Abstract
(CCG)n•(CGG)n trinucleotide repeats have been found to be associated with fragile X syndrome, the most widespread inherited cause of mental retardation in humans. The (CCG)n•(CGG)n repeats adopt i-motif conformations that are preferentially stabilized by base-pairing interactions of noncanonical proton-bound dimers of cytosine (C+•C). Halogenated cytosine residues are one form of DNA damage that may be important in altering the structure and stability of DNA or DNA–protein interactions and, hence, regulate gene expression. Previously, we investigated the effects of 5-halogenation and 1-methylation of cytosine on the base-pairing energies (BPEs) using threshold collision-induced dissociation (TCID) techniques. In the present study, we extend our work to include proton-bound homo- and heterodimers of cytosine, 1-methyl-5-fluorocytosine, and 1-methyl-5-bromocytosine. All modifications examined here are found to produce a decrease in the BPEs. However, the BPEs of all of the proton-bound dimers examined significantly exceed those of Watson-Crick G•C, neutral C•C base pairs, and various methylated variants such that DNA i-motif conformations should still be preserved in the presence of these modifications. The proton affinities (PAs) of the halogenated cytosines are also obtained from the experimental data by competitive analysis of the primary dissociation pathways that occur in parallel for the proton-bound heterodimers. 5-Halogenation leads to a decrease in the N3 PA of cytosine, whereas 1-methylation leads to an increase in the N3 PA. Thus, the 1-methyl-5-halocytosines exhibit PAs that are intermediate.

ᅟ
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cytosine methylation in human cells is an important element of cellular coordination of gene expression [1]. Generally, in the context of gene promoters, hypomethylation of cytosine residues is associated with active, constitutively expressed genes, whereas hypermethylation of cytosine residues is associated with silenced genes [2]. Indeed, cytosine methylation is a major contributor to the generation of disease-causing germline mutations [3] and somatic mutations that cause cancer [4]. Previous studies have shown that Fragile X syndrome is caused by 5-hypermethylation of cytosine, which is the most widespread inherited cause of mental retardation in humans, and results in intellectual disabilities and physical deformities [5, 6]. It has previously been shown that the expansion of (CCG)n•(CGG)n trinucleotide repeats beyond 230 trinucleotides leads to marked methylation of both the CGG repeats and the fragile X mental retardation 1 (FMR1) gene promoter on the X chromosome, resulting in a failure to express the fragile X mental retardation protein (FMRP), which is required for normal neural development [7–9]. The proposed structure of single (CCG)n and (CCG)n•(CGG)n strands involve noncanonical DNA structures such as the DNA i-motif that may be the cause of the disease [10, 11]. The DNA i-motif conformation was first discovered in 1993 by Gehring and coworkers [12] and later in (CCG)n•(CGG)n trinucleotide repeats [13]. The secondary structure of the DNA i-motif is a four-stranded structure consisting of parallel-stranded DNA duplexes zipped together in an anti-parallel orientation by intercalated proton-bound dimers of cytosine (C+•C) [12]. Since the discovery of the DNA i-motif, the biological roles of i-motif structures as well as their potential applications in nanotechnology have drawn great attention. Recently, studies have shown that the structure of the i-motif is conserved in the gas phase when electrospray ionization (ESI) is used as the ionization technique [14], indicating that gas-phase studies may be used to provide insight into solution-phase structure and function.
Because cytosine methylation is a critical player in the epigenetic control of gene expression, it is not surprising that alterations or perturbations of cytosine methylation patterns have been implicated in the development of human cancer. Indeed, a substantial and growing number of human genes display altered methylation status in human tumors [15]. One form of DNA damage that may prove particularly important in altering methylation status is halogenated cytosine residues. Previous studies have shown that halogen atoms, particularly bromine, can mimic the behavior of a methyl group in DNA–protein interactions [16, 17]. For example, oligonucleotides containing 5-bromocytosine have been found to exhibit similar binding affinities for methyl-CpG binding proteins that selectively bind methylated DNA [18]. Recent studies have determined that 5-chlorocytosine and 5-bromocytosine are formed through endogenous processes in areas of tissue inflammation, which have long been associated with cancer and, thus, suggesting that halogenation of nucleic acids can also be a significant form of DNA damage in living organisms [19–21]. In addition, 5-chlorocytosine and 5-bromocytosine can be potential sources of 5-chlorouracil and 5-bromouracil, two common known mutagens [22, 23]. The smaller fluorine substituent is a mimic of hydrogen with respect to size. However, the electron-withdrawing capacity of fluorine distinguishes it from hydrogen in its influence on enzymatic reactions. For instance, 5-fluorocytosine residues of oligonucleotides covalently bind DNA methyltransferases from both bacteria and mammals [24–26].
The structure of the proton-bound dimer of cytosine has been shown to be conserved upon 1-methylation or 5-halogentation via infrared multiple photon dissociation (IRMPD) action spectroscopy techniques [27, 28]. Theoretical calculations at the B3LYP/def2-TZVPPD level of theory predict the base-pairing energy (BPE) of the proton-bound dimer of cytosine (C+•C) as 170.1 kJ/mol, 79% and 163% greater than those of the canonical G•C and neutral C•C base pairs, respectively, indicating that the stronger base-pairing interactions in the C+•C homodimer are likely the major factor that stabilizes noncanonical DNA i-motif conformations [29–31]. Given the important biological roles that DNA i-motif conformations may play in several human diseases and cancer, including lung carcinoma [32], breast carcinoma [33], and Burkitt’s lymphomas [34], a comprehensive study is needed to determine the influence of halogenation on the strengths of base-pairing interactions in proton-bound cytosine dimers. A previous X-ray crystallography study suggests that cytosine protonation, required for the formation of the proton-bound C+•C dimers, is affected by a decrease of pKa of cytosine upon halogenation [35]. Quantitative determination of the BPEs of proton-bound homo- and heterodimers of C, 1MeC, and 5xC, where x = F, Br, and I, was performed using threshold collision-induced dissociation techniques (TCID) [29–31], It was determined that 1-methylation exerts very little influence on the BPE, whereas theory suggests that 1-methylation should lead to a decrease in the BPE. Halogenation of one or both cytosine residues at the 5-position decreases the BPE and should therefore destabilize DNA i-motif conformations. In the present work, we expand the complexes of interest to include 1-methyl-5-fluorocytosine (1Me5FC) and 1-methyl-5-bromocytosine (1Me5BrC), such that the effects of the 5-halogen substituents on the BPE in the presence of 1-methylation can be elucidated as well. The structures of cytosine and the 1-methyl-5-halocytosines as well as those of the proton-bound homodimers of the 1-methyl-5-halocytosines and the proton-bound heterodimers of the 1-methyl-5-halocytosines with cytosine are shown in Scheme 1. The BPEs of these proton-bound dimers generated by ESI are determined using TCID techniques in a guided ion beam tandem mass spectrometer. Relative N3 proton affinities (PAs) of the 1-methyl-5-halocytosines are also extracted from the experimental data from competitive analyses of the two primary dissociation pathways that occur in parallel for the proton-bound heterodimers of cytosine and the 1-methyl-5-halocytosines. Absolute N3 PAs of the 1-methyl-5-halocytosines are also obtained using the relative PAs determined here and the PA of C [29, 36–38] reported in the literature. The measured values are compared with theoretical results calculated at the B3LYP and MP2(full) levels of theory to evaluate the ability of each level of theory for predicting accurate energetics.
