Abstract
Autism spectrum disorder (ASD) is a heterogeneous group of early-onset neurodevelopmental disorders known to be highly heritable with a complex genetic architecture. Abnormal brain developmental trajectories that impact synaptic functioning, excitation-inhibition balance and brain connectivity are now understood to play a central role in ASD. Ongoing efforts to identify the genetic underpinnings still prove challenging, in part due to phenotypic and genetic heterogeneity.
This review focuses on parent-of-origin effects (POEs), where the phenotypic effect of an allele depends on its parental origin. POEs include genomic imprinting, transgenerational effects, mitochondrial DNA, sex chromosomes and mutational transmission bias. The motivation for investigating these mechanisms in ASD has been driven by their known impacts on early brain development and brain functioning, in particular for the most well-documented POE, genomic imprinting. Moreover, imprinting is implicated in syndromes such as Angelman and Prader-Willi, which frequently share comorbid symptoms with ASD. In addition to other regions in the genome, this comprehensive review highlights the 15q11-q13 and 7q chromosomal regions as well as the mitochondrial DNA as harbouring the majority of currently identified POEs in ASD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autism spectrum disorder (ASD) represents a heterogeneous group of common (prevalence of 0.83% (Vos et al. 2017)), early-onset, neurodevelopmental conditions (American Psychiatric Association 2013). ASD is characterised by differences in social interactions and communication as well as by restricted and repetitive behaviours and interests (Hodges et al. 2020). Much of the evidence to date points to the aetiology of ASD being related to brain development, converging on the abnormal development of synaptic functioning, excitation-inhibition balance and brain connectivity (Betancur et al. 2009; McFadden and Minshew 2013; Sohal and Rubenstein 2019).
ASD is known to be highly heritable (heritability approx. 80% (Bai et al. 2019)) with a complex genetic architecture. The majority of heritability is due to common variation, with rare and de novo structural variation contributing to a lesser extent (Searles Quick et al. 2021). ASD is typically grouped into either syndromic or non-syndromic ASD. Syndromic ASD, ASD plus or complex ASD occurs in approximately 25% of patients with the remaining 75% having what is referred to as essential or non-syndromic ASD (Carter and Scherer 2013). Syndromic ASD presents with additional phenotypes and/or dysmorphic features and is typically associated with chromosomal abnormalities or mutations in a single gene with diagnosis usually determined through some form of genetic testing (Fernandez and Scherer 2017; Sztainberg and Zoghbi 2016). For the majority of non-syndromic ASD cases, the genetic aetiology is unknown and is still proving challenging due to the considerable phenotypic and genetic heterogeneity present (Rylaarsdam and Guemez-Gamboa 2019). Other, more complex mechanisms of transmission may explain some of the as-yet unaccounted for heritability (Yoon et al. 2020). For instance, parent-of-origin effects (POEs) are a group of complex genetic effects that alter the phenotype in the offspring through a variety of mechanisms that modify gene expression in a parent-of-origin specific manner (Guilmatre and Sharp 2012). The most well-known mechanism responsible for POEs is genomic imprinting; however, other mechanisms include transgenerational effects (for example, maternal genetic effects and mother–offspring interaction effects), mitochondrial DNA, sex chromosomes and mutational transmission bias (for example, triplet-repeat-associated diseases) (see Guilmatre and Sharp 2012 for a general POE review). Advances in the field of ASD genomic research over the last 10 years, coupled with new and emerging evidence, make this an excellent time for this original review, highlighting the role of POEs in ASD aetiology. In this review, we introduce different POE mechanisms, giving motivation for their potential roles in ASD, and subsequently detail specific evidence for particular genes and genomic regions showing POEs in ASD. Throughout the review, we use the term autism spectrum disorder (ASD) but acknowledge that studies referred to may have had different definitions/diagnostic criteria and use the term autism for example.
Parent of origin mechanisms
Genomic imprinting
We inherit one copy of each autosomal gene from each of our parents. For the vast majority of these genes, both copies are capable of being expressed; however, for a small subset (128 genes in humans, geneimprint.com (imprint status = Imprinted-All), January 2022), one copy does not function (either partially or completely) due to the epigenetic mechanism, genomic imprinting. This mechanism silences one chromosome region in a parent-of-origin-dependent manner, leading to expression of only one copy of the gene. Imprinting is regulated by parental-specific epigenetic markers, including methylation and histone modifications established in gametogenesis and early embryonic development (Court et al. 2014; Hutter et al. 2010). Here we refer to imprinted genes as either maternally imprinted when the maternal copy of the allele is silenced or paternally imprinted when the paternal copy of the allele is silenced.
Imprinting is often tissue- and/or temporal-specific (Abramowitz and Bartolomei 2012; Reik and Walter 2001) and is necessary for optimal functioning of this small set of genes, most of which are involved in early development (including embryonic, placental and post-natal development), and behaviour, especially social behaviour (Falls et al. 1999; Lawson et al. 2013). Copy number variations (deletions or duplications), uniparental disomies (both chromosomes, or parts of a chromosome, are inherited from only one parent), aberrant methylation marks (epimutations) and point mutations can disrupt imprinting and have been identified as being associated with a range of human diseases and disorders, including neurodevelopmental and neuropsychiatric disorders (Soellner et al. 2017).
There are several motivations for considering imprinting in ASD. Early work on mouse chimaeras showed that imprinted genes have effects on general early growth, and on brain development in particular, including brain size and cell composition (Allen et al. 1995; Keverne et al. 1996), providing some of the first indications for imprinted genes having a potential role in neurodevelopment. Additionally, behavioural studies conducted on the adult stage of these mouse chimaeras pointed to imprinted genes in the brain as having an impact on behaviours such as aggression (Allen et al. 1995). In humans, many of the identified imprinted genes are expressed in the adult, influencing both cognitive and behavioural phenotypes (Isles and Wilkinson 2000; Davies et al. 2001, 2005)).
The most convincing hypothesis for the evolution of imprinting in mammals is the genetic conflict or parental tug-of-war hypothesis. This hypothesis suggests that maternally and paternally imprinted genes differentially regulate resources passed through the placenta from the mother to the develo** embryo and subsequently exert an influence on the expression of traits in the offspring (Moore and Haig 1991). The conflict has advantages when females are likely to have more than one mate. In such a scenario, it is in the father’s genetic interest to maximise resources for the current pregnancy, as he cannot be certain if future offspring from this mother will share his DNA. In contrast, it is in the mother’s genetic interest to limit the resources shared with each offspring from each pregnancy, as they will all inherit half of her DNA. Thus, genes promoting growth are favoured by the father, whereas the mother will benefit from silencing her copy of such genes (Moore and Haig 1991). A good example of this is the maternally imprinted early growth-promoting gene IGF2 (Constância et al. 2002). When maternal imprinting of IGF2 is disrupted, there is an increase in birth weight, by up to 50% when both copies are expressed, resulting in the imprinting disorder Beckwith‐Wiedemann syndrome (Byars et al. 2014). Whereas when both copies are silenced, Silver-Russell syndrome results, which is characterised by undergrowth (Eggermann et al. 2010). In a study examining behavioural phenotypes, a higher proportion of children with Beckwith‐Wiedemann syndrome than expected also presented with ASD (6.8%) (Kent et al. 2008).
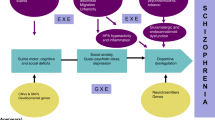
In the case of ASD, this parental tug-of-war hypothesis has been further extended to an imprinted brain developmental theory for ASD and schizophrenia. This theory proposes that imbalances during brain development (resulting from either enhanced effects of maternally imprinted genes, deficits in effects of paternally imprinted genes, or the action of both) can lead to ASD phenotypes. When the imbalance is in the opposite direction, with enhanced maternally biased effects, the risk of schizophrenia increases (Badcock and Crespi 2006, 2008; Byars et al. 2014; Crespi 2013; Crespi and Badcock 2008; Úbeda and Gardner 2010, 2011).
Syndromic forms of ASD have provided additional evidence for imprinted genes having a role in neurodevelopment. The syndromic forms of ASD, Angelman syndrome and Prader-Willi syndrome, are two well-characterised reciprocal chromosome 15q11-q13 imprinting disorders. These disorders present with mental, physical, cognitive and behavioural impacts on their phenotypes, and in a number of cases, with comorbid ASD (Hogart et al. 2010).
Transgenerational effects: maternal genetic effects
Maternal genetic effects occur when the mother’s genotype exerts an influence on the offspring’s phenotype independent of the offspring’s genotype (Wolf and Wade 2009), for example, through the intrauterine environment. Several mechanisms, including the folate- and homocysteine-related pathways and immunity or inflammation, play critical roles in the development of the foetal nervous system in utero and can be impacted by maternal genetic effects. There is established and emerging evidence for their role in ASD (Azzini et al. 2020; Edmiston et al. 2017; Johnson 2003).
Mutational transmission bias: trinucleotide repeats
Large expansions of specific trinucleotide repeat motifs result in trinucleotide-repeat-associated diseases. These motifs are often present in the general population at harmless levels but can expand to reach pathogenic levels when meiotic transmission to offspring takes place. As the rate of contraction and expansion during transmission is often different between males and females for many of these repeats, this results in POEs (Guilmatre and Sharp 2012). The neurodevelopmental disorder, Fragile X syndrome, is an X-linked dominant trinucleotide-repeat disorder and considered a syndromic form of ASD (Abbeduto et al. 2014).
