulinase while EDTA, SDS, Tween 60, and β mercaptoethanol reduced the activity of the enzyme. Nguyen et al. [37] also reported that the addition of Triton × 100 enhanced the activity of Thermomyces lanuginosus IMI 140,524 inulinase. Lu et al. [54] could observe that EDTA and SDS decreased the inulinase activity of Nocardiopsis sp. DN-K15 at different concentrations. Inulinase produced by Paecilomyces variotii was found to be negatively affected by the presence of EDTA and β mercaptoethanol [36]. Rawat et al. [23] also confirmed the inhibitory effects of EDTA on the activity of Aspergillus fumigatus inulinase.
Most organic solvents are known to deactivate various enzymes [55]. However, the activity of the inulinase enzyme from Nothophoma anigozanthi JAM was enhanced by the solvent isoamyl alcohol and the remaining solvents suppressed the activity of the enzyme, as shown in Fig. 8c. Rawat et al. [23] also have studied the effect of various solvents such as methanol, chloroform, butanol, toluene, ethanol, hexane, and glycerol on the activity of inulinase produced by Aspergillus fumigatus and found toluene to possess a stimulatory effect on the inulinase activity to some degree.
3.5.4 Study on enzyme reaction kinetics
Km and Vmax are two important parameters that determine an enzyme’s kinetic behavior as a function of substrate concentration [56, 57]. The kinetic parameters were determined from the Lineweaver–Burk plot. The Vmax and Km are estimated to be 0.333 μmol/L.min and 205 mM, respectively (Fig. 8d). The result of the Lineweaver–Burk plot revealed that the crude inulinase enzyme had a higher affinity for inulin as well as a faster reaction velocity. The lower value of Km indicates that the enzyme has reached maximum catalytic efficiency at a lower substrate concentration. Rawat et al. [23] obtained a Km and Vmax value of 5.88 mM and 1.66 µM/min, respectively, for the purified exo-inulinase isolated from Aspergillus fumigatus. Germec et al. [58] reported an experimental Vmax value of 1055.55 U/mL and Km of 16.13 mg/mL determined using a Lineweaver–Burk plot when inulin was used as a substrate for Aspergillus niger inulinase.
4 Conclusion
In this study, the inulinase enzyme was produced from the novel fungal strain Nothophoma anigozanthi JAM, isolated from the Jamun tree soil, with novel substrate banana plant shoot powder through SSF. The study of the statistical design of experiments for inulinase production has led to an eightfold increase (2.578 ± 0.166 U/gds) when compared to unoptimized conditions. Banana plant shoot powder was found to be the best alternative source of the synthetic and expensive inulin substrate; it may also find its application in biofuel sectors. TLC, osazone, and Seliwanoff tests validated the production of inulinase by Nothophoma anigozanthi JAM. The inulinase activity was increased in the presence of Ca+2 ions (co-factor), isoamyl alcohol (organic solvent), and Triton × 100 (additives) and also was active between the range 40 to 60 °C and pH 5.0. These properties of inulinase may be beneficial for the food and beverage industries. Further studies can be carried out to purify the inulinase with different techniques such as precipitation, dialysis, size exclusion chromatography, and ion exchange chromatography to enhance the enzymatic activity. Applicational studies of the enzyme in regard to theproduction of biofuels, organic acids, and single-cell proteins are also recommended.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding authors upon reasonable request.