Experimental and Computation
General Procedures
TCID of four proton-bound dimers, (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC), is studied using a guided ion beam tandem mass spectrometer that has been described in detail previously [39]. The proton-bound dimers are generated by ESI from solutions containing 0.5–1 mM of the 1-methyl-5-halogenated cytosines and/or cytosine and 1% (v/v) acetic acid in an approximately 50%:50% MeOH:H2O mixture. The proton-bound dimer ions are desolvated, focused, and thermalized in a radio frequency (rf) ion funnel and hexapole ion guide collision cell interface. The thermalized ions emanating from the hexapole ion guide are extracted, accelerated, and focused into a magnetic sector momentum analyzer for mass analysis. Mass-selected ions are decelerated to a desired kinetic energy and focused into a rf octopole ion beam guide that acts as an efficient radial trap [40–42] for ions such that scattered reactant and products ions are not lost as they drift toward the end of the octopole. The octopole passes through a static gas cell where the proton-bound dimer ions undergo collision-induced dissociation (CID) with Xe [43–45] under nominally single collision conditions, ~0.05–0.10 mTorr. Product and unreacted proton-bound dimer ions drift to the end of the octopole, where they are focused into a quadrupole mass filter for mass analysis. The ions are detected using a secondary electron scintillation (Daly) detector and standard pulse counting techniques. Cytosine was purchased from Alfa Aesar (Ward Hill, MA, USA); the 1-methyl-5-halocytosines were synthesized in the laboratory of Professor T. H. Morton of the University of California, Riverside.
Theoretical Calculations
The stable low-energy tautomeric conformations of C and H+(C) have previously been examined as described in detail elsewhere [30]. In the present study, geometry optimizations and frequency analyses of the low-energy tautomeric conformations of 1Me5XC, H+(1Me5XC), where 1Me5XC = 1Me5FC and 1Me5BrC, and four proton-bound dimers including (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC), were performed using Gaussian 09 [46] at the B3LYP/6-31G*, B3LYP/def2-TZVPPD, and MP2(full)/6-31G* levels of theory. The def2-TZVPPD basis set [47] is a balanced basis set on all atoms at the triple zeta level and includes polarization and diffuse functions. The def2-TZVPPD basis set was obtained from the EMSL basis set exchange library [48, 49]. The polarizabilities of the neutral nucleobases required for threshold analyses were calculated at the PBE1PBE/6-311+G(2d,2p) level of theory, which has been shown to provide polarizabilities that exhibit better agreement with experimental values than the B3LYP functional employed here for structures and energetics [50]. Relaxed potential energy surface (PES) scans were performed at the B3LYP/6-31G* level of theory to provide candidate structures for the transition states (TSs) for dissociation of the ground-state conformations of the proton-bound dimers to produce ground-state O2-protonated, I + , and neutral, i, products. The actual TSs were obtained using the quasi-synchronous transit method, QST3 [51] at the B3LYP/6-31G*, B3LYP/def2-TZVPPD, and MP2(full)/6-31G* levels of theory, using the input from the relevant minima (reactant and products) and an estimate of the TS obtained from the relaxed PES scans. Single point energies for 1Me5XC, H+(1Me5XC), TSs, and four proton-bound dimers were determined at the B3LYP/6-311+G(2d,2p), B3LYP/def2-TZVPPD, and MP2(full)/6-311+G(2d,2p) levels of theory using geometries optimized at the B3LYP/6-31G*, B3LYP/def2-TZVPPD, and MP2(full)/6-31G* levels, respectively. Frequency analyses at the MP2(full)/def2-TZVPPD require computational resources beyond those available to us; therefore, single point energy calculations performed at the MP2(full)/def2-TZVPPD make use of the B3LYP/def2-TZVPPD optimized structures. Zero-point energy (ZPE) corrections were determined using vibrational frequencies calculated at the B3LYP and MP2(full) levels of theory and scaled by factors of 0.9804 and 0.9646, respectively [52]. To obtain accurate energetics, basis set superposition error corrections (BSSEs) are also included in the calculated BPEs using the full counterpoise approach [53, 54].
Thermochemical Analysis
The threshold regions of the measured CID cross-sections are modeled using procedures developed elsewhere [55–62] that have been found to reproduce CID cross-sections well [63–67]. Details regarding data handling and analysis procedures, which includes explicitly accounting for the internal and translational energy distributions of the reactant proton-bound dimers, the effects of multiple ion-neutral collisions, and the lifetime of the dissociating proton-bound dimers, are summarized in the Supplementary Information.
Results and Discussion
Cross-Sections for Collision-Induced Dissociation
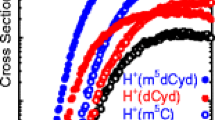
Experimental cross sections were obtained for the interaction of Xe with four proton-bound dimers, (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC). The energy-dependent CID cross-sections of all four proton-bound dimers are shown in Figure 1. Over the collision energy range examined, typically ~0–6 eV, the only dissociation pathway observed for the proton-bound homodimers corresponds to cleavage of the three hydrogen bonds responsible for the binding in these species resulting in loss of the neutral nucleobase in the CID Reactions 1.
Cross sections for collision-induced dissociation of the (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC) proton-bound dimers with Xe as a function of collision energy in the center-of-mass frame (lower x-axis) and laboratory frame (upper x-axis). Data are shown for the Xe pressure of ~0.1 mTorr
CID of the (C)H+(1Me5XC) proton-bound heterodimers leads to two dissociation pathways that occur in parallel and compete with each other, Reactions 2 and 3.
This behavior is consistent with fragmentation via IRMPD [27, 28] and CID [29–31] of similar proton-bound dimers. Production of the protonated nucleobase having the higher PA is energetically favored over production of the protonated nucleobase with the lower PA. The apparent CID thresholds suggest that the N3 PAs of these nucleobases follow the order: C ~ 1Me5FC > 1Me5BrC. However, kinetic effects may alter the relative order as a result of the tight competition between the dissociation pathways of Reactions 2 and 3.