Mitochondrial DNA
Mitochondrial DNA (mtDNA), consisting of 37 genes, is exclusively maternally inherited and, together with many nuclear DNA genes, is responsible for generating the energy needed to power cells (Rossignol and Frye 2012). The complex role of mitochondria across all tissues and organs (only red blood cells have no mitochondria) means that mitochondrial disorders are often multi-systemic and multi-symptomatic. The number of mitochondria per cell varies widely (heteroplasmy), but the brain is known to have a very high demand on mitochondrial energy, in particular at neural synapses (Pei and Wallace 2018). If this energy supply is in any way disrupted, it can impact brain function and by extension could increase risk for neuropsychiatric disorders such as ASD (Chen et al. 2015; Giulivi et al. 2010; Pei and Wallace 2018; Yoo et al. 2017).
Identified parent-of-origin effects in ASD
In the following sections, we detail specific chromosomes, genomic regions and genes where evidence of POEs in ASD has been identified. The key findings outlined here are also summarised in Table 1.
Chromosome 7q
In a linkage study involving ASD-affected families and sib-pairs, significant evidence of paternal identity-by-descent sharing was identified on chromosome 7q32.3–34 (Ashley-Koch et al. 1999). In addition, the authors also detected significant linkage disequilibrium with paternal transmissions in multiplex and simplex ASD families. A genome-wide parent-of-origin linkage analysis conducted in affected sib-pairs identified two distinct regions, at 7q21.1–22.2 and at 7q32.1–32.3, as showing an excess of paternal and maternal identity-by-descent sharing in ASD, respectively (Lamb et al. 2005). Although neither of these peaks overlap with a known imprinted region, they lie close to imprinted gene clusters (Schanen 2006).
CADPS2 gene, chromosome 7q31.32
CADPS2 shows tissue-specific mono-allelic expression (maternally inherited allele expressed in blood and specific brain regions; amygdala) (Bonora et al. 2014). Cadps2-knockout mice show deficits in neuronal development and abnormal social behaviour (Sadakata et al. 2007, 2012; Sadakata and Furuichi 2009, 2010). Given its prominent role in the nervous system and the evidence from mouse studies, it has been considered an excellent candidate ASD gene, and evidence for downregulation in ASD compared to non-ASD brains has been shown (Voineagu et al. 2011). Bonora et al. (2014) identified a novel intragenic deletion in CADPS2 in two siblings with mild intellectual disability (ID) and epilepsy (Bonora et al. 2014). In a follow-up mutation screening study in the same article, the authors discovered a missense variant of maternal origin that disrupts the interaction of CADPS2 and the dopamine receptor D2DR, in a cohort of ASD/ID patients.
MEST gene, chromosome 7q32.2
MEST has long been known to be maternally imprinted with mono-allelic expression exclusively of the paternal copy (Pilvar et al. 2019). Maternal methylation appears to be driving this imprinting effect (Court et al. 2014; Partida et al. 2018; Schneider et al. 2012). Inactivation of the paternal allele for MEST in mice resulted in embryonic growth retardation and abnormal maternal behaviour suggesting a role in adult behaviour; additionally, methylation levels of MEST have been linked to cognitive ability (Lefebvre et al. 1998; Lorgen-Ritchie et al. 2019). Using whole-genome sequencing, an association was found between ASD and recurrent paternal rare cis-regulatory structural variants overlap** variant-intolerant genes (Brandler et al. 2018), including but not limited to CNTN4, LEO1, RAF1 and MEST. Furthermore, these were transmitted to affected offspring more frequently than to their unaffected siblings. Positive association was found between MEST and ASD in a Korean male case–control cohort (Kwack et al. 2008), while a targeted sequencing study of the 7q32 region containing MEST, COPG2 and KLF14 showed nominal positive association (which did not survive Bonferroni correction) between two haploblocks and ASD in a study of 7q32-linked ASD families (Korvatska et al. 2011).
CNTNAP2 gene, chromosome 7q35-36.1
Using a combination of linkage and association analyses, two linkage peaks and a common genetic variant (displaying maternal over-transmission) significantly associated with ASD susceptibility were identified in the CNTNAP2 gene (Arking et al. 2008). CNTNAP2 is one of the largest genes in the human genome and is a member of the neurexin family, playing a role in brain development; regulating interactions between neurons and glia cells and contributing to the development of neuron axon structures (Waterhouse 2013). However, evidence for imprinting of CNTNAP2 is mixed, some studies argue against CNTNAP2 imprinting, at least in the adult human brain (Schneider et al. 2014), and others suggest imprinting might regulate expression of CNTNAP2 under certain tissue-specific and/or developmental-stage specific conditions (I. S. Lee et al. 2015; Lin et al. 2012). A number of other studies have identified associations between this gene and ASD (in particular language development) but have either not tested for or have not identified a POE (Alarcón et al. 2008; Anney et al. 2012; Bakkaloglu et al. 2008; Chiocchetti et al. 2015; Li et al. 1998; Guffanti et al. 2011; Nurmi et al. 2001, 2003).
The majority of Angelman syndrome cases are caused by deletions which lead to a decrease in expression of UBE3A, the remaining due to mutations in UBE3A, paternal uniparental disomy (pUPD) and imprinting defects (Dagli and Williams 2017). For Angelman syndrome, the clinical characteristics include motor dysfunction, intellectual disability, speech impairment and seizures (Greer et al. 2010; Rangasamy et al. 2013). High rates of comorbidity with ASD have been shown for Angelman syndrome (Mertz et al. 2014).
As mentioned, Prader-Willi syndrome individuals with a mUPD show more ASD traits than Prader-Willi syndrome individuals with paternal deletions. Using human post-mortem brain samples, cortical tissue expression of UBE3A was shown to be substantially higher for Prader-Willi syndrome individuals with mUPD than for those with a deletion form (Hogart et al. 2007).
Iossifov et al. reported a de novo T485A missense mutation in UBE3A in an ASD male proband; however, they did not test whether the variant was present on the paternal or maternal copy of the chromosome (Iossifov et al. 2014). A functional investigation of this mutation in UBE3A in human cell culture experiments showed that the T485A variant inhibits UBE3A self-regulation, leading to increased UBE3A activity, and increases dendritic spine formation (Yi et al. 2015).
SNRPN gene, chromosome 15q11.2
SNRPN is a maternally imprinted gene that regulates expression of Nr4a1, a nuclear receptor critical for cortical neuron development (Barr et al. 1995; H. Li et al. 2016; Reed and Leff 1994). Several studies have reported uniparental methylation POEs for this gene (Court et al. 2014; Partida et al. 2018). SNRPN has been proposed as a likely candidate gene for Prader-Willi syndrome (Cassidy et al. 2000). In a trio study design, two SNPs in SNRPN showed marginal imprinting effects (Kim et al. 2008) and a balanced chromosomal abnormality was identified in an individual with ASD without Angelman syndrome or Prader-Willi syndrome (Talkowski et al. 2012). Through a post-mortem brain tissue analysis, Hogart et al. 2009 identified deficiencies in the paternally expressed transcripts of SNRPN in a female individual with ASD and milder Prader-Willi-like characteristics (Hogart et al. 2009). In addition, methylation levels of SNRPN have been linked to cognitive ability (Lorgen-Ritchie et al. 2019).
FMR1 gene, X chromosome
Fragile X syndrome is predominantly caused by a CGG triplet repeat mutation expansion in the promoter region of the FMR1 gene at the chromosome Xq27.3 locus (Kaufmann et al. 2017). This syndrome is understood to be the most common single-gene cause of ASD (1–6% of ASD cases) (Kaufmann et al. 2017). As Fragile X syndrome is X-linked, it affects more males than females and more severely (Kaufmann et al. 2017). Fragile X syndrome is characterised by males almost always exhibiting moderate intellectual disability, having a characteristic appearance (macrocephaly with a prominent forehead, long face, large protruding ears and a prominent chin, the dysmorphism is milder in females) and behaviour (Fernandez and Scherer 2017; Sherman et al. 2005). Males carrying a full mutation (> 200 repeats) do not produce offspring, whereas males with an intermediate (45–54 repeats) or premutation (45–54 repeats) will pass it on, but only daughters will inherit the mutation (Nolin et al. 2019). For maternal transmissions, pre-mutations can expand to full mutations, which is rarely the case for paternal transmissions (Nolin et al. 2003, 2019). This results in a greater chance of offspring develo** Fragile X syndrome when the mother is the premutation carrier compared to the father. In addition, there is a paternal bias for contraction of the premutation triplet repeats (Nolin et al. 2019). This triplet repeat mutation results in abnormal methylation of FMR1 and either partial or complete silencing of the gene (Kidd et al. 2019). The upshot is decreased production of the Fragile X mental retardation protein (FMRP) which has a vital role in synaptic plasticity and brain development as it regulates protein synthesis at the synapses (Penagarikano et al. 2007).
Turner syndrome, X chromosome
Turner syndrome results when one of the X chromosomes (maternal or paternal) is either partially or fully missing and shows an increased rate of ASD (Creswell and Skuse 1999; Donnelly et al. 2000; Skuse et al. 1997). Turner syndrome clinical characteristics include growth disorders, reproductive system and cardiovascular abnormalities and autoimmune diseases (Cui et al. 2018). From a POE perspective, the phenotype varies depending on whether the single X chromosome has been maternally or paternally inherited (Donnelly et al. 2000; Skuse et al. 1997). Turner syndrome girls with a paternally derived X chromosome were shown to be more socially adept than girls with a maternally derived chromosome (Skuse et al. 1997). This led to Skuse et al. (1997) hypothesising the presence of a maternally imprinted genetic locus for social cognition on the X chromosome, which was further supported by Good et al. (Good et al. 2003; Skuse et al. 1997). Given the higher prevalence of ASD in males, and as they do not inherit the paternal X chromosome, Skuse has also speculated that a paternally expressed locus may be present on the X chromosome that gives a protective effect against ASD (Skuse 2000).
