Abstract
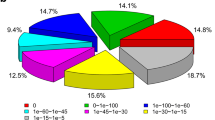
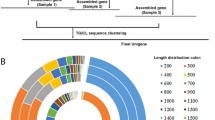
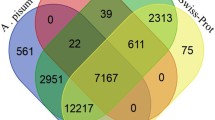
We performed the Illumina sequencing of the yellow mealworm beetle, Tenebrio molitor L., a global stored grain pest and a model for insect physiological studies, and analyzed the transcriptome dataset. We obtained 82,001,576 clean reads that were assembled into 31,785 unigenes with a mean size of 1,022 bp. Among these unigenes, 16,025 (50.41 %) showed significant similarity (E value <10−5) to known proteins in the NCBI non-redundant database. Gene ontology, Cluster of Orthologous Groups and Kyoto Encyclopedia of Genes and Genomes analyses were used to classify functions of the unigenes. We identified 27 putative glutathione S-transferase (GST) genes, including 2 delta, 13 epsilon, 1 omega, 5 sigma, 1 theta and 1 zeta class GSTs, and 2 microsomal GSTs and 2 “unclassified” GSTs. Quantitative reverse transcription-PCR assay showed that 2 and 3 GST genes were mainly expressed in larval and pupal stages, respectively; while 10 genes were mainly expressed in the adult stage. Tissue-specific expression analysis found that, among the 27 GST genes, 8 were expressed predominately in the midgut, 13 were mainly detected in Malpighian tubules, and only one gene specifically distributed in the fat body. This work provides the first insights into the T. molitor GSTs at the molecular level: the results may assist in a functional study for the GSTs in this insect species.




Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Arrese EL, Soulages JL (2010) Insect fat body: energy, metabolism, and regulation. Annu Rev Entomol 55:207–225
Després L, David J-P, Gallet C (2007) The evolutionary ecology of insect resistance to plant chemicals. Trends Ecol Evol 22:298–307
Ding Y, Ortelli F, Rossiter L, Hemingway J, Ranson H (2003) The Anopheles gambiae glutathione transferase supergene family: annotation, phylogeny and expression profiles. BMC Genomics 4:35
Dow JAT, Davies SA (2006) The Malpighian tubule: rapid insights from post-genomic biology. J Insect Physiol 52:365–378
Enayati AA, Ranson H, Hemingway J (2005) Insect glutathione transferases and insecticide resistance. Insect Mol Biol 14:3–8
Friedman R (2011) Genomic organization of the glutathione S-transferase family in insects. Mol Phylogenet Evol 61:924–932
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Hayes JD, Flanagan JU, Jowsey IR (2005) Glutathione transferases. Annu Rev Pharmacol Toxicol 45:51–88
Hemingway J, Ranson H (2000) Insecticide resistance in insect vectors of human disease. Annu Rev Entomol 45:371–391
Higgins LG, Hayes JD (2011) Mechanisms of induction of cytosolic and microsomal glutathione transferase (GST) genes by xenobiotics and pro-inflammatory agents. Drug Metab Rev 43:92–137
Hu F, Dou W, Wang J-J, Jia F-X, Wang J-J (2014) Multiple glutathione S-transferase genes: identification and expression in oriental fruit fly, Bactrocera dorsalis. Pest Manag Sci 70:295–303
Iseli C, Jongeneel CV, Bucher P (1999) ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. In: Proceedings for the 7th International Conference on Intelligent Systems for Molecular Biology, pp 138–148
Keeling C, Yuen M, Liao N, Roderick Docking T, Chan S, Taylor G, Palmquist D, Jackman S, Nguyen A, Li M, Henderson H, Janes J, Zhao Y, Pandoh P, Moore R, Sperling F, Huber DW, Birol I, Jones S, Bohlmann J (2013) Draft genome of the mountain pine beetle, Dendroctonus ponderosae Hopkins, a major forest pest. Genome Biol 14:R27
Ketterman AJ, Saisawang C, Wongsantichon J (2011) Insect glutathione transferases. Drug Metab Rev 43:253–265
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Lumjuan N, Stevenson BJ, Prapanthadara LA, Somboon P, Brophy PM, Loftus BJ, Severson DW, Ranson H (2007) The Aedes aegypti glutathione transferase family. Insect Biochem Mol Biol 37:1026–1035
Nakamura C, Yajima S, Miyamoto T, Sue M (2013) Structural analysis of an epsilon-class glutathione transferase from housefly, Musca domestica. Biochem Biophys Res Commun 430:1206–1211
Qin G, Jia M, Liu T, Xuan T, Yan Zhu K, Guo Y, Ma E, Zhang J (2011) Identification and characterisation of ten glutathione S-transferase genes from oriental migratory locust, Locusta migratoria manilensis (Meyen). Pest Manag Sci 67:697–704
Qin G, Jia M, Liu T, Zhang X, Guo Y, Zhu KY, Ma E, Zhang J (2012) Heterologous expression and characterization of a sigma glutathione S-transferase involved in carbaryl detoxification from oriental migratory locust, Locusta migratoria manilensis (Meyen). J Insect Physiol 58:220–227