References
Bender JP, Mazutti MA, Treichel H, Di Luccio M (2006) Inulinase production by Kluyveromyces marxianus NRRL Y-7571 using solid state fermentation. Appl Biochem Biotechnol 32:951–958. https://doi.org/10.1385/abab:132:1:951
Camelia NB, Gabriela B (2011) Inulinases – a versatile tool for biotechnology. Inn Rom Food Biotech 9:1–11. https://www.gup.ugal.ro/ugaljournals/index.php/IFRB/article/view/3373. Accessed 02 May 2022
Guerrero-Urrutia C, Volke-Sepulveda T, Figueroa-Martinez F, Favela-Torres E (2021) Solid-state fermentation enhances inulinase and invertase production by Aspergillus brasiliensis. Process Biochem 108:169–175. https://doi.org/10.1016/j.procbio.2021.06.014
Das D, Bhat MR, Selvaraj R (2019) Review of inulinase production using solid-state fermentation. Ann Microbiol 69(3):201–209. https://doi.org/10.1007/s13213-019-1436-5
Singh RS, Singh T, Hassan M, Kennedy JF (2020) Updates on inulinases: structural aspects and biotechnological applications. Int J Biol Macromol 164:193–210. https://doi.org/10.1016/j.ijbiomac.2020.07.078
Singh RS, Chauhan K (2018) Production, purification, characterization, and applications of fungal inulinases. Curr Biotechnol 7(3):242–260. https://doi.org/10.2174/2211550105666160512142330
Singh RS, Chauhan K, Kennedy JF (2019) Fructose production from inulin using fungal inulinase immobilized on 3-aminopropyl-triethoxysilane functionalized multiwalled carbon nanotubes. Int J Biol Macromol 125:41–52. https://doi.org/10.1016/j.ijbiomac.2018.11.281
Vigneswaran C, Pavithra V, Gayathri V, Mythili K (2015) Banana fiber: scope and value added product development. J Text Appar Tech Manag 9(2). https://jtatm.textiles.ncsu.edu/index.php/JTATM/article/viewFile/6825/3586. Accessed 20 Dec 2022
Alwi H, Idris J, Musa M, Ku Hamid KH (2013) A preliminary study of banana stem juice as a plant-based coagulant for treatment of spent coolant wastewater. J Chem Article ID 165057. https://doi.org/10.1155/2013/165057
Thomas L, Larroche C, Pandey A (2013) Current developments in solid-state fermentation. Biochem Eng J 81:146–161. https://doi.org/10.1016/j.bej.2013.10.013
El Sheikha AF, Ray RC (2017) Potential impacts of bioprocessing of sweet potato. Crit Rev Food Sci Nutr 57(3):455–471
Singhania RR, Patel AK, Soccol CR, Pandey A (2009) Recent advances in solid-state fermentation. Biochem Eng J 44(1):13–18. https://doi.org/10.1016/j.bej.2008.10.019
Pandey A, Selvakumar P, Soccol CR, Nigam P (1999) Solid state fermentation for the production of industrial enzymes. Curr Sci 149–162. http://www.jstor.org/stable/24102923. Accessed 18 Apr 2022
El Sheikha AF, Ramesh CR (2022) Bioprocessing of horticultural wastes by solid-state fermentation into value-added/innovative bioproducts: a review. Food Rev Int. https://doi.org/10.1080/87559129.2021.2004161
Prabhu G, Bhat D, Bhat RM, Selvaraj S (2022) A critical look at bioproducts co-cultured under solid state fermentation and their challenges and industrial applications. Waste Biomass Valoriz 13:3095–3111. https://doi.org/10.1007/s12649-022-01721-0
Das D, Bhat MR, Selvaraj R (2020) Optimization of inulinase production by a newly isolated Penicillium Amphipolaria strain using solid-state fermentation of hardy sugarcane stems. Biocatal Agric Biotechnol 30:101875. https://doi.org/10.1016/j.bcab.2020.101875
de Oliveira RL, da Silva SP, Converti A, Porto TS (2022) Production, biochemical characterization, and kinetic/thermodynamic study of inulinase from Aspergillus terreus URM4658. Molecules 27(19):6418. https://doi.org/10.3390/molecules27196418