Theoretical Results
As discussed above, the stable tautomeric conformations of neutral cytosine, C, and protonated cytosine, H+(C), and various 1- and 5-methylated and 5-halogenated derivatives have previously been examined at the B3LYP/6-31G*, B3LYP/def2-TZVPPD, MP2(full)/6-31G* and MP2(full)/def2-TZVPPD levels of theory in the IRMPD and TCID studies we previously reported [29–31]. These calculations are expanded here to include structures for 1Me5FC and 1Me5BrC as well as the proton-bound dimers, (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC), and were optimized at the same levels of theory as described in the theoretical calculations section. The geometry-optimized structures of the three most stable tautomeric conformations of C, 1Me5FC, and 1Me5BrC and their relative Gibbs free energies at 298 K are included in Scheme 2, whereas the analogous results for the protonated nucleobases, H+(C), H+(1Me5FC), and H+(1Me5BrC) are shown in Scheme 3. Enthalpies at 0 K and relative Gibbs free energies of these neutral and protonated nucleobases are summarized in Supplementary Table S3. To differentiate the various stable low-energy tautomeric conformations of these species, lower case Roman numerals are used to describe the tautomeric conformations of the neutral nucleobase, whereas upper case Roman numerals with a “+” sign are used to describe the tautomeric conformations of the protonated nucleobase, and both are ordered based on the relative Gibbs free energies at 298 K of the low-energy tautomeric conformations of C and H+(C). The B3LYP/def2-TZVPPD optimized structures of the ground-state conformations of the four proton-bound dimers examined here are shown in Scheme 4. As can be seen in the scheme, the ground-state structures of all four proton-bound dimers involve three hydrogen bonds and adopt an anti-parallel configuration of the protonated and neutral nucleobases, corresponding to the most commonly observed conformation in multi-stranded DNAs. In the ground-state tautomeric conformation of the heterodimers, the excess proton is bound to the nucleobase computed to have the higher PA. These ground-state conformers are designated as II + •••i_3a to indicate that the excited II + tautomeric conformation of the protonated nuceleobase binds to the ground-state i tautomeric conformation of the neutral nucleobase. The underscore 3a designation indicates that the binding occurs via three hydrogen-bonding interactions and the protonated and neutral nucleobases are bound in an anti-parallel configuration. It is unclear whether tautomerization to the O2-protonated nucleobases, I +, will occur during the dissociation of these complexes. Therefore, PES and TS calculations were performed to determine the height of the tautomerization barriers. The reaction coordinate diagrams for dissociation of the (1Me5FC)H+(1Me5FC) and (C)H+(1Me5FC) proton-bound dimers to produce neutral i and N3-protonated II + or O2-protonated I + products are shown in Figure 2, along with the PES for the excited i•••II + _3a to dissociate to the i and II + products. Parallel results were obtained for the bromine-containing proton-bound dimers and are included in Supplementary Figure S1. The relative energies along the PESs for these dissociation pathways determined at all four levels of theory for all four proton-bound dimers are summarized in Supplementary Table S4. In the TSs of all four proton-bound dimers, the excess proton is chelating with the O2 and N3 atoms. As can be seen in the PESs of Figure 2 and Supplementary Figure S1, the tautomerization barriers (176.4–189.5 kJ/mol) exceed the dissociation energies for simple cleavage of the three hydrogen bonds (160.8–166.1 kJ/mol) by 9.4–25.5 kJ/mol, indicating that at threshold, tautomerization will not occur. The tautomerization barriers were also determined at the B3LYP/6-311+G(2d,2p), MP2(full)/6-311+G(2d,2p), and MP2(full)/def2-TZVPPD levels to ensure that the barriers computed are not highly sensitive to the basis sets and level of theory employed. As can be seen in Supplementary Table S4, the computed tautomerization barriers exceed the dissociation energies for simple cleavage of the three hydrogen bonds for all four proton-bound dimers (diabatic dissociation) regardless of the level of theory employed, confirming that tautomerization will not occur upon dissociation at threshold energies, and indicating that BPEs involving simple cleavage of the three hydrogen bonds of the proton-bound dimers and N3 PAs of the nucleobases are measured in the experiments. At elevated energies, dissociation accompanied by tautomerization will also contribute to the observed dissociation behavior. However, the pathways associated with these tight TS pathways will be much slower and, therefore, will be minor contributors to the overall rate of dissociation, and thus should not exert a significant impact on the threshold determinations. BPEs including ZPE and BSSE corrections calculated for the dissociation pathways that produce the N3-protonated products (II +) at the B3LYP and MP2(full) levels of theory using the 6-311+G(2d,2p) and def2-TZVPPD basis sets are summarized in Table 1.
B3LYP/def2-TZVPPD optimized geometries of the three most stable tautomeric conformations of protonated cytosine, H+(C), protonated 1-methyl-5-fluorocytosine, H+(1Me5FC), and protonated 1-methyl-5-bromocytosine, H+(1Me5BrC). Relative Gibbs free energies at 298 K calculated at the B3LYP/def2-TZVPPD level of theory are also shown
B3LYP/def2-TZVPPD potential energy surfaces for dissociation of the ground-state II + •••I_3a conformations of the (1Me5FC)H+(1Me5FC) and (C)H+(1Me5FC) proton-bound dimers to produce 1Me5FC_i or C_i and protonated H+(1Me5FC)_I + products and neutral 1Me5FC_i or C_i and protonated H+(1Me5FC)_II + products, and dissociation of the first excited i•••II+_3a conformation of (C)H+(1Me5FC) proton-bound dimer to produce C_i and protonated H+(1Me5FC)_II + products, parts (a) through (c), respectively
Threshold Analysis
As described in the Supplementary Information, the model of Equation S1 was used to analyze the thresholds for Reaction 1 for the (1Me5FC)H+(1Me5FC) and (1Me5BrC)H+(1Me5BrC) proton-bound homodimers, whereas Equation S2 was used to analyze the thresholds for Reactions 2 and 3 for the (C)H+(1Me5FC) and (C)H+(1Me5BrC) proton-bound heterodimers. As concluded from the theoretical results, tautomerization will not occur upon CID at threshold energies and, therefore, the tautomeric forms of the protonated and neutral nucleobase products are the same as in the proton-bound dimers, II + and i, respectively. Theoretical calculations also found that the hydrogen bond involving the excess proton provides ~100 kJ/mol of stabilization energy for the proton-bound dimer, whereas the two additional neutral hydrogen bonds each add ~30 kJ/mol additional stabilization. Therefore, the reaction coordinate involves lengthening of the N3−H+ •••N3 hydrogen bond, which leads to simultaneous lengthening and cleavage of the other two neutral hydrogen bonds. Based on the computational results, a loose phase space limit transition state (PSL TS) model [60] is applied. The results of these analyses are summarized in Supplementary Table S5 and shown in Figure 3. The threshold energies determined are also summarized in Table 1. For the homodimers, the experimental cross sections for Reaction 1 are accurately reproduced using a loose PSL TS [60] model for the (1Me5XC)H+(1Me5XC)_II + •••i_3a → H+(1Me5XC)_II + + 1Me5X_i CID pathway. In the cases of the heterodimers, the experimental cross sections for Reactions 2 and 3 are accurately reproduced using the loose PSL TS model for the (C)H+(1Me5XC)_II + •••i_3a → H+(1Me5XC)_II + + C_i and (C)H+(1Me5XC)_II + •••i_3a → H+(C)_II + + 1Me5X_i CID pathways, respectively, confirming our assumption that tautomerization does not occur upon dissociation at or near threshold energies, and indicating that the ground-state II + •••i_3a structures are accessed in the experiments. However, the TCID experiments suggest that production of H+(1Me5FC) and H+(1Me5BrC) is energetically favored over production of H+(C), whereas theory suggests the opposite trend. To establish whether or not these differences could be the result of an inappropriate treatment of the TSs for dissociation in these proton-bound heterodimer systems, we comprehensively examined a variety of alternative TS models, where either the TS for the H+(1Me5XC) + C pathway was made tighter or the TS associated with the H+(C) + 1Me5XC pathway was made looser. However, in all cases, these models led to poorer fits to the data and poorer agreement with theory and, thus, we believe that these differences represent problems with the theoretical models.
Zero-pressure-extrapolated cross sections for collision-induced dissociation of the (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC) proton-bound dimers with Xe in the threshold region as a function of kinetic energy in the center-of-mass frame (lower x-axis) and the laboratory frame (upper x-axis), parts (a) through (d), respectively. The solid lines show the best fits to the data using the models of Equations S1 and S2 convoluted over the neutral and ion kinetic and internal energy distributions. The dotted lines show the model cross sections in the absence of experimental kinetic energy broadening for the complexes with an internal temperature of 0 K. The data and models are shown expanded by a factor of 10 and offset from zero in the insets
The relative N3 PAs of cytosine and the 5-halocytosines and 1-methyl-5-halocytosines are also obtained from competitive analyses of these dissociation pathways for the proton-bound heterodimers examined here and those previously examined [31] and are summarized in Table 2. Supplementary Table S5 also includes threshold values, E 0, obtained without inclusion of the RRKM lifetime analysis. Comparison of these results with the E 0(PSL) values provides a measurement of the kinetic shift associated with the finite experimental time window.