Qin G, Jia M, Liu T, Zhang X, Guo Y, Zhu KY, Ma E, Zhang J (2013) Characterization and functional analysis of four glutathione S-transferases from the migratory locust, Locusta migratoria. PLoS One 8:e58410
Rajarapu SP, Mittapalli O (2013) Glutathione-S-transferase profiles in the emerald ash borer, Agrilus planipennis. Comp Biochem Phys B 165:66–72
Ramsey JS, Rider DS, Walsh TK, De Vos M, Gordon KHJ, Ponnala L, Macmil SL, Roe BA, Jander G (2010) Comparative analysis of detoxification enzymes in Acyrthosiphon pisum and Myzus persicae. Insect Mol Biol 19:155–164
Ranson H, Collins F, Hemingway J (1998) The role of alternative mRNA splicing in generating heterogeneity within the Anopheles gambiae class I glutathione S-transferase family. Proc Natl Acad Sci USA 95:14284–14289
Ranson H, Rossiter L, Ortelli F, Jensen B, Wang X, Roth CW, Collins FH, Hemingway J (2001) Identification of a novel class of insect glutathione S-transferases involved in resistance to DDT in the malaria vector Anopheles gambiae. Biochem J 359:295–304
Rogers ME, Jani MK, Vogt RG (1999) An olfactory-specific glutathione-S-transferase in the sphinx moth Manduca sexta. J Exp Biol 202:1625–1637
Shi H, Pei L, Gu S, Zhu S, Wang Y, Zhang Y, Li B (2012) Glutathione S-transferase (GST) genes in the red flour beetle, Tribolium castaneum, and comparative analysis with five additional insects. Genomics 100:327–335
Singh SP, Coronella JA, Beneš H, Cochrane BJ, Zimniak P (2001) Catalytic function of Drosophila melanogaster glutathione S-transferase DmGSTS1-1 (GST-2) in conjugation of lipid peroxidation end products. Eur J Biochem 268:2912–2923
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tang Q-Y, Zhang C-X (2013) Data Processing System (DPS) software with experimental design, statistical analysis and data mining developed for use in entomological research. Insect Sci 20:254–260
Toba G, Aigaki T (2000) Disruption of the microsomal glutathione S-transferase-like gene reduces life span of Drosophila melanogaster. Gene 253:179–187
Vontas JG, Small GJ, Hemingway J (2001) Glutathione S-transferases as antioxidant defence agents confer pyrethroid resistance in Nilaparvata lugens. Biochem J 357:65–72
Wang Y, Qiu L, Ranson H, Lumjuan N, Hemingway J, Setzer WN, Meehan EJ, Chen L (2008) Structure of an insect epsilon class glutathione S-transferase from the malaria vector Anopheles gambiae provides an explanation for the high DDT-detoxifying activity. J Struct Biol 164:228–235
Wongsantichon J, Ketterman AJ (2005) Alternative splicing of glutathione S-transferases. In: Helmut S, Lester P (eds) Methods in enzymology. Academic Press, San Diego, pp 100–116
Yamamoto K, Zhang PB, Banno Y, Fujii H (2006) Identification of a sigma-class glutathione-S-transferase from the silkworm, Bombyx mori. J Appl Entomol 130:515–522
Yamamoto K, Usuda K, Kakuta Y, Kimura M, Higashiura A, Nakagawa A, Aso Y, Suzuki M (2012) Structural basis for catalytic activity of a silkworm delta-class glutathione transferase. Biochim Biophys Acta 1820:1469–1474
Yamamoto K, Higashiura A, Suzuki M, Aritake K, Urade Y, Uodome N, Nakagawa A (2013) Crystal structure of a Bombyx mori sigma-class glutathione transferase exhibiting prostaglandin E synthase activity. Biochim Biophys Acta 1830:3711–3718
Yu Q, Lu C, Li B, Fang S, Zuo W, Dai F, Zhang Z, **ang Z (2008) Identification, genomic organization and expression pattern of glutathione S-transferase in the silkworm, Bombyx mori. Insect Biochem Mol Biol 38:1158–1164
Zhou W-W, Li X-W, Quan Y-H, Cheng J, Zhang C-X, Gurr G, Zhu Z-R (2012) Identification and expression profiles of nine glutathione S-transferase genes from the important rice phloem sap-sucker and virus vector Laodelphax striatellus (Fallén) (Hemiptera: Delphacidae). Pest Manag Sci 68:1296–1305
Zhou W-W, Liang Q-M, Xu Y, Gurr GM, Bao Y-Y, Zhou X-P, Zhang C-X, Cheng J, Zhu Z-R (2013) Genomic insights into the glutathione S-transferase gene family of two rice planthoppers, Nilaparvata lugens (Stål) and Sogatella furcifera (Horváth) (Hemiptera: Delphacidae). PLoS One 8:e56604
Acknowledgments
This work was supported by the Industry Project of the Ministry of Agriculture of China (201403030), the National Natural Science Foundation of China (31401734, 31371935), and grants from Anhui Agricultural University (2013ZR008, YJ2014-2).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, S., Shi, XX., Jiang, YD. et al. De novo analysis of the Tenebrio molitor (Coleoptera: Tenebrionidae) transcriptome and identification of putative glutathione S-transferase genes. Appl Entomol Zool 50, 63–71 (2015). https://doi.org/10.1007/s13355-014-0305-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13355-014-0305-8