Mukesh K, Shiny R, Vikas B, Raj KS (2015) Optimization of tannase production by a novel Klebsiella pneumonia KP715242 using central composite design. Biotechnol Rep 7:128–134. https://doi.org/10.1016/j.btre.2015.06.002
Yupanqui-Mendoza SL, de Arruda PV, da Silva GMC (2022) Statistical sequential optimization of process parameters for inulinase production by Kluyveromyces marxianus ATCC 36907 in solid-state fermentation using beer residue. Biocat Agri Biotech 39:102252. https://doi.org/10.1016/j.bcab.2021.102252
Subbalaxmi S, Vytla RM (2016) Process optimization for tannase production by Bacillus gottheilii M2S2 on inert polyurethane foam support. Biocatal Agri Biotechnol 7:48–55. https://doi.org/10.1016/j.bcab.2016.05.004
Golunski S, Silva MF, Marques CT, Rosseto V, Kaizer RR, Mossi AJ, Rigo D, Dallago RM, Di Luccio M, Treichel H (2017) Purification of inulinases by changing the ionic strength of the medium and precipitation with alcohols. An Acad Bras Cienc 89:57–63. https://doi.org/10.1590/0001-3765201720160367
Housseiny MM (2014) Production of an endo-inulinase from Aspergillus niger AUMC 9375, by solid state fermentation of agricultural wastes, with purification and characterization of the free and immobilized enzyme. J Microbiol 52(5):389–398. https://doi.org/10.1007/s12275-014-3561-y
Rawat HK, Soni H, Suryawanshi RK et al (2021) Exo-inulinase production from Aspergillus fumigatus NFCCI 2426: purification, characterization, and immobilization for continuous fructose production. J Food Sci 86(5):1778–1790. https://doi.org/10.1111/1750-3841.15681
Lasano NF, Ramli NS, Hamid AH, Karim R, Dek MS, Shukri R (2019) Effects of different extraction solvents on polyphenols and antioxidant capacity of peel, pulp and seed kernel of kuini (Mangifera odorata). Orient Pharm Exp Med 19(3):277–286. https://doi.org/10.1007/s13596-019-00383-z
Simonovska B (2000) Determination of inulin in foods. J AOAC Int 83(3):675–678. https://doi.org/10.1093/jaoac/83.3.675
Lu W, Li D, Wu Y (2003) Influence of water activity and temperature on xylanase biosynthesis in pilot-scale solid-state fermentation by Aspergillus sulphureus. Enzyme and Microb Technol 32(2):305–311. https://doi.org/10.1016/S0141-0229(02)00292-2
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. J Anal Chem 31:426–428. https://doi.org/10.1021/ac60147a030
Dinarvand M, Rezaee M, Masomian M et al (2013) Effect of C/N ratio and media optimization through response surface methodology on simultaneous productions of intra-and extracellular inulinase and invertase from Aspergillus niger ATCC 20611. BioMed Res Int. https://doi.org/10.1155/2013/508968
Bonciu CN, Constantin O, Bahrim GE (2011) Screening of biotechnological parameters for fructofuranosidases production by a newly isolated fungal strain using plackett-burman design. Not Bot Horti Agrobot Cluj Napoca 39(2):271–275. https://doi.org/10.15835/nbha3926323
Singh RS, Chauhan K (2017) Inulinase production from a new inulinase producer, Penicillium oxalicum BGPUP-4. Biocatal Agri Biotechnol 9:1–10. https://doi.org/10.1016/j.bcab.2016.10.012
Sanchez-Viesca F, Gómez R (2019) On the mechanism of the Froehde reaction. World J Org Chem 7(1):1–4. https://doi.org/10.12691/wjoc-7-1-3
Shah T, Modi N (2016) Utility of osazone test to identify sugars. J Med Sci Clin Res 04: 14361–14365. https://doi.org/10.18535/jmscr/v4i12.14
Temkov M, Strinska H, Dobrev G, Velickova E, Muresan V, Krastanov A (2018) Purification of bacterial inulinase in aqueous two-phase systems. Eng Life Sci 18(11):840–850. https://doi.org/10.1002/elsc.201700174
Garuba EO, Onilude AA (2020) Exo-Inulinase Production by a catabolite repression-resistant mutant Thermophilic Aspergillus tamarii-U4 in solid state fermentation. Biotechnol J Int 24:21–31. https://doi.org/10.9734/bji/2020/v24i430110