Mitochondrial DNA mutations
While mtDNA mutations are mainly associated with classical mitochondrial diseases (MELAS syndrome, MERRF syndrome, Leigh syndrome, Leber’s hereditary optic neuropathy, etc.), the majority of these diseases have some form of neurological, neurodevelopmental or psychiatric component, not surprising considering the energy demand of both the central nervous system and the brain (Bressan and Kramer 2021; C. S. Dela Cruz and Kang 2018). Studies of biochemical markers of abnormal aerobic respiration, such as elevated lactate levels, have provided indirect evidence of mitochondrial dysfunction in ASD (Correia et al. 2006; Žigman et al. 2021). Cohorts of individuals with mitochondrial disease have also been shown to have increased risk of ASD in comparison to the general population (10–20% rates of ASD compared to 2% in the general population), and the reverse is also true; individuals with ASD have been shown to have increased rates of mitochondrial disease compared to the general population (1–5% rates of mitochondrial disease compared to 0.05% in the general population) (Legido et al. 2013). Together, these findings suggest a real relationship between ASD and mitochondrial diseases. Interestingly, some imprinted genes also appear to affect mitochondrial function (Bressan and Kramer 2021; Panov et al. 2020; Urraca et al. 2013; Victor et al. 2021; Yazdi et al. 2013).
A systematic review by Cruz et al. (2019) identified a number of studies showing that genetic variations in mtDNA are associated with neurological disorders, including neurodevelopmental disorders such as ASD. A whole-exome sequencing study of ASD-affected individuals, together with their unaffected siblings and mothers, showed that the transmission of mtDNA point mutations of suspected high pathogenicity was greater between mothers and affected children than between mothers and unaffected siblings, with a higher frequency of these mutations in the ASD probands with lower Intelligence quotient (IQ) and/or deficit in social behaviour (Wang et al. 2016). Two mtDNA genes, MT-ND5 and MT-ATP6, have been linked to ASD through a whole-exome sequencing study of ten multiplex families (Patowary et al. 2017). This builds on other research that identified mutations in the MT-ATP6 gene as linked to ASD (Piryaei et al. 2012; Rossi et al. 2017). A mutation of the MT-TK gene was found to be associated with members of a family (mother and three children) with multiple neurological disorders, including a boy with an autism-like phenotype and was suggested as the basis for his ASD (Graf et al. 2000). For the MT-TL1 mtDNA gene (associated with mitochondrial encephalopathy, lactic acidosis and stroke-like episodes (MELAS) syndrome which can coexist with ASD (Griffiths and Levy 2017)), two mutations have been potentially linked to ASD: A3243G in five mother–offspring pairs (Pons et al. 2004) and A3260G in a single family (Connolly et al. 2010). In an alternative approach, Chalkia et al. examined mitochondrial lineages (ASD-affected individuals, their parents and siblings) which encompass ancient mtDNA functional polymorphisms for association with ASD risk (Chalkia et al. 2017). They found evidence that particular European, Asian and Native American haplogroups showed a significant increase in risk for ASD when compared with the most common European haplogroup.
Maternal genetic effects
Johnson et al. identified a maternal genetic effect in mothers of ASD offspring for the HLA-DR4 (chromosome 6p21.3) allele, suggesting that this finding supports the possibility of an immune component to ASD acting during pregnancy (Johnson et al. 2009). An over-transmission of the GSTP1*A haplotype on chromosome 11q13.2 to mothers of ASD offspring was identified by Williams et al. 2007 (Williams et al. 2007). GSTP1 is associated with oxidative stress, furthering the evidence for inflammation in the intrauterine environment impacting on ASD risk. SHANK3 (chromosome 22q13.33) encodes a synaptic scaffold protein, essential in the postsynaptic density, and several studies have found associations with ASD (Boccuto et al. 2013; Durand et al. 2007; Leblond et al. 2014; Sanders et al. 2015). Connolly et al. presented evidence to suggest that a mutation in the mother’s SHANK3 gene could increase the likelihood of her offspring having ASD (Connolly et al. 2017). The folate pathway supports the change between cell proliferation and differentiation during the early stages of development. If the availability of folate derivatives in the intrauterine environment is altered, neurodevelopment can be disrupted (James et al. 2010). Through a case–control analysis followed by a trio design, James et al. found that the maternal genotype carrying a functional polymorphism in RFC1 (involved in folate metabolism, chromosome 21) was associated with an increased risk of ASD, whereas the offspring genotype was not (James et al. 2010).
Discussion
The aetiology of ASD is driven by a combination of environmental and genetic factors (Searles Quick et al. 2021), though a large proportion of the genetic risk remains unexplained. To improve our understanding of ASD pathogenesis, we need to investigate alternative models of inheritance. POEs are good examples of such alternative forms that should be pursued to help explain some of the missing heritability of this complex disorder. Despite this, it remains an under-studied branch of ASD research.
In this review, we have described different mechanisms of POEs and presented a range of evidence for their role in ASD. We have summarised evidence supporting both maternal and paternal imprinting at the 7q32.1–32.3 and 15q11-13 chromosomal regions, related to both ASD and the syndromic forms of ASD, Angelman and Prader-Willi syndromes, showing how crucial the imprinted brain is for the developmental and behaviour phenotypes associated with ASD. Indeed, a recent study has shown that both imprinting and brain development correlate with ASD (Li et al. 2018); however, increasing sample size and/or replicating in many scenarios is not feasible. The recent generation of whole genome and whole-exome sequencing data from large ASD cohorts, such as the studies by Iossifov et al. and Brandler et al., provides an even greater opportunity to identify rare ASD variants which may have parent-of-origin effects; however, care must be taken to ensure the right statistical tests are performed to identify POEs in these studies (Brandler et al. 2018; S. Connolly and Heron 2014; Iossifov et al. 2014).
We see in this review that POEs play a role in several syndromic forms of ASD (Angelman, Prader-Willi and Fragile X syndromes). Although syndromic forms account for only a small proportion of ASD cases (~ 5%), the biological insights gained from studying syndromic ASD may offer avenues for the understanding of non-syndromic ASD (Sztainberg and Zoghbi 2016). However, care needs to be taken with identifying ASD comorbidity, as syndromic forms of ASD often have different developmental trajectories from non-syndromic ASD (Sztainberg and Zoghbi 2016). Furthermore, care needs to be applied in the choice of ASD diagnostic tool, as diagnosing ASD when the mental age is low is difficult with standard tools (Lord et al. 2000; Miller et al. 2019). In the case of Angelman syndrome for example, Hogart et al. note that the comorbidity studies should be interpreted with caution due to the severity of the cognitive and language impairments and the low mental age range which could be resulting in an over-estimate of ASD comorbidity in Angelman syndrome (Hogart et al. 2010; Trillingsgaard and Østergaard 2004). The assumption here is also that the same set of genes identified in these syndromes are also involved in the comorbid ASD. When IQ or mental age is low, there is the possibility that the ID could be mimicking aspects of the ASD phenotype (Grafodatskaya et al. 2010).
As is clear from the evidence presented here, determining POEs in ASD is an evolving field. For example, with regard to imprinting, it is not a simple case of taking a list of imprinted genes and determining if ASD is linked or associated with these genes. Rather, the imprinting status for a number of these genes is also an evolving area of research. The temporal- and tissue-specific nature of imprinting poses challenges for determining whether or not a gene is imprinted. Authors have also noted and demonstrated that imprinting and maternal genetic effects are known to mimic each other, making determination of the POE type more difficult (Connolly and Heron 2014; Wolf and Wade 2009).
To conclude, the evidence reviewed here converges on the role of ASD POEs in brain development and brain functioning. In particular, a number of the POEs show involvement with synaptic functioning (for example, mutations in FMR1, UBE3A, SHANK3, mtDNA). This is an exciting and expanding field of research, linking together what, on the surface, appear to be quite disparate mechanisms. It is curious that such distinct mechanisms all appear to converge on similar developmental and functional pathways, leading to similar etiological outcomes/presentations. Given the early stages of our understanding of POEs in ASD, we are yet to fully feel their implications in the clinical setting. No doubt as genomic technologies develop, including, for example, technologies that allow for epigenetic profiling, diagnostic testing for POEs will become more straightforward which has the potential to impact diagnostic testing in ASD. In addition, POEs offer another avenue to further tease apart the complex nature of the ASD phenotype with the ultimate goal being to identify usable drug targets and biomarkers that will have tangible impacts for ASD in terms of treatments and interventions. One current example of this is Nr4a1, which has been proposed as a possible drug target for SNRPN-related neurodevelopment disorders, including Prader-Willi syndrome and ASD (H. Li et al. 2016) and is already under consideration as a drug target in cancer (Hedrick et al. 2015; S. O. Lee et al. 2014). We are interested to see how the field develops in the future and how our understanding of ASD and related disorders will change with a better understanding of the contributions of POEs to ASD.
Data availability
As this is a review article no data were analysed.
