Abstract
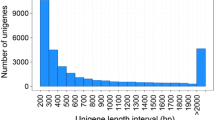
Philomycus bilineatus is a highly common gastropod mollusk pest in China and is also utilized to treat infectious diseases. However, no genomic resources are available for this non-model species. In the present study, the transcriptomic analysis of P. bilineatus was completed. After sequencing using the next generation sequencing technology, 9.11 Gb of clean reads were obtained, which led to the assembly and annotation of 145,523 transcripts and 125,690 unigenes. Unigenes were functionally classified using Gene Ontology (GO), euKaryotic Ortholog Groups of proteins (KOG), and Kyoto Encyclopedia of Genes and Genomes (KEGG). A total of 27,554 unigenes were assigned into 55 GO terms, 13,989 unigenes were differentiated into 26 KOG categories, and 16,368 unigenes were assigned to 229 KEGG pathways. Furthermore, 16,614 simple sequence repeats (SSRs), 38 olfactory genes, and 40 antimicrobial peptide/protein genes were identified. The transcriptome profile of P. bilineatus will provide a valuable genomic resource for further study, will promote the development of new pest management strategies through interference of chemosensory communication, and will support potential medicinal uses of this species.








Similar content being viewed by others
References
Bjorstad A, Fu H, Karlsson A, Dahlgren C, Bylund J (2005) Interleukin-8-derived peptide has antibacterial activity antimicrob agents. Chemother 49:3889–3895. https://doi.org/10.1128/AAC.49.9.3889-3895.2005
Brito NF, Moreira MF, Melo AC (2016) A look inside odorant-binding proteins in insect chemoreception. J Insect Physiol 95:51–65. https://doi.org/10.1016/j.**sphys.2016.09.008
Bruhn KW, Spellberg B (2015) Transferrin-mediated iron sequestration as a novel therapy for bacterial and fungal infections. Curr Opin Microbiol 27:57–61. https://doi.org/10.1016/j.mib.2015.07.005
Cattaneo AM (2018) Current status on the functional characterization of chemosensory receptors of Cydia pomonella (Lepidoptera: Tortricidae). Front Behav Neurosci 12:189. https://doi.org/10.3389/fnbeh.2018.00189
Chipman DM, Sharon N (1969) Mechanism of lysozyme action Science 165:454–465
Deng YW et al (2014) De novo assembly, gene annotation, and simple sequence repeat marker development using Illumina paired-end transcriptome sequences in the pearl oyster Pinctada maxima. Biosci Biotechnol Biochem 78:1685–1692. https://doi.org/10.1080/09168451.2014.936351
Du XJ, Wang JX, Liu N, Zhao XF, Li FH, **ang JH (2006) Identification and molecular characterization of a peritrophin-like protein from fleshy prawn (Fenneropenaeus chinensis). Mol Immunol 43:1633–1644. https://doi.org/10.1016/j.molimm.2005.09.018
Du L, Zhao X, Liang X, Gao X, Liu Y, Wang G (2018) Identification of candidate chemosensory genes in Mythimna separata by transcriptomic analysis. BMC Genomics 19:518. https://doi.org/10.1186/s12864-018-4898-0
Fan J, Francis F, Liu Y, Chen JL, Cheng DF (2011) An overview of odorant-binding protein functions in insect peripheral olfactory reception. Genet Mol Res 10:3056–3069. https://doi.org/10.4238/2011.December.8.2
Fiedler TJ, Hudder A, McKay SJ, Shivkumar S, Capo TR, Schmale MC, Walsh PJ (2010) The transcriptome of the early life history stages of the California Sea Hare Aplysia californica. Comp Biochem Physiol Part D Genomics Proteomics 5:165–170. https://doi.org/10.1016/j.cbd.2010.03.003
Fu N, Wang Q, Shen HL (2013) De novo assembly, gene annotation and marker development using Illumina paired-end transcriptome sequences in celery (Apium graveolens L.). PLoS One 8:e57686. https://doi.org/10.1371/journal.pone.0057686