The entropy of activation, ΔS †, is a measure of the looseness of the TS and also a reflection of the complexity of the system. ΔS † is largely determined from the molecular constants used to model the energized complex and the TS, but also depends on the threshold energy, E 0(PSL). The ΔS †(PSL) values at 1000 K are listed in Supplementary Table S5, and vary between 91 and 100 J•K–1•mol–1 across these systems. The large positive entropies of activation determined result from the fact that while the two neutral hydrogen bonds contribute to the stability, they also conformationally constrain the reactant proton-bound dimer such that the loose PSL TS is a product-like structure that occurs at the centrifugal barrier for dissociation.
Discussion
Comparison of Experiment and Theory
BPEs of the four proton-bound dimers at 0 K measured here by TCID techniques are summarized in Table 1. Also listed in Table 1 are the BPEs of the proton-bound dimers calculated at the B3LYP and MP2(full) levels of theory using the 6-311+G(2d,2p) and def2-TZVPPD basis sets, and including ZPE and BSSE corrections. The agreement between the measured and B3LYP/def2-TZVPPD calculated BPEs of the proton-bound homo- and heterodimers is illustrated in Figure 4, parts a and b, respectively, whereas results for all four levels of theory are compared in Supplementary Figure S2. The TCID experiments suggest that Reaction 2 (loss of neutral C) is energetically favored over Reaction 3 (loss of neutral 1Me5XC) for both proton-bound heterodimers, (C)H+(1Me5FC) and (C)H+(1Me5BrC), whereas theory suggests that Reaction 3 is the lowest energy dissociation pathway for these two complexes. The BPEs listed in Table 1 and plotted in Figure 4b and Supplementary Figure S2 correspond to those for the lowest energy dissociation pathways. In order to understand the effects of 1-methylation and 5-halogentation on the BPEs, the measured and calculated BPEs of the (C)H+(C), (1MeC)H+(1MeC), (5FC)H+(5FC), (5BrC)H+(5BrC), (C)H+(1MeC), (C)H+(5FC), and (C)H+(5BrC) complexes are also included for comparison [29–31]. The mean absolute deviations (MADs) between theory and experiment for the B3LYP/def2-TZVPPD and B3LYP/6-311+G(2d,2p) levels of theory are 3.9 ± 1.7 and 4.4 ± 2.5 kJ/mol, respectively. The MADs for the B3LYP results are smaller than the average experimental uncertainty (AEU) in these values, 4.7 ± 0.5 kJ/mol, suggesting that the B3LYP level of theory accurately describes the hydrogen-bonding interactions responsible for the binding in these proton-bound dimers, with the def2-TZVPPD results being slightly more accurate. The MP2(full) level of theory does not perform nearly as well. The MADs between the MP2(full)/def2-TZVPPD and MP2(full)/6-311+G(2d,2p) results and the measured values are 39.9 ± 7.5 and 36.3 ± 5.0 kJ/mol, respectively, significantly greater than the MADs for the B3LYP values and the AEU. The agreement between the MP2(full) calculated and TCID measured values improves to 26.0 ± 6.7 and 17.0 ± 5.0 kJ/mol when BSSE corrections are not included, consistent with previous TCID studies on similar proton-bound dimers [29–31]. This is also consistent with previous theoretical studies of hydrogen-bonded complexes [68–75], which have shown that at least triple-zeta-quality basis sets are required to accurately describe systems where there can be significant intramolecular noncovalent interactions, and the BSSE corrections can get rather large for MP2 calculations when flexible but still unsaturated basis sets are used. Based on the comparisons between theory and experiment, it is clear that B3LYP theory describes the base-pairing interactions in the proton-bound dimers more accurately, whereas MP2(full) underestimates the strength of the base-pairing interactions in all complexes.
TCID measured BPEs of (xC)H+(xC) homodimers at 0 K (in kJ/mol), where xC = C, 1MeC, 5FC, 5BrC, 1Me5FC, and 1Me5BrC, plotted versus B3LYP/def2-TZVPPD calculated values including ZPE and BSSE corrections. BPEs of the (5FC)H+(5FC), (5BrC)H+(5BrC), and (1MeC)H+(1MeC) proton-bound dimers are taken from references [29] and [30], part (a). TCID measured BPEs of (C)H+(xC) heterodimers at 0 K (in kJ/mol) plotted versus B3LYP/def2-TZVPPD calculated values including ZPE and BSSE corrections. BPEs of the (C)H+(C), (C)H+(5FC), (C)H+(5BrC), and (C)H+(1MeC) proton-bound dimers are taken from references [29–31], part (b). Values determined in this work are taken from Table 1. The black solid diagonal line indicates the values for which the calculated and measured BPEs are equal
The measured and B3LYP/def2-TZVPPD calculated relative N3 PAs are listed in Table 2 and compared pictorially in Figure 5a. As can be seen in Figure 5a, the B3LYP/def2-TZVPPD level of theory provides good estimates for the relative N3 PAs of C versus 1MeC, 5FC, and 5BrC, whereas theory underestimates the relative N3 PAs of C versus 1Me5FC and 1Me5BrC. The MAD between theory and experiment for the relative N3 PAs is 6.0 ± 7.4 kJ/mol, larger than the AEU in these values, 1.4 ± 0.9 kJ/mol. The B3LYP/def2-TZVPPD calculations indicate that the N3 PAs of 1Me5FC and 1Me5BrC are smaller than C, whereas TCID experiments suggest that the N3 PAs of 1Me5FC and 1Me5BrC exceed that of C. However, both theory and experiment find that the PA of 1Me5FC lies between those of 1MeC and 5FC. Likewise, both theory and experiment find that the PA of 1Me5BrC lies between those of 1MeC and 5BrC.
Absolute N3 PAs at 298 K of the four halogenated cytosines, 5FC, 5BrC, 1Me5FC, and 1Me5BrC, are derived from TCID of the (C)H+(1Me5FC) and (C)H+(1Me5BrC) proton-bound heterodimers examined here as well as the (C)H+(5FC) and (C)H+(5BrC) heterodimers previously investigated [31] and the PA of C previously determined [29–38]. The results of these analyses are summarized in Table 2. The N3 PAs of C, 1MeC, and the halogenated cytosines follow the order: 1MeC (964.7 ± 2.9 kJ/mol) > 1Me5BrC (959.9 ± 3.3 kJ/mol) > 1Me5FC (955.7 ± 3.3 kJ/mol) > C (949.2 ± 2.8 kJ/mol) > 5BrC (930.9 ± 3.6 kJ/mol) > 5FC (926.3 ± 3.5 kJ/mol). The absolute N3 PAs determined are compared with B3LYP/def2-TZVPPD calculated values in Figure 5b. The MAD between theory and experiment for the absolute N3 PAs is 6.1 ± 3.6 kJ/mol, almost twice the AEU in these values, 3.3 ± 0.3 kJ/mol, and is largely the result of theory underestimating the N3 PAs of 1Me5FC and 1Me5BrC.