References
Abbeduto L, McDuffie A, Thurman AJ (2014) The fragile X syndrome-autism comorbidity: what do we really know? Front Genet 5:355. https://doi.org/10.3389/fgene.2014.00355
Abramowitz LK, Bartolomei MS (2012) Genomic imprinting: recognition and marking of imprinted loci. Curr Opin Genet Dev 22(2):72–78. https://doi.org/10.1016/j.gde.2011.12.001
Alarcón M, Abrahams BS, Stone JL, Duvall JA, Perederiy JV, Bomar JM, Sebat J, Wigler M, Martin CL, Ledbetter DH, Nelson SF, Cantor RM, Geschwind DH (2008) Linkage, association, and gene-expression analyses identify CNTNAP2 as an autism-susceptibility gene. Am J Hum Genet 82(1):150–159. https://doi.org/10.1016/j.ajhg.2007.09.005
Allen ND, Logan K, Lally G, Drage DJ, Norris ML, Keyerne EB (1995) Distribution of parthenogenetic cells in the mouse brain and their influence on brain development and behavior. Proc Natl Acad Sci USA 92(23):10782–10786. https://doi.org/10.1073/pnas.92.23.10782
American Psychiatric Association (2013) DSM 5. In Diagnostic and Statistical Manual of Mental Disorders 11(2):1–992. American Psychiatric Association https://doi.org/10.1176/appi.books.9780890425596.dsm05
Anney R, Klei L, Pinto D, Almeida J, Bacchelli E, Baird G, Bolshakova N, Bölte S, Bolton PF, Bourgeron T, Brennan S, Brian J, Casey J, Conroy J, Correia C, Corsello C, Crawford EL, de Jonge M, Delorme R, Duketis E, Duque F, Estes A, Farrar P, Fernandez BA, Folstein SE, Fombonne E, Gilbert J, Gillberg C, Glessner JT, Green A, Green J, Guter SJ, Heron EA, Holt R, Howe JL, Hughes G, Hus V, Igliozzi R, Jacob S, Kenny GP, Kim C, Kolevzon A, Kustanovich V, Lajonchere CM, Lamb JA, Law-Smith M, Leboyer M, Le Couteur A, Leventhal BL, Liu XQ, Lombard F, Lord C, Lotspeich L, Lund SC, Magalhaes TR, Mantoulan C, McDougle CJ, Melhem NM, Merikangas A, Minshew NJ, Mirza GK, Munson J, Noakes C, Nygren G, Papanikolaou K, Pagnamenta AT, Parrini B, Paton T, Pickles A, Posey DJ, Poustka F, Ragoussis J, Regan R, Roberts W, Roeder K, Roge B, Rutter ML, Schlitt S, Shah N, Sheffield VC, Soorya L, Sousa I, Stoppioni V, Sykes N, Tancredi R, Thompson AP, Thomson S, Tryfon A, Tsiantis J, Van Engeland H, Vincent JB, Volkmar F, Vorstman JA, Wallace S, Wing K, Wittemeyer K, Wood S, Zurawiecki D, Zwaigenbaum L, Bailey AJ, Battaglia A, Cantor RM, Coon H, Cuccaro ML, Dawson G, Ennis S, Freitag CM, Geschwind DH, Haines JL, Klauck SM, McMahon WM, Maestrini E, Miller J, Monaco AP, Nelson SF, Nurnberger JI Jr, Oliveira G, Parr JR, Pericak-Vance MA, Piven J, Schellenberg GD, Scherer SW, Vicente AM, Wassink TH, Wijsman EM, Betancur C, Buxbaum JD, Cook EH, Gallagher L, Gill M, Hallmayer J, Paterson AD, Sutcliffe JS, Szatmari P, Vieland VJ, Hakonarson H, Devlin B (2012) Individual common variants exert weak effects on the risk for autism spectrum disorders. Hum Mol Genet 21(21):4781–92. https://doi.org/10.1093/hmg/dds301
Arking DE, Cutler DJ, Brune CW, Teslovich TM, West K, Ikeda M, Rea A, Guy M, Lin S, Cook EH, Chakravarti A (2008) A common genetic variant in the neurexin superfamily member CNTNAP2 increases familial risk of autism. Am J Hum Genet 82(1):160–164. https://doi.org/10.1016/j.ajhg.2007.09.015
Ashley-Koch A, Wolpert CM, Menold MM, Zaeem L, Basu S, Donnelly SL, Ravan SA, Powell CM, Qumsiyeh MB, Aylsworth AS, Vance JM, Gilbert JR, Wright HH, Abramson RK, Delong GR, Cuccaro ML, Pericak-Vance MA (1999) Genetic studies of autistic disorder and chromosome 7. Genomics 61(3):227–236. https://doi.org/10.1006/geno.1999.5968
Azzini E, Ruggeri S, Polito A (2020) Homocysteine: its possible emerging role in at-risk population groups. Int J Mol Sci 21(4):1421. https://doi.org/10.3390/ijms21041421
Badcock C, Crespi B (2006) Imbalanced genomic imprinting in brain development: an evolutionary basis for the aetiology of autism. J Evol Biol 19(4):1007–1032. https://doi.org/10.1111/j.1420-9101.2006.01091.x
Badcock C, Crespi B (2008) Battle of the sexes may set the brain. Nature 454(7208):1054–1055. https://doi.org/10.1038/4541054a. (Nature Publishing Group)
Bai D, Yip BHK, Windham GC, Sourander A, Francis R, Yoffe R, Glasson E, Mahjani B, Suominen A, Leonard H, Gissler M, Buxbaum JD, Wong K, Schendel D, Kodesh A, Breshnahan M, Levine SZ, Parner ET, Hansen SN, Hultman C, Reichenberg A, Sandin S (2019) Association of genetic and environmental factors with autism in a 5-country cohort. JAMA Psychiat 76(10):1035–1043. https://doi.org/10.1001/jamapsychiatry.2019.1411
Bakkaloglu B, O’Roak BJ, Louvi A, Gupta AR, Abelson JF, Morgan TM, Chawarska K, Klin A, Ercan-Sencicek AG, Stillman AA, Tanriover G, Abrahams BS, Duvall JA, Robbins EM, Geschwind DH, Biederer T, Gunel M, Lifton RP, State MW (2008) Molecular cytogenetic analysis and resequencing of contactin associated protein-like 2 in autism spectrum disorders. Am J Hum Genet 82(1):165–173. https://doi.org/10.1016/j.ajhg.2007.09.017
Barr JA, Jones J, Glenister PH, Cattanach BM (1995) Ubiquitous expression and imprinting of Snrpn in the mouse. Mamm Genome 6(6):405–407. https://doi.org/10.1007/BF00355641
Betancur C, Sakurai T, Buxbaum JD (2009) The emerging role of synaptic cell-adhesion pathways in the pathogenesis of autism spectrum disorders. Trends Neurosci 32(7):402–412. https://doi.org/10.1016/j.tins.2009.04.003
Boccuto L, Lauri M, Sarasua SM, Skinner CD, Buccella D, Dwivedi A, Orteschi D, Collins JS, Zollino M, Visconti P, Dupont B, Tiziano D, Schroer RJ, Neri G, Stevenson RE, Gurrieri F, Schwartz CE (2013) Prevalence of SHANK3 variants in patients with different subtypes of autism spectrum disorders. Eur J Hum Genet 21(3):310–316. https://doi.org/10.1038/ejhg.2012.175
Bolton PF, Dennis NR, Browne CE, Thomas NS, Veltman MWM, Thompson RJ, Jacobs P (2001) The phenotypic manifestations of interstitial duplications of proximal 15q with special reference to the autistic spectrum disorders. Am J Med Genet 105(8):675–685. https://doi.org/10.1002/ajmg.1551
Bonora E, Graziano C, Minopoli F, Bacchelli E, Magini P, Diquigiovanni C, Lomartire S, Bianco F, Vargiolu M, Parchi P, Marasco E, Mantovani V, Rampoldi L, Trudu M, Parmeggiani A, Battaglia A, Mazzone L, Tortora G, Maestrini E, Seri M, Romeo G (2014) Maternally inherited genetic variants of CADPS 2 are present in autism spectrum disorders and intellectual disability patients. EMBO Mol Med 6(12):1639–1639. https://doi.org/10.15252/emmm.201404829
Boyar FZ, Whitney MM, Lossie AC, Gray BA, Keller KL, Stalker HJ, Zori RT, Geffken G, Mutch J, Edge PJ, Voeller KS, Williams CA, Driscoll DJ (2001) A family with a grand-maternally derived interstitial duplication of proximal 15q. Clin Genet 60(6):421–430. https://doi.org/10.1034/j.1399-0004.2001.600604.x
Brandler WM, Antaki D, Gujral M, Kleiber ML, Whitney J, Maile MS, Hong O, Chapman TR, Tan S, Tandon P, Pang T, Tang SC, Vaux KK, Yang Y, Harrington E, Juul S, Turner DJ, Thiruvahindrapuram B, Kaur G, Wang Z, Kingsmore SF, Gleeson JG, Bisson D, Kakaradov B, Telenti A, Venter JC, Corominas R, Toma C, Cormand B, Rueda I, Guijarro S, Messer KS, Nievergelt CM, Arranz MJ, Courchesne E, Pierce K, Muotri AR, Iakoucheva LM, Hervas A, Scherer SW, Corsello C, Sebat J (2018) Paternally inherited cis-regulatory structural variants are associated with autism. Science 360(6386):327–331. https://doi.org/10.1126/science.aan2261
Bressan P, Kramer P (2021) Mental Health, Mitochondria, and the Battle of the Sexes. Biomedicines 9(2):116. https://doi.org/10.3390/biomedicines9020116
Browne CE, Dennis NR, Maher E, Long FL, Nicholson JC, Sillibourne J, Barber JCK (1997) Inherited interstitial duplications of proximal 15q: Genotype-phenotype correlations. Am J Hum Genet 61(6):1342–1352. https://doi.org/10.1086/301624