Trivedi S, Divecha J, Shah A (2012) Optimization of inulinase production by a newly isolated Aspergillus tubingensis CR16 using low cost substrates. Carbohydr Polym 90(1):483–490. https://doi.org/10.1016/j.carbpol.2012.05.068
Gao Z, Wang X, Lu N et al (2019) Enzymatic properties of thermoacidophilic inulinase from Paecilomyces variotii to be used in production of high fructose syrup Sh. Kexue/Food Sci 40(8):94–101
Nguyen DQ, Sujtó NM, Bujna E, Hoschke Á, Rezessyné SJ (2013) Effects of medium composition and process parameters on the production of extracellular inulinase by Thermomyces lanuginosus. Food Technol Biotechnol 51(1):36–44
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56(3):658–666. https://doi.org/10.1021/ja01318a036
Surti A, Mhatre S (2021) Optimization of inulinase production by a fungal species isolated from rotten garlic samples. Appl Biotechnol Rep 8(2):164–171. https://doi.org/10.30491/jabr.2020.238953.1253
Singh RS, Chauhan K, Kennedy JF (2017) A panorama of bacterial inulinases: production, purification, characterization and industrial applications. Int J Biol Macromol 96:312–322. https://doi.org/10.1016/j.ijbiomac.2016.12.004
Rawat HK, Ganaie MA, Kango N (2015) Production of inulinase, fructosyltransferase and sucrase from fungi on low-value inulin-rich substrates and their use in generation of fructose and fructo-oligosaccharides. Antonie Van Leeuwenhoek 107(3):799–811. https://doi.org/10.1007/s10482-014-0373-3
Corrado I, Cascelli N, Ntasi G, Birolo L, Sannia G, Pezzella C (2021) Optimization of inulin hydrolysis by Penicillium lanosocoeruleum inulinases and efficient conversion into polyhydroxyalkanoates. Front Bioeng Biotech 108. https://doi.org/10.3389/fbioe.2021.616908
Selvaraj S, Vytla RM, Vijay GS, Natarajan K (2019) Modeling and optimization of tannase production with Triphala in packed bed reactor by response surface methodology, genetic algorithm, and artificial neural network. 3 Biotech 9(7):1–12. https://doi.org/10.1007/s13205-019-1763-z
Sheng J, Chi Z, Yan K, Wang X, Gong F, Li J (2009) Use of response surface methodology for optimizing process parameters for high inulinase production by the marine yeast Cryptococcus aureus G7a in solid-state fermentation and hydrolysis of inulin. Bioprocess Biosyst Eng 32(3):333–339. https://doi.org/10.1007/s00449-008-0252-2
Yazici SO, Sahin S, Biyik HH, Geroglu Y, Ozmen I (2021) Optimization of fermentation parameters for high-activity inulinase production and purification from Rhizopus oryzae by Plackett-Burman and Box-Behnken. J Food Sci Tech 58(2):739–751
Kamble PP, Kore MV, Patil SA, Jadhav JP, Attar YC (2018) Statistical optimization of process parameters for inulinase production from Tithonia weed by Arthrobacter mysorens strain no.1. J Microbiol Methods 149:55–66. https://doi.org/10.1016/j.mimet.2018.04.019
**ao R, Tanida M, Takao S (1989) Purification and some properties of endoinulinase from Chrysosporium pannorum. J Ferment Bioeng 67(4):244–248. https://doi.org/10.1016/0922-338X(89)90226-2
Gill PK, Manhas RK, Singh P (2006) Purification and properties of a heat-stable exoinulinase isoform from Aspergillus fumigatus. Bioresour Technol 97(7):894–902. https://doi.org/10.1016/j.biortech.2005.04.034
Pessoni RA, Braga MR, Figueiredo-Ribeiro RDCL (2007) Purification and properties of exo-inulinases from Penicillium janczewskii growing on distinct carbon sources. Mycologia 99(4):493–503. https://doi.org/10.1080/15572536.2007.11832543