Influence of 1-Methylation and 5-Halogenation on the N3 PA
As can be seen in Figure 5, 5-halogenation leads to a decrease in the N3 PA of cytosine, whereas 1-methylation leads to an increase in the N3 PA. This is the expected behavior and is easily understood based on the electronic properties of the methyl and halogen substituents. The methyl substituent is an electron donating moiety and, therefore, increases the electron density within the aromatic ring, leading to stabilization of the positive charge associated with the excess proton. The halogens are electron withdrawing and, therefore, decrease the electron density within the aromatic ring, resulting in destabilization of the positive charge associated with the excess proton. This behavior is also consistent with observations made in previous TCID studies of the proton-bound dimers of 1-methylated cytosines [29]. The effect of the 1-methyl substituent on the N3 PA is rather consistent. The increases in the N3 PAs from C to 1MeC, 5FC to 1Me5FC, and 5BrC to 1Me5BrC are 15.5, 19.4, and 20.0 kJ/mol, respectively. The TCID measured N3 PAs of cytosine and the modified cytosines follow the order: 1MeC > 1Me5BrC > 1Me5FC > C > 5BrC > 5FC. This order differs slightly from the trend suggested by the apparent thresholds because the competition between the two dissociation pathways is tight such that kinetic effects alter the relative order of the dissociation onsets.
Influence of 1-Methylation and 5-Halogenation on the BPEs
The measured and calculated BPEs at 0 K of the four proton-bound dimers measured here along with values reported for the proton-bound homo- and heterodimers of C, 1MeC, 5FC, and 5BrC, are listed in Table 1 and shown in Figure 4. The BPEs of the proton-bound dimers of the 5-halogenated cytosines are smaller than that of the (C)H+(C) homodimer [30], indicating that 5-halogenation decreases the base-pairing interactions in the proton-bound dimers. Experimentally, 1-permethylation is found to exert very little influence on the BPE, whereas theory suggests that 1-permethylation of cytosine leads to a small decrease in the BPE. 1-Methylation of a single cytosine residue decreases the BPE [29]. For the proton-bound dimers of the 1-methyl-5-halocytosines, both theory and experiments suggest a decrease in the BPE. However, the decrease in the BPE is smaller than the uncertainties in these measurements. Thus, 5-halogenation of cytosine residues should result in destabilization of DNA i-motif conformations, but the effects of 5-halogenation are much less significant when cytosine is 1-methylated.
Implications for the Stability of DNA i-Motif Conformations
The base-pairing interactions in the proton-bound dimer of cytosine are the major forces responsible for stabilization of DNA i-motif conformations. Previous TCID studies of proton-bound homodimers of cytosine, and 1-methylated and 5-halogenated cytosines found that 1-hypermethylation of cytosine produces a very slight increase in the BPE, and should, therefore, result in minor stabilization of DNA i-motif conformations. In contrast, 5-hyperhalogenation of cytosine leads to a small decrease in BPE of the proton-bound dimer and would, therefore, tend to destabilize DNA i-motif conformations [29, 30]. In contrast, the present TCID results indicate that 5-halogentation of cytosine residues has almost no effect on the strength of the base-pairing interactions when cytosine is methylated at the N1 position and, thus, should have little or no effect on the stability of DNA i-motif conformations. In the case of proton-bound heterodimers, 1-methylation [29], 5-halogenation [30], and 1-methyl-5-halogenation of a single cytosine residue lead to a decrease in the BPE. However, the decrease in the BPE upon 1-methyl-5-halogenation of a single cytosine is smaller than the uncertainties in these measurements. Thus, 5-halogention of a single cytosine residue should not significantly alter the stabilization of DNA i-motif conformations. By extension, these results also suggest that the BPE of the proton-bound dimer of 5-halo-2′-deoxycytidine (x5Cyd) should be roughly equal to that of C. However, polarizability effects may also play a role such that this conclusion must be experimentally (and theoretically) verified and is the subject of future investigations. However, the BPEs of all of the proton-bound heterodimers examined to date are still much greater than those of canonical Watson-Crick G•C and neutral C•C base pairs, suggesting that DNA i-motif conformations are still favored over conventional base pairing. Thus, although halogenation of cytosine at the C5 positions tends to weaken the base-pairing interactions in the proton-bound dimers of cytosine, the effects are sufficiently small that i-motif conformations should be stable to such modifications. Although the change in the BPE induced by halogenation is not large for a single proton-bound dimer, the accumulated effect could be dramatic in diseased state trinucleotide repeats associated with the fragile-X syndrome where more than 230 trinucleotides and hundreds of halogenated proton-bound dimers could be present. Because 5-halogenation at any cytosine residue may lead to a decrease in the BPE, the influence of halogenation will be seen in the number of trinucleotide repeats required to induce structural conversion from canonical Watson-Crick base pairing to DNA i-motif conformations.
To further probe the influence of modifications on the stability of DNA i-motif conformations, other factors that play roles in stabilizing/destabilizing these noncanonical structures such as nucleobase-stacking interactions and steric effects associated with nucleobase orientation and the folding of the nucleic acid strands must also be considered. Follow-up work to examine how these base-pairing interactions evolve in increasingly larger model systems including proton-bound dimers of the analogous 2′-deoxycytidine nucleosides [76] and nucleotides and extending to (CCG) n trinucleotide repeats that are associated with the formation of i-motif conformations and fragile X syndrome are being pursued. Present results indicate that the B3LYP level of theory provides accurate estimates for the energetics of binding in such proton-bound dimers and, therefore, may be suitable for investigating larger and more biologically relevant model systems. 1-Methylation, a mimic for the 2′-deoxyribose moiety as well as the actual 2′-deoxyribose moiety have almost no effect on the BPE of the proton-bound dimer and, hence, should exert almost no effect on the stability of DNA i-motif conformations. In contrast, 5-halogenation of cytosine exerts a larger more destabilizing influence and would tend to destabilize DNA i-motif conformations, thereby requiring more trinucleotide repeats for structural conversion of Watson-Crick base-paired DNA to i-motif conformations. Information provided by this work including structures, the energy-dependent dissociation behavior, and relative stabilities of these proton-bound dimers should also facilitate experiments and data interpretation for studies of larger and more biologically relevant model systems.