Gifford JL, Hunter HN, Vogel HJ (2005) Lactoferricin: a lactoferrin-derived peptide with antimicrobial, antiviral, antitumor and immunological properties. Cell Mol Life Sci 62:2588–2598. https://doi.org/10.1007/s00018-005-5373-z
Grabherr MG et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652. https://doi.org/10.1038/nbt.1883
Guo Y, He H (2014) Identification and characterization of a goose-type lysozyme from sewage snail Physa acuta fish shellfish. Immunol. 39:321–325. https://doi.org/10.1016/j.fsi.2014.05.029
He R et al (2017) Molecular characteristics and serodiagnostic potential of chitinase-like protein from Sarcoptes scabiei. Oncotarget 8:83995–84005. https://doi.org/10.18632/oncotarget.21056
Heyland A, Vue Z, Voolstra CR, Medina M, Moroz LL (2011) Developmental transcriptome of Aplysia californica. J Exp Zool B Mol Dev Evol 316B:113–134. https://doi.org/10.1002/jez.b.21383
Hirose T, Sunazuka T, Omura S (2010) Recent development of two chitinase inhibitors, Argifin and Argadin, produced by soil microorganisms. P Jpn Acad B-Phys 86:85–102. https://doi.org/10.2183/pjab.86.85
Huang Y, Ma F, Wang W, Ren Q (2015) Identification and molecular characterization of a peritrophin-like gene, involved in the antibacterial response in Chinese mitten crab, Eriocheir sinensis. Dev Comp Immunol 50:129–138. https://doi.org/10.1016/j.dci.2015.01.002
Hultmark D, Steiner H, Rasmuson T, Boman HG (1980) Insect immunity. Purification and properties of three inducible bactericidal proteins from hemolymph of immunized pupae of Hyalophora cecropia. Eur J Biochem 106:7–16
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kang SW et al (2016) Transcriptome sequencing and de novo characterization of Korean endemic land snail, Koreanohadra kurodana for functional transcripts and SSR markers. Mol Gen Genomics 291:1999–2014. https://doi.org/10.1007/s00438-016-1233-9
Kang SW et al (2017) Sequencing and de novo assembly of visceral mass transcriptome of the critically endangered land snail Satsuma myomphala: annotation and SSR discovery. Comp Biochem Phys D 21:77–89. https://doi.org/10.1016/j.cbd.2016.10.004
Kang K et al (2018) Identification of putative fecundity-related gustatory receptor genes in the brown planthopper Nilaparvata lugens. BMC Genomics 19:970. https://doi.org/10.1186/s12864-018-5391-5
Lesoway MP, Abouheif E, Collin R (2016) Comparative transcriptomics of alternative developmental phenotypes in a marine gastropod. J Exp Zool Part B 326:151–167. https://doi.org/10.1002/jez.b.22674
Li X, Li M, Hou L, Zhang ZY, Pang XM, Li YY (2018) De novo transcriptome assembly and population genetic analyses for an endangered Chinese endemic Acer miaotaiense (Aceraceae). Genes-Basel. 9:378. https://doi.org/10.3390/genes9080378
Li Z, Meng M, Li S, Deng B (2019) The transcriptome analysis of Protaetia brevitarsis Lewis larvae. PLoS One 14:e0214001. https://doi.org/10.1371/journal.pone.0214001
Liu NY, Li ZB, Zhao N, Song QS, Zhu JY, Yang B (2018) Identification and characterization of chemosensory gene families in the bark beetle, Tomicus yunnanensis. Comp Biochem Physiol Part D Genomics Proteomics 25:73–85. https://doi.org/10.1016/j.cbd.2017.11.003
Magister S, Kos J (2013) Cystatins in immune system. J Cancer 4:45–56. https://doi.org/10.7150/jca.5044
Moroz LL et al (2006) Neuronal transcriptome of Aplysia: neuronal compartments and circuitry. Cell 127:1453–1467. https://doi.org/10.1016/j.cell.2006.09.052
Mutz KO, Heilkenbrinker A, Lonne M, Walter JG, Stahl F (2013) Transcriptome analysis using next-generation sequencing. Curr Opin Biotechnol 24:22–30. https://doi.org/10.1016/j.copbio.2012.09.004
Parmakelis A, Kotsakiozi P, Kontos CK, Adamopoulos PG, Scorilas A (2017) The transcriptome of a “slee**” invader: de novo assembly and annotation of the transcriptome of aestivating Cornu aspersum. BMC Genomics 18:491. https://doi.org/10.1186/s12864-017-3885-1