Gong F, Zhang T, Chi Z, Sheng J, Li J, Wang X (2008) Purification and characterization of extracellular inulinase from a marine yeast Pichia guilliermondii and inulin hydrolysis by the purified inulinase. Biotechnol Bioprocess Eng 13(5):533–539. https://doi.org/10.1007/s12257-007-0177-7
Guimarães LHS, Terenzi HF, de Moraes MDLT, Jorge JA (2007) Production and characterization of a thermostable extracellular β-D-fructofuranosidase produced by Aspergillus ochraceus with agroindustrial residues as carbon sources. Enzyme Microb Technol 42(1):52–57. https://doi.org/10.1016/j.enzmictec.2007.07.021
Chen HQ, Chen XM, Li Y, Wang J, ** ZY, Xu XM et al (2009) Purification and characterisation of exo-and endo-inulinase from Aspergillus ficuum JNSP5-06. Food Chem 115(4):1206–1212. https://doi.org/10.1016/j.foodchem.2009.01.067
Yuan B, Hu N, Sun J, Wang SA, Li FL (2012) Purification and characterization of a novel extracellular inulinase from a new yeast species Candida kutaonensis sp. nov. KRF1T. Appl Microbiol Biotechnol 96(6):1517–1526. https://doi.org/10.1007/s00253-012-4108-y
Lu WD, Li AX, Guo QL (2014) Production of novel alkalitolerant and thermostable inulinase from marine actinomycete Nocardiopsis sp. DN-K15 and inulin hydrolysis by the enzyme. Ann Microbiol 64(2):441–449. https://doi.org/10.1007/s13213-013-0674-1
Laane C, Boeren S, Vos K, Veeger C (1987) Rules for optimization of biocatalysis in organic solvents. Biotechnol Bioeng 30(1):81–87. https://doi.org/10.1002/bit.260300112
Roskoski R (2015) Michaelis-Menten kinetics. Reference Module in Biomed Sci Elsevier Inc. https://doi.org/10.1016/B978-0-12-801238-3.05143-6
Chandra MRGS, Madakka M (2018) Recent developments in applied microbiology and biochemistry. Comparative biochemistry and kinetics of microbial ignocellulolytic enzymes. In: Recent developments in applied microbiology and biochemistry, Academic press. https://doi.org/10.1016/B978-0-12-816328-3.00011-8
Germec M, Turhan I (2021) Predicting the experimental data of the substrate specificity of Aspergillus niger inulinase using mathematical models, estimating kinetic constants in the Michaelis–Menten equation, and sensitivity analysis. Biomass Convers Biorefin 1–12. https://doi.org/10.1007/s13399-021-01830-1
Acknowledgements
The authors of this study are thankful to the Vision Group on Science and Technology (VGST), Government of Karnataka, India, for sponsoring the work (GRD 773) and Manipal Institute of Technology, MAHE, for providing the facilities to conduct the research work.
Funding
Open access funding provided by Manipal Academy of Higher Education, Manipal Open access funding is provided by the Manipal Academy of Higher Education (MAHE), Manipal. This study was supported by a fund provided by the Vision Group on Science and Technology (VGST), Government of Karnataka, India (GRD 773).
Author information
Authors and Affiliations
Contributions
Material preparation, experimentation, data collection, and validation of this work were performed by Nisarga Tippanavar, Divya Bhat, Orline Rebello, and Girisa Prabhu. The study was supervised, and the draft manuscript was written and revised by Subbalaxmi Selvaraj and Ramananda M Bhat. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethical approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tippanavar, N., Bhat, D., Rebello, O. et al. Bio-utilization of agricultural residue banana plant shoot through solid state fermentation for production of inulinase using newly isolated Nothophoma anigozanthi JAM. Biomass Conv. Bioref. (2023). https://doi.org/10.1007/s13399-023-03835-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13399-023-03835-4