Conclusions
5-Halogenation of cytosine, one of the most common DNA damage pathways, can regulate gene expression by altering the structure and stability of DNA or DNA–protein interactions. In order to understand the effects of 5-halogenation of 1-methylcytosine on the base-pairing interactions responsible for stabilizing DNA i-motif conformations and the proton affinities of the modified nucleobases, the threshold collision-induced dissociation behaviors of four proton-bound dimers, (1Me5FC)H+(1Me5FC), (1Me5BrC)H+(1Me5BrC), (C)H+(1Me5FC), and (C)H+(1Me5BrC), are examined in a guided ion beam tandem mass spectrometer. The only dissociation pathway observed for the proton-bound homodimers corresponds to cleavage of the three hydrogen bonds responsible for the binding in these species resulting in loss of the neutral nucleobase. For the proton-bound heterodimers, two dissociation pathways involving production of the two protonated nucleobases occur in parallel and compete with each other. PESs were calculated to determine the heights of tautomerization barriers for dissociation of the ground-state conformations of the proton-bound dimers to produce O2-protonated nucleobase products (I +). The calculations confirm that the tautomerization barriers exceed the dissociation energy for production of the N3-protonated nucleobases (II +) such that tautomerization will not occur upon dissociation at or near threshold energies. Thresholds corresponding to BPEs for CID reactions that produce the N3-protonated nucleobase are determined after careful consideration of the effects of the kinetic and internal energy distributions of the proton-bound dimer and Xe reactants, multiple collisions with Xe, and the lifetime of the activated proton-bound dimers using a loose PSL TS model. Competitive threshold analyses of the two dissociation pathways that occur in parallel for the proton-bound heterodimers provide the relative N3 PAs of cytosine and the halogenated cytosines. Theoretical estimates for the BPEs of the proton-bound dimers and the N3 PAs of 5-halogenated and 1-methyl-5-halogenated cytosines are determined from calculations performed at the B3LYP and MP2(full) levels of theory using the 6-311+G(2d,2p) and def2-TZVPPD basis sets. Reasonably good agreement between experimental and theoretical BPEs is found for the B3LYP level of theory, whereas MP2(full) theory produces values that are systematically low, even when BSSE corrections are not included in the computed BPEs. Reasonable agreement is also achieved for the measured and B3LYP/def2-TZVPPD calculated relative and absolute N3 PAs of cytosine and 5-halogenated cytosines. However, theory seems to underestimate the N3 PAs of 1Me5FC and 1Me5BrC. These results suggest that calculations at the B3LYP/def2-TZVPPD level of theory can be employed to provide reliable energetic predictions for related systems that bind via multiple hydrogen bonds. Halogenation clearly influences the base-pairing interactions in the proton-bound dimers. 5-Halogenation is found to decrease the BPE of the (C)H+(C) proton-bound dimer [30, 31] but exert a much less dramatic effect on the BPE when the cytosine residues are 1-methylated. These results suggest that DNA i-motif conformations should be destabilized under 5-halogenation conditions. However, the BPEs of all halogenated proton-bound dimers still significantly exceed those of canonical Watson-Crick G•C and neutral C•C base pairs, suggesting that the effects of halogenation are not sufficient to destroy DNA i-motif conformations but may alter the number of trinucleotide repeats necessary to induce structural conversion from canonical Watson-Crick base-pairing to DNA i-motif conformations. Halogenation is found to decrease the N3 PA of cytosine. The N3 PAs of C, 1MeC, and the halogenated cytosines follow the order: 1MeC (964.7 ± 2.9 kJ/mol) > 1Me5BrC (959.9 ± 3.3 kJ/mol) > 1Me5FC (955.7 ± 3.3 kJ/mol) > C (949.2 ± 2.8 kJ/mol) > 5BrC (930.9 ± 3.6 kJ/mol) > 5FC (926.3 ± 3.5 kJ/mol), indicating that 1-methylation has a greater influence on the N3 PAs than C5-halogenation, whereas theory underestimates the N3 PA of 1Me5BrC and 1Me5FC and suggests that the order of N3 PAs is: 1MeC > C > 1Me5BrC > 1Me5FC > 5BrC >5FC.
References
Riggs, A.D., Jones, P.A.: 5-Methylcytosine, gene regulation, and cancer. Adv. Cancer Res. 40, 1–30 (1983)
Bird, A.: DNA methylation patterns and epigenetic memory. Gene. Dev. 16, 6–21 (2002)
Cooper, D.N., Youssoufian, H.: The CpG dinucleotide and human genetic disease. Hum. Genet. 78, 151–155 (1988)
Rideout III, W.M., Coetzee, G.A., Olumi, A.F., Jones, P.A.: 5-Methylcytosine as an endogenous mutagen in the human LDL receptor and p53 genes. Science 249, 1288–1290 (1990)
McLenan, Y., Polussa, J., Tassone, F., Hagerman, R.: Fragile X syndrome. Curr. Genom. 12, 216–224 (2011)
Garber, K.B., Visootsak, J., Warren, S.T.: Fragile X syndrome. Eur. J. Hum. Genet. 16, 666–672 (2008)
Sutcliffe, J.S., Nelson, D.L., Zhang, F., Pieretti, M., Caskey, C.T., Saxe, D., Warren, W.T.: DNA methylation represses FMR-1 transcription in fragile X syndrome. Hum. Mol. Genet. 1, 397–400 (1992)
Oberlé, I., Rousseau, F., Heitz, D., Kretz, C., Devys, D., Hanauer, A., Boué, J., Bertheas, M.F., Mandel, J.L.: Instability of a 550-base pair DNA segment and abnormal methylation in fragile X syndrome. Science 252, 1097–1102 (1991)
Pieretti, M., Zhang, F., Fu, Y.-H., Warren, S.T., Oostra, B.A., Caskey, C.T., Nelson, D.L.: Absence of expression of the FMR-1 gene in fragile X syndrome. Cell 66, 817–822 (1991)
Darlow, J.M., Leach, D.R.F.: Secondary structure in d(CGG) and d(CCG) repeat tracts. J. Mol. Biol. 275, 3–16 (1998)
Mitas, M.: Trinucleotide repeats associated with human disease. Nucleic Acids Res. 25, 2245–2253 (1997)
Gehring, K., Leroy, J.-L., Guéron, M.: A tetrameric DNA structure with protonated cytosine–cytosine base pairs. Nature 363, 561–565 (1993)
Fotjík, P., Vorlícková, M.: The fragile X chromosome (GCC) repeat folds into a DNA tetraplex at neutral pH. Nucleic Acids Res. 29, 4684–4690 (2001)
Rosu, F., Gabelica, V., Joly, L., Grégoire, G., Pauw, E.D.: Zwitterionic i-motif structures are preserved in DNA negatively charged ions produced by electrospray mass spectrometry. Phys. Chem. Chem. Phys. 12, 13448–13454 (2010)
Esteller, M.: Aberrant dna methylation as a cancer inducing mechanism. Annu. Rev. Pharmacol. Toxicol. 45, 629–656 (2005)
Brennan, C.A., Van Cleve, M.D., Gumport, R.I.: The effects of base analogue substitutions on the methylation by the EcoRI modification methylase of octadeoxyribonucleotides containing modified EcoRI recognition sequences. J. Biol. Chem. 261, 7279–7286 (1986)