Byars SG, Stearns SC, Boomsma JJ (2014) Opposite risk patterns for autism and schizophrenia are associated with normal variation in birth size: phenotypic support for hypothesized diametric gene-dosage effects. Proc Biol Sci 281(1794):20140604. https://doi.org/10.1098/rspb.2014.0604
Carter M, Scherer S (2013) Autism spectrum disorder in the genetics clinic: a review. Clin Genet 83(5):399–407. https://doi.org/10.1111/cge.12101
Cassidy SB, Dykens EM, Williams CA (2000) Prader-Willi and Angelman syndromes: sister imprinted disorders. Am J Med Genet Semin Med Genet 97(2):136–146. https://doi.org/10.1002/1096-8628(200022)97:2%3c136::AID-AJMG5%3e3.0.CO;2-V
Cassidy SB, Schwartz S, Miller JL, Driscoll DJ (2012) Prader-Willi syndrome. Genet Med 14(1):10–26. https://doi.org/10.1038/gim.0b013e31822bead0. (Nature Publishing Group)
Chalkia D, Singh LN, Leipzig J, Lvova M, Derbeneva O, Lakatos A, Hadley D, Hakonarson H, Wallace DC (2017) Association between mitochondrial DNA haplogroup variation and autism spectrum disorders. JAMA Psychiat 74(11):1161–1168. https://doi.org/10.1001/jamapsychiatry.2017.2604
Chaste P, Sanders SJ, Mohan KN, Klei L, Murtha MT, Hus V, Lowe JK, Willsey AJ, Yu TW, Fombonne E, Geschwind D, Dorothy E, Morrow EM, Walsh CA, Sutcliffe JS, State MW (2014) Modest impact on risk for autism spectrum disorder of rare copy number variants at 15q11.2, specifically breakpoints 1 to 2. Autism Res 7(3):355–362. https://doi.org/10.1002/aur.1378.Modest
Chen S, Li Z, He Y, Zhang F, Li H, Liao Y, Wei Z, Wan G, **ang X, Hu M, **a K, Chen X, Tang J (2015) Elevated mitochondrial DNA copy number in peripheral blood cells is associated with childhood autism. BMC Psychiatry 15:50. https://doi.org/10.1186/s12888-015-0432-y
Chiocchetti AG, Kopp M, Waltes R, Haslinger D, Duketis E, Jarczok TA, Poustka F, Voran A, Graab U, Meyer J, Klauck SM, Fulda S, Freitag CM (2015) Variants of the CNTNAP2 5’ promoter as risk factors for autism spectrum disorders: a genetic and functional approach. Mol Psychiatry 20(7):839–849. https://doi.org/10.1038/mp.2014.103
Connolly BS, Feigenbaum ASJ, Robinson BH, Dipchand AI, Simon DK, Tarnopolsky MA (2010) MELAS syndrome, cardiomyopathy, rhabdomyolysis, and autism associated with the A3260G mitochondrial DNA mutation. Biochem Biophys Res Commun 402(2):443–447. https://doi.org/10.1016/j.bbrc.2010.10.060
Connolly S, Anney R, Gallagher L et al (2017) A genome-wide investigation into parent-of-origin effects in autism spectrum disorder identifies previously associated genes including SHANK3. Eur J Hum Genet 25:234–239. https://doi.org/10.1038/ejhg.2016.153
Connolly S, Heron EA (2014) Review of statistical methodologies for the detection of parent-of-origin effects in family trio genome-wide association data with binary disease traits. Brief Bioinform 16(3):429–448. https://doi.org/10.1093/bib/bbu017
Constância M, Hemberger M, Hughes J, Dean W, Ferguson-Smith A, Fundele R, Stewart F, Kelsey G, Fowden A, Sibley C, Reik W (2002) Placental-specific IGF-II is a major modulator of placental and fetal growth. Nature 417(6892):945–948.https://doi.org/10.1038/nature00819
Cook EH, Courchesne RY, Cox NJ, Lord C, Gonen D, Guter SJ, Lincoln A, Nix K, Haas R, Leventhal BL, Courchesne E (1998) Linkage-disequilibrium map** of autistic disorder, with 15q11-13 markers. Am J Hum Genet 62(5):1077–1083. https://doi.org/10.1086/301832
Cook EH, Lindgren V, Leventhal BL, Courchesne R, Lincoln A, Shulman C, Lord C, Courchesne E (1997) Autism or atypical autism in maternally but not paternally derived proximal 15q duplication. Am J Hum Genet 60(4):928–934
Correia C, Coutinho AM, Diogo L, Grazina M, Marques C, Miguel T, Ataíde A, Almeida J, Borges L, Oliveira C, Oliveira G, Vicente AM (2006) Brief report: high frequency of biochemical markers for mitochondrial dysfunction in autism: no association with the mitochondrial aspartate/glutamate carrier SLC25A12 gene. J Autism Dev Disord 36(8):1137–1140. https://doi.org/10.1007/s10803-006-0138-6
Court F, Tayama C, Romanelli V, Martin-Trujillo A, Iglesias-Platas I, Okamura K, Sugahara N, Simón C, Moore H, Harness JV, Keirstead H, Sanchez-Mut JV, Kaneki E, Lapunzina P, Soejima H, Wake N, Esteller M, Ogata T, Hata K, Nakabayashi K, Monk D (2014) Genome-wide parent-of-origin DNA methylation analysis reveals the intricacies of human imprinting and suggests a germline methylation-independent mechanism of establishment. Genome Res 24(4):554–569. https://doi.org/10.1101/gr.164913.113
Crespi B (2013) Diametric gene-dosage effects as windows into neurogenetic architecture. Curr Opin Neurobiol 23(1):143–151. https://doi.org/10.1016/j.conb.2012.08.005. (Elsevier Current Trends)
Crespi B, Badcock C (2008) Psychosis and autism as diametrical disorders of the social brain. Behav Brain Sci 31(3):241–320. https://doi.org/10.1017/S0140525X08004214
Creswell CS, Skuse DH (1999) Autism in association with Turner syndrome: genetic implications for male vulnerability to pervasive developmental disorders. Neurocase 5(6):511–518. https://doi.org/10.1080/13554799908402746
Cruz ACP, Ferrasa A, Muotri AR, Herai RH (2019) Frequency and association of mitochondrial genetic variants with neurological disorders. Mitochondrion 46:345–360. https://doi.org/10.1016/j.mito.2018.09.005. (Elsevier B.V)
Cui X, Cui Y, Shi L, Luan J, Zhou X, Han J (2018) A basic understanding of Turner syndrome: incidence, complications, diagnosis, and treatment. Intractable Rare Dis Res 7(4):223–228. https://doi.org/10.5582/irdr.2017.01056. (International Advancement Center for Medicine and Health Research)
Dagli AI, Williams CA (2017) NCBI Bookshelf - GeneReviewsTM. Angelman Syndrome. Scielo 2:2–5. http://www.ncbi.nlm.nih.gov/books/NBK1144/. Accessed Aug 2022
Davies W, Isles AR, Wilkinson LS (2001) Imprinted genes and mental dysfunction. Ann Med 33(6):428–436. https://doi.org/10.3109/07853890108995956
Davies W, Isles AR, Wilkinson LS (2005) Imprinted gene expression in the brain. Neurosci Biobehav Rev 29(3):421–430. https://doi.org/10.1016/j.neubiorev.2004.11.007
Dela Cruz CS, Kang MJ (2018) Mitochondrial dysfunction and damage associated molecular patterns (DAMPs) in chronic inflammatory diseases. Mitochondrion 41:37–44. https://doi.org/10.1016/j.mito.2017.12.001. (Elsevier B.V)
Depienne C, Moreno-De-Luca D, Heron D, Bouteiller D, Gennetier A, Delorme R, Chaste P, Siffroi JP, Chantot-Bastaraud S, Benyahia B, Trouillard O, Nygren G, Kopp S, Johansson M, Rastam M, Burglen L, Leguern E, Verloes A, Leboyer M, Brice A, Gillberg C, Betancur C (2009) Screening for genomic rearrangements and methylation abnormalities of the 15q11-q13 region in autism spectrum disorders. Biol Psychiat 66(4):349–359. https://doi.org/10.1016/j.biopsych.2009.01.025
Donnelly SL, Wolpert CM, Menold MM, Bass MP, Gilbert JR, Cuccaro ML, DeLong GR, Pericak-Vance MA (2000) Female with autistic disorder and monosomy X (Turner syndrome): Parent-of-origin effect of the X chromosome. Am J Med Genet Neuropsychiatr Genet 96(3):312–316. https://doi.org/10.1002/1096-8628(20000612)96:3%3c312::AID-AJMG16%3e3.0.CO;2-8
Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F, Nygren G, Rastam M, Gillberg IC, Anckarsäter H, Sponheim E, Goubran-Botros H, Delorme R, Chabane N, Mouren-Simeoni MC, De Mas P, Bieth E, Rogé B, Héron D, Burglen L, Gillberg C, Leboyer M, Bourgeron T (2007) Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 39(1):25–27. https://doi.org/10.1038/ng1933
Dykens EM, Lee E, Roof E (2011) Prader-Willi syndrome and autism spectrum disorders: an evolving story. J Neurodev Disord 3(3):225–237. https://doi.org/10.1007/s11689-011-9092-5