Pelosi P, Iovinella I, Zhu J, Wang G, Dani FR (2018) Beyond chemoreception: diverse tasks of soluble olfactory proteins in insects. Biol Rev Camb Philos Soc 93:184–200. https://doi.org/10.1111/brv.12339
Tagai C, Morita S, Shiraishi T, Miyaji K, Iwamuro S (2011) Antimicrobial properties of arginine- and lysine-rich histones and involvement of bacterial outer membrane protease T in their differential mode of actions. Peptides 32:2003–2009. https://doi.org/10.1016/j.peptides.2011.09.005
Tellam RL, Wijffels G, Willadsen P (1999) Peritrophic matrix proteins. Insect Biochem Mol Biol 29:87–101
Tian Z et al (2018) Antennal transcriptome analysis of the chemosensory gene families in Carposina sasakii (Lepidoptera: Carposinidae). BMC Genomics 19:544. https://doi.org/10.1186/s12864-018-4900-x
Tonk M, Vilcinskas A, Rahnamaeian M (2016) Insect antimicrobial peptides: potential tools for the prevention of skin cancer. Appl Microbiol Biotechnol 100:7397–7405. https://doi.org/10.1007/s00253-016-7718-y
Venthur H, Zhou JJ (2018) Odorant receptors and odorant-binding proteins as insect pest control targets: a comparative analysis front. Physiol. 9:1163. https://doi.org/10.3389/fphys.2018.01163
Vogt RG et al (2009) The insect SNMP gene family insect. Biochem Molec 39:448–456. https://doi.org/10.1016/j.ibmb.2009.03.007
Vosshall LB, Hansson BS (2011) A unified nomenclature system for the insect olfactory coreceptor. Chem Senses 36:497–498. https://doi.org/10.1093/chemse/bjr022
Wang W, Hui JHL, Chan TF, Chu KH (2014) De novo transcriptome sequencing of the snail echinolittorina malaccana: identification of genes responsive to thermal stress and development of genetic markers for population studies. Mar Biotechnol 16:547–559. https://doi.org/10.1007/s10126-014-9573-0
Wu Q, Patocka J, Kuca K (2018) Insect antimicrobial peptides, a mini review. Toxins (Basel) 10. https://doi.org/10.3390/toxins10110461
Yi HY, Chowdhury M, Huang YD, Yu XQ (2014) Insect antimicrobial peptides and their applications. Appl Microbiol Biotechnol 98:5807–5822. https://doi.org/10.1007/s00253-014-5792-6
Yi X, Qi J, Zhou X, Hu MY, Zhong GH (2017) Differential expression of chemosensory-protein genes in midguts in response to diet of Spodoptera litura. Sci Rep 7:296. https://doi.org/10.1038/s41598-017-00403-5
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14. https://doi.org/10.1186/gb-2010-11-2-r14
Yuvaraj JK, Andersson MN, Zhang DD, Lofstedt C (2018) Antennal transcriptome analysis of the chemosensory gene families from trichoptera and basal lepidoptera. Front Physiol 9:1365. https://doi.org/10.3389/fphys.2018.01365
Zhang G et al (2012) The oyster genome reveals stress adaptation and complexity of shell formation. Nature. 490:49–54. https://doi.org/10.1038/nature11413
Zhang L, Li L, Zhu Y, Zhang G, Guo X (2014) Transcriptome analysis reveals a rich gene set related to innate immunity in the Eastern oyster (Crassostrea virginica). Mar Biotechnol (NY) 16:17–33. https://doi.org/10.1007/s10126-013-9526-z
Funding
This work was supported by The National Natural Science Foundation of China (grant number: 81703475).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All animal collection and utility protocols were approved by the Henan University of Science and Technology of Biology Animal Use Ethics Committee.
Additional information
Communicated by: Maciej Szydlowski
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, Z., Yuan, Y., Meng, M. et al. De novo transcriptome of the whole-body of the gastropod mollusk Philomycus bilineatus, a pest with medical potential in China. J Appl Genetics 61, 439–449 (2020). https://doi.org/10.1007/s13353-020-00566-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-020-00566-4