Brennan, C.A., Van Cleve, M.D., Gumport, R.I.: The effects of base analogue substitutions on the cleavage by the EcoRI restriction endonuclease of octadeoxyribonucleotides containing modified EcoRI recognition sequences. J. Biol. Chem. 261, 7270–7278 (1986)
Valinluck, V., Liu, P., Kang Jr., J.I., Burdzy, A., Sowers, L.C.: 5-Halogenated pyrimidine lesions within a CpG sequence context mimic 5-methylcytosine by enhancing the binding of the methyl-CpG-binding domain of methyl-CpG-binding protein 2 (MeCP2). Nucleic Acids Res. 33, 3057–3064 (2005)
Whiteman, M., Jenner, A., Halliwell, B.: Hypochlorous acid-induced base modifications in isolated calf thymus DNA. Chem. Res. Toxicol. 10, 1240–1246 (1997)
Henderson, J.P., Byun, J., Williams, M.V., Mueller, D.M., McCormick, M.L., Heinecke, J.W.: Production of brominating intermediates by myeloperoxidase. A transhalogenation pathway for generating mutagenic nucleobases during inflammation. J. Biol. Chem. 276, 7867–7875 (2001)
Badouard, C., Masuda, M., Nishino, H., Cadet, J., Favier, A., Ravanat, J.L.: Detection of chlorinated DNA and RNA nucleosides by HPLC coupled to tandem mass spectrometry as potential biomarkers of inflammation. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 827, 26–31 (2005)
Morris, S.M.: The genetic toxicology of 5-bromodeoxyuridine in mammalian cells. Mutat. Res. 258, 161–188 (1991)
Morris, S.M.: The genetic toxicology of 5-fluoropyrimidines and 5-chlorouracil. Mutat. Res. 297, 39–51 (1993)
Vanetich, K.M., Santi, D.V.: Thymidylate synthase and fluorouracil. Adv. Exp. Med. Biol. 244, 113–124 (1998)
Smith, S.S., Kaplan, B.E., Sowers, L.C., Newman, E.M.: Mechanism of human methyl-directed DNA methyltransferase and the fidelity of cytosine methylation. Proc. Natl. Acad. Sci. U. S. A. 89, 4744–4748 (1992)
Klimasauskas, S., Kumar, S., Roberts, R.J., Cheng, X.: HhaI methyltransferase flips its target base out of the DNA helix. Cell 76, 357–369 (1994)
Yang, B., Wu, R.R., Berden, G., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of proton-bound dimers of cytosine and modified cytosines: Effects of modifications on gas-phase conformations. J. Phys. Chem. B 117, 14191–14201 (2013)
Oomens, J., Moehlig, A.R., Morton, T.H.: Infrared multiple photon dissociation (IRMPD) spectroscopy of the proton-bound dimer of 1-methylcytosine in the gas phase. J. Phys. Chem. Lett. 1, 2891–2897 (2010)
Yang, B., Moehlig, A.R., Frieler, C.E., Rodgers, M.T.: Base-pairing energies of protonated nucleobase pairs and proton affinities of 1-methylated cytosines: Model systems for the effects of the sugar moiety on the stability of DNA i-motif conformations. J. Phys. Chem. B 119, 1857–1868 (2014)
Yang, B., Wu, R.R., Rodgers, M.T.: Base-pairing energies of proton-bound homodimers determined by guided ion beam tandem mass spectrometry: Application to cytosine and 5-substituted cytosines. Anal. Chem. 85, 11000–11006 (2013)
Yang, B., Rodgers, M.T.: Base-pairing energies of proton-bound heterodimers of cytosine and modified cytosines: Implications for the stability of DNA i-motif conformations. J. Am. Chem. Soc. 136, 282–290 (2014)
Little, C.D., Nau, M.M., Carney, D.N., Gazdar, A.F., Minna, J.D.: Amplification and expression of the c-myc oncogene in human lung cancer cell lines. Nature 306, 194–196 (1983)
Munzel, P., Marx, D., Kochel, H., Schauer, A., Bock, K.W.: Genomic alterations of the c-myc protooncogene in relation to the overexpression of c-erbB2 and Ki-67 in human breast and cervix carcinomas. J. Cancer Res. Clin. Oncol. 117, 603–607 (1991)
Clark, H.M., Yano, T., Otsuki, T., Jaffe, E.S., Shibata, D., Raffeld, M.: Mutations in the coding region of c-myc in AIDS-associated and other aggressive lymphomas. Cancer Res. 54, 3383–3386 (1994)
Sunami, T., Kondo, J., Hirao, I., Watanabe, K., Miura, K., Takenaka, A.: Structure of d(GCGAAGC) (hexagonal form): A base-intercalated duplex as a stable structure. Acta Crystallogr. D Biol. Crystallogr. 60, 90–96 (2004)
Hunter, E.P., Lias, S.G.: Evaluated gas phase basicities and proton affinities of molecules: An update. J. Phys. Chem. Ref. Data 27, 413–656 (1998)
NIST Chemistry Webbook, available at http://webbook.nist.gov/chemistry/. Accessed 1 Oct 2014
Liu, M., Li, T., Amegayibor, S., Cardoso, D.S., Fu, Y., Lee, J.K.: Gas-phase thermochemical properties of pyrimidine nucleobases. J. Org. Chem. 73, 9283–9291 (2008)
Rodgers, M.T.: Substituent effects in the binding of alkali metal ions to pyridines studied by threshold collision-induced dissociation and ab initio theory: The methylpyridines. J. Phys. Chem. A 105, 2374–2383 (2001)
Teloy, E., Gerlich, D.: Integral cross sections for ion-molecule reactions. I. The guided ion beam technique. Chem. Phys. 4, 417–427 (1974)
Gerlich, D: Inhomogeneous rf fields: A versatile tool for the study of processes with slow ions. Thesis [Diplomarbeit] University of Freiburg, Federal Republic of Germany] (1971)
Gerlich, D.: in State-selected and state-to-state ion-molecule reaction dynamics, Part I. Experiment. Ng, C.-Y.; Baer, M. Eds. Adv. Chem. Phys. 82, 1–176 (1992)
Dalleska, N.F., Honma, K., Armentrout, P.B.: Stepwise solvation enthalpies of protonated water clusters: Collision-induced dissociation as an alternative to equilibrium studies. J. Am. Chem. Soc. 115, 12125–12131 (1993)
Aristov, N., Armentrout, P.B.: Collision-induced dissociation of vanadium monoxide ion. J. Phys. Chem. 90, 5135–5140 (1986)
Hales, D.A., Armentrout, P.B.: Effect of internal excitation on the collision-induced dissociation and reactivity of Co2 +. J. Clust. Sci. 1, 127–142 (1990)
Frisch, M.J., Trucks, G.W., Schlegel, H.B., Scuseria, G.E., Robb, M.A., Cheeseman, J.R., Scalmani, G., Barone, V., Mennucci, B., Petersson, G.A., Nakatsuji, H., Caricato, M., Li, X., Hratchian, H.P., Izmaylov, A.F., Bloino, J., Zheng, G., Sonnenberg, J.L., Hada, M., Ehara, M., Toyota, K., Fukuda, R., Hasegawa, J., Ishida, M., Nakajima, T., Honda, Y., Kitao, O., Nakai, H., Vreven, T., Montgomery Jr., J.A., Peralta, J.E., Ogliaro, F., Bearpark, M., Heyd, J.J., Brothers, E., Kudin, K.N., Staroverov, V.N., Kobayashi, R., Normand, J., Raghavachari, K., Rendell, A., Burant, J.C., Iyengar, S.S., Tomasi, J., Cossi, M., Rega, N., Millam, J.M., Klene, M., Knox, J.E., Cross, J.B., Bakken, V., Adamo, C., Jaramillo, J., Gomperts, R., Stratmann, R.E., Yazyev, O., Austin, A.J., Cammi, R., Pomelli, C., Ochterski, J.W., Martin, R.L., Morokuma, K., Zakrzewski, V.G., Voth, G.A., Salvador, P., Dannenberg, J.J., Dapprich, S., Daniels, A.D., Farkas, Ö., Foresman, J.B., Ortiz, J.V., Cioslowski, J., Fox, D.J.: Gaussian 09, Revision A.1. Gaussian, Inc, Wallingford (2009)
Weigend, F., Ahlrichs, R.: Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: Design and assessment of accuracy. Phys. Chem. Chem. Phys. 7, 3297–3305 (2005)