Dykens EM, Roof E, Hunt-Hawkins H, Dankner N, Lee EB, Shivers CM, Daniell C, Kim SJ (2017) Diagnoses and characteristics of autism spectrum disorders in children with Prader-Willi syndrome. J Neurodev Disord 9(1):18. https://doi.org/10.1186/s11689-017-9200-2
Edmiston E, Ashwood P, Van de Water J (2017) Autoimmunity, autoantibodies, and autism spectrum disorder. Biol Psychiatry 81(5):383–390. Elsevier USA. https://doi.org/10.1016/j.biopsych.2016.08.031
Eggermann T, Begemann M, Binder G, Spengler S (2010) Silver-Russell syndrome: genetic basis and molecular genetic testing. Orphanet J Rare Dis 5(1):19. https://doi.org/10.1186/1750-1172-5-19. (BioMed Central)
Falls JG, Pulford DJ, Wylie AA, Jirtle RL (1999) Genomic imprinting: implications for human disease. Am J Pathol 154(3):635–647. https://doi.org/10.1016/S0002-9440(10)65309-6. (American Society for Investigative Pathology Inc)
Fernandez BA, Scherer SW (2017) Syndromic autism spectrum disorders: moving from a clinically defined to a molecularly defined approach. www.dialogues-cns.org. Accessed Dec 2022
Finucane BM, Lusk L, Arkilo D, Chamberlain S, Devinsky O, Dindot S, Jeste SS, LaSalle JM, Reiter LT, Schanen NC, Spence SJ (2016) 15q duplication syndrome and related disorders. GeneReviews 1993–2019
Giulivi C, Zhang YF, Omanska-Klusek A, Ross-Inta C, Wong S, Hertz-Picciotto I, Tassone F, Pessah IN (2010) Mitochondrial dysfunction in autism. JAMA J Am Med Assoc 304(21):2389–2396. https://doi.org/10.1001/jama.2010.1706
Good CD, Lawrence K, Simon Thomas N, Price CJ, Ashburner J, Friston KJ, Frackowiak RSJ, Oreland L, Skuse DH, Skuse D (2003) Dosage-sensitive X-linked locus influences the development of amygdala and orbitofrontal cortex, and fear recognition in humans. Brain 126:2431–2446. https://doi.org/10.1093/brain/awg242
Graf WD, Marin-Garcia J, Gao HG, Pizzo S, Naviaux RK, Markusic D, Barshop BA, Courchesne E, Haas RH (2000) Autism associated with the mitochondrial DNA G8363A transfer RNA(Lys) mutation. J Child Neurol 15(6):357–361. https://doi.org/10.1177/088307380001500601
Grafodatskaya D, Chung B, Szatmari P, Weksberg R (2010) Autism spectrum disorders and epigenetics. J Am Acad Child Adolesc Psychiatry 49(8):794–809. https://doi.org/10.1016/j.jaac.2010.05.005. (Elsevier)
Greer PL, Hanayama R, Bloodgood BL, Mardinly AR, Lipton DM, Flavell SW, Kim TK, Griffith EC, Waldon Z, Maehr R, Ploegh HL, Chowdhury S, Worley PF, Steen J, Greenberg ME (2010) The Angelman syndrome protein Ube3A regulates synapse development by ubiquitinating arc. Cell 140(5):704–716. https://doi.org/10.1016/j.cell.2010.01.026
Griffiths KK, Levy RJ (2017) Evidence of mitochondrial dysfunction in autism: biochemical links, genetic-based associations, and non-energy-related mechanisms. Oxid Med Cell Longev 2017:4314025. https://doi.org/10.1155/2017/4314025
Guffanti G, Lievers LS, Bonati MT, Marchi M, Geronazzo L, Nardocci N, Estienne M, Larizza L, Macciardi F, Russo S (2011) Role of UBE3A and ATP10A genes in autism susceptibility region 15q11-q13 in an Italian population: a positive replication for UBE3A. Psychiatry Res 185(1–2):33–38. https://doi.org/10.1016/j.psychres.2010.04.057
Guilmatre A, Sharp AJ (2012) Parent of origin effects. Clin Genet 81(3):201–209. https://doi.org/10.1111/j.1399-0004.2011.01790.x
Hedrick E, Lee SO, Kim G, Abdelrahim M, ** UH, Safe S, Abudayyeh A (2015) Nuclear receptor 4A1 (NR4A1) as a drug target for renal cell adenocarcinoma. PLoS One 10(6):e0128308. https://doi.org/10.1371/journal.pone.0128308
Herzing LBK (2002) Allele-specific expression analysis by RNA-FISH demonstrates preferential maternal expression of UBE3A and imprint maintenance within 15q11- q13 duplications. Hum Mol Genet 11(15):1707–1718. https://doi.org/10.1093/hmg/11.15.1707
Hodges H, Fealko C, Soares N (2020) Autism spectrum disorder: definition, epidemiology, causes, and clinical evaluation. Transl Pediatr 9(Suppl 1):S55–S65. AME Publishing Company. https://doi.org/10.21037/tp.2019.09.09
Hogart A, Leung KN, Wang NJ, Wu DJ, Driscoll J, Vallero RO, Schanen NC, LaSalle JM (2009) Chromosome 15q11-13 duplication syndrome brain reveals epigenetic alterations in gene expression not predicted from copy number. J Med Genet 46(2):86–93. https://doi.org/10.1136/jmg.2008.061580
Hogart A, Nagarajan RP, Patzel KA, Yasui DH, LaSalle JM (2007) 15q11-13 GABAA receptor genes are normally biallelically expressed in brain yet are subject to epigenetic dysregulation in autism-spectrum disorders. Hum Mol Genet 16(6):691–703. https://doi.org/10.1093/hmg/ddm014
Hogart A, Wu D, LaSalle JM, Schanen NC (2010) The comorbidity of autism with the genomic disorders of chromosome 15q11.2–q13. In Neurobiol Dis 38(2):181–191. https://doi.org/10.1016/j.nbd.2008.08.011
Hsiao JS, Germain ND, Wilderman A, Stoddard C, Wojenski LA, Villafano GJ, Core L, Cotney J, Chamberlain SJ (2019) A bipartite boundary element restricts UBE3A imprinting to mature neurons. Proc Natl Acad Sci USA 116(6):2181–2186. https://doi.org/10.1073/pnas.1815279116
Hutter B, Bieg M, Helms V, Paulsen M (2010) Divergence of imprinted genes during mammalian evolution. BMC Evol Biol 10(1):116. https://doi.org/10.1186/1471-2148-10-116
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, Stessman HA, Witherspoon KT, Vives L, Patterson KE, Smith JD, Paeper B, Nickerson DA, Dea J, Dong S, Gonzalez LE, Mandell JD, Mane SM, Murtha MT, Sullivan CA, Walker MF, Waqar Z, Wei L, Willsey AJ, Yamrom B, Lee YH, Grabowska E, Dalkic E, Wang Z, Marks S, Andrews P, Leotta A, Kendall J, Hakker I, Rosenbaum J, Ma B, Rodgers L, Troge J, Narzisi G, Yoon S, Schatz MC, Ye K, McCombie WR, Shendure J, Eichler EE, State MW, Wigler M (2014) The contribution of de novo coding mutations to autism spectrum disorder. Nature 515(7526):216–221. https://doi.org/10.1038/nature13908
Isles AR, Wilkinson LS (2000) Imprinted genes, cognition and behaviour. Trends Cogn Sci 4(8):309–318. https://doi.org/10.1016/S1364-6613(00)01504-7
James SJ, Melnyk S, Jernigan S, Pavliv O, Trusty T, Lehman S, Seidel L, Gaylor DW, Cleves MA (2010) A functional polymorphism in the reduced folate carrier gene and DNA hypomethylation in mothers of children with autism. Am J Med Genet B Neuropsychiatr Genet 153(6):1209–1220. https://doi.org/10.1002/ajmg.b.31094
Johnson WG (2003) Teratogenic alleles and neurodevelopmental disorders. BioEssays 25(5):464–477. https://doi.org/10.1002/bies.10268
Johnson WG, Buyske S, Mars AE, Sreenath M, Stenroos ES, Williams TA, Stein R, Lambert GH (2009) HLA-DR4 as a risk allele for autism acting in mothers of probands possibly during pregnancy. Arch Pediatr Adolesc Med 163(6):542–546. https://doi.org/10.1001/archpediatrics.2009.74
Kaufmann WE, Kidd SA, Andrews HF, Budimirovic DB, Esler A, Haas-Givler B, Stackhouse T, Riley C, Peacock G, Sherman SL, Brown WT, Berry-Kravis E (2017) Autism spectrum disorder in Fragile X syndrome: cooccurring conditions and current treatment. Pediatrics 139(Suppl 3):S194–S206. https://doi.org/10.1542/peds.2016-1159F
Kent L, Bowdin S, Kirby GA, Cooper WN, Maher ER (2008) Beckwith Weidemann syndrome: a behavioral phenotype-genotype study. Am J Med Genet B Neuropsychiatr Genet 147(7):1295–1297. https://doi.org/10.1002/ajmg.b.30729
Keverne EB, Fundele R, Narasimha M, Barton SC, Surani MA (1996) Genomic imprinting and the differential roles of parental genomes in brain development. Dev Brain Res 92(1):91–100.https://doi.org/10.1016/0165-3806(95)00209-X
Kidd SA, Berry-Kravis E, Choo TH, Chen C, Esler A, Hoffmann A, Andrews HF, Kaufmann WE (2019) Improving the diagnosis of autism spectrum disorder in Fragile X syndrome by adapting the social communication questionnaire and the social responsiveness scale-2. J Autism Dev Disord 50(9):3276–3295. https://doi.org/10.1007/s10803-019-04148-0