Feller, D.: The role of databases in support of computational chemistry calculations. J. Comput. Chem. 17, 1571–1586 (1996)
Schuchardt, K.L., Didier, B.T., Elsethagen, T., Sun, L., Gurumoorthi, V., Chase, J., Li, L., Windus, T.L.: Basis set exchange: a community database for computational science. J. Chem. Inf. Mol. 47, 1045–1052 (2007)
Smith, S.M., Markevitch, A.N., Romanov, D.A., Li, X., Levis, R.J., Schlegel, H.B.: Static and dynamic polarizabilities of conjugated molecules and their cations. J. Phys. Chem. A 108, 11063–11072 (2004)
Peng, C., Schlegel, H.B.: Combining synchronous transit and quasi-Newton methods for finding transition states. Isr. J. Chem. 33, 449–454 (1993)
Exploring chemistry with electronic structures methods, 2nd ed.; Foresman, J.B.; Frisch, Æ. Gaussian: Pittsburg p. 64 (1996)
Boys, S.F., Bernardi, R.: Calculation of small molecular interactions by differences of separate total energies—Some procedures with reduced errors. Mol. Phys. 19, 553–566 (1979)
van Duijneveldt, F.B., van Duijneveldt-van de Rijdt, J.G.C.M., van Lenthe, J.H.: State of the art in counterpoise theory. Chem. Rev. 94, 1873–1885 (1994)
Muntean, F., Armentrout, P.B.: Guided ion beam study of collision-induced dissociation dynamics: Integral and differential cross sections. J. Chem. Phys. 115, 1213–1228 (2001)
Beyer, T.S., Swinehart, D.S.: Number of multiply-restricted partitions [A1]. Commun. ACM 16, 379 (1973)
Stein, S.E., Rabinovitch, B.S.: Accurate evaluation of internal energy level sums and densities including anharmonic oscillators and hindered rotors. J. Chem. Phys. 58, 2438–2445 (1973)
Stein, S.E., Rabinovitch, B.S.: On the use of exact state counting methods in RRKM rate calculations. Chem. Phys. Lett. 49, 183–188 (1997)
Khan, F.A., Clemmer, D.E., Schultz, R.H., Armentrout, P.B.: Sequential bond energies of Cr(CO)x +, x = 1–6. J. Phys. Chem. 97, 7978–7987 (1993)
Rodgers, M.T., Ervin, K.M., Armentrout, P.B.: Statistical modeling of collision-induced dissociation thresholds. J. Chem. Phys. 106, 4499–4508 (1997)
Chesnavich, W.J., Bowers, M.T.: Theory of translationally driven reactions. J. Phys. Chem. 83, 900–905 (1979)
Rodgers, M.T., Armentrout, P.B.: Statistical modeling of competitive threshold collision-induced dissociation. J. Chem. Phys. 109, 1787–1800 (1998)
Armentrout, P.B., Yang, B., Rodgers, M.T.: Metal cation dependence of interactions with amino acids: Bond energies of Rb+ and Cs+ to Met, Phe, Tyr, and Trp. J. Phys. Chem. B 117, 3771–3781 (2013)
Nose, H., Chen, Y., Rodgers, M.T.: Energy-resolved collision-induced dissociation studies of 1,10-phenanthroline complexes of the late first-row divalent transition-metal cations: Determination of the third sequential binding energies. J. Phys. Chem. A 117, 4316–4330 (2013)
Nose, H., Rodgers, M.T.: Energy-resolved collision-induced dissociation studies of 2,2′-bipyridine complexes of the late first-row divalent transition-metal cations: Determination of the third sequential binding energies. Chem. Plus Chem. 78, 1109–1123 (2013)
Armentrout, P.B., Austin, C.A., Rodgers, M.T.: Alkali metal cation interactions with 15-crown-5 in the gas phase: Revisited. J. Phys. Chem. A 118, 8088–8097 (2014)
Yang, B., Rodgers, M.T.: Alkali metal cation binding affinities of cytosine in the gas phase: Revisited. Phys. Chem. Chem. Phys. 16, 16110–16220 (2014)
Sim, F., St-Amant, A., Pápai, I., Salahub, D.R.: Gaussian density functional calculations on hydrogen-bonded systems. J. Am. Chem. Soc. 114, 4391–4400 (1992)
Laasonen, K., Parrinello, M., Car, R., Lee, C., Vanderbilt, D.: Structure of small water clusters using gradient-corrected density functional theory. Chem. Phys. Lett. 207, 208–213 (1993)
Kim, K., Jordan, K.D.: Comparison of density functional and MP2 calculations on the water monomer and dimer. J. Phys. Chem. 98, 10089–10094 (1994)
Del Bene, J.E., Person, W.B., Szczepaniak, K.: Properties of hydrogen-bonded complexes obtained from the B3LYP functional with 6–31G(d, p) and 6–31+G(d, p) basis sets: Comparison with MP2/6-31+G(d, p) results and experimental data. J. Phys. Chem. 99, 10705–10707 (1995)
Novoa, J.J., Sosa, C.: Evaluation of the density functional approximation on the computation of hydrogen bond interactions. J. Phys. Chem. 99, 15837–15845 (1995)
Hobza, P., Šponer, J., Reschel, T.: Density functional theory and molecular clusters. J. Comput. Chem. 16, 1315–1325 (1995)
Suhai, S.: Density functional studies of the hydrogen-bonded network in an infinite water polymer. J. Phys. Chem. 99, 1172–1181 (1995)
Valdés, H., Klusák, V., Pitoňák, M., Exner, O., Starý, I., Hobza, P., Rulisek, L.: Evaluation of the intramolecular basis set super-position error in the calculations of large molecules: [n]Helicenes and Phe-Gly-Phe tripeptide. J. Comput. Chem. 29, 861–870 (2008)
Yang, B., Rodgers, M.T.: Base-pairing energies of protonated nucleoside base pairs of dCyd and m5dCyd: Implications for the stability of DNA i-motif conformations. J. Am. Soc. Mass Spectrom. (2015). doi:10.1007/s13361-015-1144-8
Acknowledgments
The authors acknowledge financial support for this work provided by the National Science Foundation, grant CHE-1409420. The authors thank Wayne State University C&IT for computer time. B.Y. also gratefully acknowledges support from a Wayne State University Thomas C. Rumble Graduate Fellowship and Summer Dissertation Fellowship. Thanks are also due to T. H. Morton for providing the 1-methylated-5-halocytosines.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Complete citation for reference 46. A description of data handling, thermochemical analysis, and thermal correction procedures is provided. Tables of B3LYP/6-31G* vibrational frequencies, average vibrational energies at 298 K, and rotational constants of the neutral and protonated nucleobases as well as the proton-bound nucleobase dimers, B3LYP/def-TZVPPD relative stabilities of the three low-energy tautomeric conformation of xC and H+(xC), relative energies of species along the PES for dissociation of the (1Me5BrC)H+(1Me5BrC) and (C)H+(1Me5BrC) proton-bound dimers, fitting parameters of Equations S1 and S2, and enthalpies and free energies of the base-pairing interactions at 298 K. Figures comparing TCID measured and calculated BPEs at the B3LYP and MP2(full) levels of theory using the def2-TZVPPD and 6-311+G(2d,2p) basis sets. (PDF 643 kb)
Rights and permissions
About this article
Cite this article
Yang, B., Wu, R.R. & Rodgers, M.T. Base-Pairing Energies of Proton-Bound Dimers and Proton Affinities of 1-Methyl-5-Halocytosines: Implications for the Effects of Halogenation on the Stability of the DNA i-Motif. J. Am. Soc. Mass Spectrom. 26, 1469–1482 (2015). https://doi.org/10.1007/s13361-015-1174-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-015-1174-2