Kim SJ, Brune CW, Kistner EO, Christian SL, Courchesne EH, Cox NJ, Cook EH (2008) Transmission disequilibrium testing of the chromosome 15q11-q13 region in autism. Am J Med Genet B Neuropsychiatr Genet 147(7):1116–1125. https://doi.org/10.1002/ajmg.b.30733
Korvatska O, Estes A, Munson J, Dawson G, Bekris LM, Kohen R, Yu C-E, Schellenberg GD, Raskind WH (2011) Mutations in the TSGA14 gene in families with autism spectrum disorders. Am J Med Genet B Neuropsychiatr Genet 156(3):303–311. https://doi.org/10.1002/ajmg.b.31162
Kwack K, Lee SK, Kim M, Nam M, Bang HJ, Yang JW, Choe K, Kim SK, Hong MS, Chung J, Kim HG (2008) Positive association between the mesoderm specific transcript gene and autism spectrum disorder in a Korean male population. FASEB J 22(S1):906.8–906.8. https://doi.org/10.1096/fasebj.22.1_supplement.906.8
Lamb JA, Barnby G, Bonora E, Sykes N, Bacchelli E, Blasi F, Maestrini E, Broxholme J, Tzenova J, Weeks D, Bailey AJ, Monaco AP (2005) Analysis of IMGSAC autism susceptibility loci: evidence for sex limited and parent of origin specific effects. J Med Genet 42(2):132–137. https://doi.org/10.1136/jmg.2004.025668
Lawson HA, Cheverud JM, Wolf JB (2013) Genomic imprinting and parent-of-origin effects on complex traits. Nat Rev Genet 14(9):609–617. https://doi.org/10.1038/nrg3543.Genomic
Leblond CS, Nava C, Polge A, Gauthier J, Huguet G, Lumbroso S, Giuliano F, Stordeur C, Depienne C, Mouzat K, Pinto D, Howe J, Lemière N, Durand CM, Guibert J, Ey E, Toro R, Peyre H, Mathieu A, Amsellem F, Rastam M, Gillberg IC, Bourgeron T (2014) Meta-analysis of SHANK mutations in autism spectrum disorders: a gradient of severity in cognitive impairments. PLoS Genet 10(9):1004580. https://doi.org/10.1371/journal.pgen.1004580
Lee IS, Carvalho CMB, Douvaras P, Ho SM, Hartley BJ, Zuccherato LW, Ladran IG, Siegel AJ, McCarthy S, Malhotra D, Sebat J, Rapoport J, Fossati V, Lupski JR, Levy DL, Brennand KJ (2015) Characterization of molecular and cellular phenotypes associated with a heterozygous CNTNAP2 deletion using patient-derived hiPSC neural cells. In npj Schizophrenia 1:15019. https://doi.org/10.1038/npjschz.2015.19. (Nature Publishing Group)
Lee SO, Li X, Hedrick E, ** UH, Tjalkens RB, Backos DS, Li L, Zhang Y, Wu Q, Safe S (2014) Diindolylmethane analogs bind NR4A1 and are NR4A1 antagonists in colon cancer cells. Mol Endocrinol 28(10):1729–1739. https://doi.org/10.1210/me.2014-1102
Lefebvre L, Viville S, Barton SC, Ishino F, Keverne EB, Azim Surani M (1998) Abnormal maternal behaviour and growth retardation associated with loss of the imprinted gene Mest. Nat Genet 20(2):163–169. https://doi.org/10.1038/2464
Legido A, Jethva R, Goldenthal MJ (2013) Mitochondrial dysfunction in autism. Semin Pediatr Neurol 20(3):163–175. https://doi.org/10.1016/j.spen.2013.10.008
Li H, Zhao P, Xu Q, Shan S, Hu C, Qiu Z, Xu X (2016) The autism-related gene SNRPN regulates cortical and spine development via controlling nuclear receptor Nr4a1. Sci Rep 6(1):1–10. https://doi.org/10.1038/srep29878
Li J, Lin X, Wang M, Hu Y, Xue K, Gu S, Lv L, Huang S, **e W (2020) Potential role of genomic imprinted genes and brain developmental related genes in autism. BMC Med Genomics 13(1):54. https://doi.org/10.1186/s12920-020-0693-2
Li X, Hu Z, He Y, **ong Z, Long Z, Peng Y, Bu F, Ling J, Xun G, Mo X, Pan Q, Zhao J, **a K (2010) Association analysis of CNTNAP2 polymorphisms with autism in the Chinese han population. Psychiatr Genet 20(3):113–117. https://doi.org/10.1097/YPG.0b013e32833a216f
Lin M, Hrabovsky A, Pedrosa E, Wang T, Zheng D, Lachman HM (2012) Allele-biased expression in differentiating human neurons: implications for neuropsychiatric disorders. PLoS ONE 7(8):44017. https://doi.org/10.1371/journal.pone.0044017
Lord C, Risi S, Lambrecht L, Cook EH, Leventhal BL, Dilavore PC, Pickles A, Rutter M (2000) The autism diagnostic observation schedule-generic: a standard measure of social and communication deficits associated with the spectrum of autism. J Autism Dev Disord 30(3):205–223. https://doi.org/10.1023/A:1005592401947
Lorgen-Ritchie M, Murray AD, Ferguson-Smith AC, Richards M, Horgan GW, Phillips LH, Hoad G, Gall I, Harrison K, McNeill G, Ito M, Haggarty P (2019) Imprinting methylation in SNRPN and MEST1 in adult blood predicts cognitive ability. PLoS ONE 14(2):e0211799. https://doi.org/10.1371/journal.pone.0211799
Mao R, Jalal SM, Snow K, Michels VV, Szabo SM, Babovic-Vuksanovic D (2000) Characteristics of two cases with dup(15)(q11.2–q12): one of maternal and one of paternal origin. Genet Med 2(2):131–135. https://doi.org/10.1097/00125817-200003000-00003
McFadden K, Minshew NJ (2013) Evidence for dysregulation of axonal growth and guidance in the etiology of ASD. Front Hum Neurosci 7(Oct). Frontiers Media S. A. https://doi.org/10.3389/fnhum.2013.00671
Mertz LGB, Thaulov P, Trillingsgaard A, Christensen R, Vogel I, Hertz JM, Østergaard JR (2014) Neurodevelopmental outcome in Angelman syndrome: genotype-phenotype correlations. Res Dev Disabil 35(7):1742–1747. https://doi.org/10.1016/j.ridd.2014.02.018
Miller LE, Burke JD, Robins DL, Fein DA (2019) Diagnosing autism spectrum disorder in children with low mental age. J Autism Dev Disord 49(3):1080–1095. https://doi.org/10.1007/s10803-018-3810-8
Moore T, Haig D (1991) Genomic imprinting in mammalian development: a parental tug-of-war. Trends Genet 7(2):45–49. https://doi.org/10.1016/0168-9525(91)90230-N
Moreno-De-Luca D, Sanders SJ, Willsey AJ, Mulle JG, Lowe JK, Geschwind DH, State MW, Martin CL, Ledbetter DH (2013) Using large clinical data sets to infer pathogenicity for rare copy number variants in autism cohorts. Mol Psychiatry 18(10):1090–1095. https://doi.org/10.1038/mp.2012.138
Nolin SL, Brown WT, Glicksman A, Houck GE, Gargano AD, Sullivan A, Biancalana V, Bröndum-Nielsen K, Hjalgrim H, Holinski-Feder E, Kooy F, Longshore J, Macpherson J, Mandel JL, Matthijs G, Rousseau F, Steinbach P, Väisänen ML, Von Koskull H, Sherman SL (2003) Expansion of the fragile X CGG repeat in females with premutation or intermediate alleles. Am J Hum Genet 72(2):454–464. https://doi.org/10.1086/367713
Nolin SL, Glicksman A, Tortora N, Allen E, Macpherson J, Mila M, Vianna-Morgante AM, Sherman SL, Dobkin C, Latham GJ, Hadd AG (2019) Expansions and contractions of the FMR1 CGG repeat in 5,508 transmissions of normal, intermediate, and premutation alleles. Am J Med Genet A 179(7):1148–1156. https://doi.org/10.1002/ajmg.a.61165
Nurmi EL, Amin T, Olson LM, Jacobs MM, McCauley JL, Lam AY, Organ EL, Folstein SE, Haines JL, Sutcliffe JS (2003) Dense linkage disequilibrium map** in the 15q11-q13 maternal expression domain yields evidence for association in autism. Mol Psychiatry 8(6):624–634. https://doi.org/10.1038/sj.mp.4001283
Nurmi EL, Bradford Y, Chen YH, Hall J, Arnone B, Gardiner MB, Hutcheson HB, Gilbert JR, Pericak-Vance MA, Copeland-Yates SA, Michaelis RC, Wassink TH, Santangelo SL, Sheffield VC, Piven J, Folstein SE, Haines JL, Sutcliffe JS (2001) Linkage disequilibrium at the Angelman syndrome gene UBE3A in Autism families. Genomics 77(1–2):105–113. https://doi.org/10.1006/geno.2001.6617
Panov J, Simchi L, Feuermann Y, Kaphzan H (2020) Bioinformatics analyses of the transcriptome reveal Ube3a-dependent effects on mitochondrial-related pathways. Int J Mol Sci 21(11):4156. https://doi.org/10.3390/ijms21114156
Partida GC, Laurin C, Ring SM, Gaunt TR, McRae AF, Visscher PM, Montgomery GW, Martin NG, Hemani G, Suderman M, Relton CL, Smith GD, Evans DM (2018) Genome-wide survey of parent-of-origin effects on DNA methylation identifies candidate imprinted loci in humans. Hum Mol Genet 27(16):2927–2939. https://doi.org/10.1093/hmg/ddy206
Patowary A, Nesbitt R, Archer M, Bernier R, Brkanac Z (2017) Next generation sequencing mitochondrial DNA analysis in autism spectrum disorder. Autism Res 10(8):1338–1343. https://doi.org/10.1002/aur.1792
Pei L, Wallace DC (2018) Mitochondrial etiology of neuropsychiatric disorders. Biol Psychiatry 83(9):722–730. https://doi.org/10.1016/j.biopsych.2017.11.018. (Elsevier USA)
Penagarikano O, Mulle JG, Warren ST (2007) The pathophysiology of Fragile X syndrome. Annu Rev Genomics Hum Genet 8:109–129. https://doi.org/10.1146/annurev.genom.8.080706.092249
Pilvar D, Reiman M, Pilvar A, Laan M (2019) Parent-of-origin-specific allelic expression in the human placenta is limited to established imprinted loci and it is stably maintained across pregnancy. Clin Epigenetics 11(1):94. https://doi.org/10.1186/s13148-019-0692-3
Piryaei F, Houshmand M, Aryani O, Dadgar S, Soheili ZS (2012) Investigation of the mitochondrial ATPase 6/8 and tRNA Lys genes mutations in autism. Cell J 14(2):98–101. /pmc/articles/PMC3584428/?report=abstract
Pons R, Andreu AL, Checcarelli N, Vilà MR, Engelstad K, Sue CM, Shungu D, Haggerty R, De Vivo DC, DiMauro S (2004) Mitochondrial DNA abnormalities and autistic spectrum disorders. J Pediatr 144(1):81–85. https://doi.org/10.1016/j.jpeds.2003.10.023
Rangasamy S, D’Mello SR, Narayanan V (2013) Epigenetics, autism spectrum, and neurodevelopmental disorders. Neurotherapeutics 10(4):742–756. https://doi.org/10.1007/s13311-013-0227-0
Reed ML, Leff SE (1994) Maternal imprinting of human SNRPN, a gene deleted in prader-willi syndrome. Nat Genet 6(2):163–167. https://doi.org/10.1038/ng0294-163
Reik W, Walter J (2001) Genomic imprinting: parental influence on the genome : article : nature reviews genetics. Nat Rev Genet 2(1):21–32
Rossi M, El-Khechen D, Black MH, Farwell Hagman KD, Tang S, Powis Z (2017) Outcomes of diagnostic exome sequencing in patients with diagnosed or suspected autism spectrum disorders. Pediatr Neurol 70:34-43.e2. https://doi.org/10.1016/j.pediatrneurol.2017.01.033
Rossignol DA, Frye RE (2012) Mitochondrial dysfunction in autism spectrum disorders: a systematic review and meta-analysis. In Mol Psychiatry 17(3):290–314. https://doi.org/10.1038/mp.2010.136. (Nature Publishing Group)
Rylaarsdam L, Guemez-Gamboa A (2019) Genetic causes and modifiers of autism spectrum Disorder. Front Cell Neurosci 13:385. https://doi.org/10.3389/fncel.2019.00385
Sadakata T, Furuichi T (2009) Developmentally regulated Ca2+-dependent activator protein for secretion 2 (CAPS2) is involved in BDNF secretion and is associated with autism susceptibility. Cerebellum 8(3):312–322. https://doi.org/10.1007/s12311-009-0097-5
Sadakata T, Furuichi T (2010) Ca2+-dependent activator protein for secretion 2 and autistic-like phenotypes. Neurosci Res 67(3):197–202. https://doi.org/10.1016/j.neures.2010.03.006
Sadakata T, Shinoda Y, Oka M, Sekine Y, Sato Y, Saruta C, Miwa H, Tanaka M, Itohara S, Furuichi T (2012) Reduced axonal localization of a Caps2 splice variant impairsaxonal release of BDNF and causes autistic-like behavior in mice. Proceedings of the National Academy of Sciences 109:21104–21109. https://doi.org/10.1073/pnas.1210055109
Sadakata T, Washida M, Iwayama Y, Shoji S, Sato Y, Ohkura T, Katoh-Semba R, Nakajima M, Sekine Y, Tanaka M, Nakamura K, Iwata Y, Tsuchiya KJ, Mori N, Detera-Wadleigh SD, Ichikawa H, Itohara S, Yoshikawa T, Furuichi T (2007) Autistic-like phenotypes in Cadps2-knockout mice and aberrant CADPS2 splicing in autistic patients. J Clin Investig 117(4):931–943. https://doi.org/10.1172/JCI29031
Sampath S, Bhat S, Gupta S, O’Connor A, West AB, Arking DE, Chakravarti A (2013) Defining the contribution of CNTNAP2 to autism susceptibility. PLoS One 8(10):e77906. https://doi.org/10.1371/journal.pone.0077906
Sanders SJ, He X, Willsey AJ, Ercan-Sencicek AG, Samocha KE, Cicek AE, Murtha MT, Bal VH, Bishop SL, Dong S, Goldberg AP, **lu C, Keaney JF, Klei L, Mandell JD, Moreno-De-Luca D, Poultney CS, Robinson EB, Smith L, Solli-Nowlan T, Su MY, Teran NA, Walker MF, Werling DM, Beaudet AL, Cantor RM, Fombonne E, Geschwind DH, Grice DE, Lord C, Lowe JK, Mane SM, Martin DM, Morrow EM, Talkowski ME, Sutcliffe JS, Walsh CA, Yu TW, Ledbetter DH, Martin CL, Cook EH, Buxbaum JD, Daly MJ, Devlin B, Roeder K, State MW (2015) Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 87(6):1215–1233. https://doi.org/10.1016/j.neuron.2015.09.016
Schanen NC (2006) Epigenetics of autism spectrum disorders. Hum Mol Genet 15(SUPPL. 2):138–150. https://doi.org/10.1093/hmg/ddl213
Schneider E, El Hajj N, Richter S, Roche-Santiago J, Nanda I, Schempp W, Riederer P, Navarro B, Bontrop RE, Kondova I, Scholz CJ, Haaf T (2014) Widespread differences in cortex DNA methylation of the “language gene” CNTNAP2 between humans and chimpanzees. Epigenetics 9(4):533–545. https://doi.org/10.4161/epi.27689
Schneider E, Mayer S, El Hajj N, Jensen LR, Kuss AW, Zischler H, Kondova I, Bontrop RE, Navarro B, Fuchs E, Zechner U, Haaf T (2012) Methylation and expression analyses of the 7q autism susceptibility locus genes <b><i>MEST</i></b> <b><i>COPG2,</i></b> and <b><i>TSGA14</i></b> in Human and Anthropoid Primate Cortices. Cytogenet Genome Res 136(4):278–287. https://doi.org/10.1159/000337298
Schroer RJ, Phelan MC, Michaelis RC, Crawford EC, Skinner SA, Cuccaro M, Simensen RJ, Bishop J, Skinner C, Fender D, Stevenson RE (1998) Autism and maternally derived aberrations of chromosome 15q. Am J Med Genet 76(4):327–336. https://doi.org/10.1002/(SICI)1096-8628(19980401)76:4%3c327::AID-AJMG8%3e3.0.CO;2-M
Searles Quick VB, Wang B, State MW (2021) Leveraging large genomic datasets to illuminate the pathobiology of autism spectrum disorders. Neuropsychopharmacology 46(1):55–69. https://doi.org/10.1038/s41386-020-0768-y
Sherman S, Pletcher BA, Driscoll DA (2005) Fragile X syndrome: diagnostic and carrier testing. Genet Med 7(8):584–587. https://doi.org/10.1097/01.GIM.0000182468.22666.dd. (Various)
Skuse DH (2000) Imprinting, the X-chromosome, and the male brain: explaining sex differences in the liability to autism. Pediatr Res 47(1):9–16. https://doi.org/10.1203/00006450-200001000-00006. (Lippincott Williams and Wilkins)
Skuse DH, James RS, Bishop DVM, Coppin B, Dalton P, Aamodt-Leeper G, Bacarese-Hamilton M, Creswell C, McGurk R, Jacobs PA (1997) Evidence from Turner’s syndrome of an imprinted X-linked locus affecting cognitive function. Nature 387(6634):705–708. https://doi.org/10.1038/42706
Smith SEP, Zhou YD, Zhang G, ** Z, Stoppel DC, Anderson MP (2011) Increased gene dosage of Ube3a results in autism traits and decreased glutamate synaptic transmission in mice. Sci Transl Med 3(103):103ra97. https://doi.org/10.1126/scitranslmed.3002627
Soellner L, Begemann M, Mackay DJG, Grønskov K, Tümer Z, Maher ER, Temple IK, Monk D, Riccio A, Linglart A, Netchine I, Eggermann T (2017) Recent advances in imprinting disorders. Clin Genet 91(1):3–13. https://doi.org/10.1111/cge.12827
Sohal V, Rubenstein JLR (2019) Excitation-inhibition balance as a framework for investigating mechanisms in neuropsychiatric disorders. Mol Psychiatry 24(9):1248–1257. https://doi.org/10.1038/s41380-019-0426-0
Sztainberg Y, Zoghbi HY (2016) Lessons learned from studying syndromic autism spectrum disorders. Nat Neurosci 19(11):1408–1418. https://doi.org/10.1038/nn.4420
Talkowski ME, Rosenfeld JA, Blumenthal I, Pillalamarri V, Chiang C, Heilbut A, Ernst C, Hanscom C, Rossin E, Lindgren AM, Pereira S, Ruderfer D, Kirby A, Ripke S, Harris DJ, Lee JH, Ha K, Kim HG, Solomon BD, Gropman AL, Lucente D, Sims K, Ohsumi TK, Borowsky ML, Loranger S, Quade B, Lage K, Miles J, Wu BL, Shen Y, Neale B, Shaffer LG, Daly MJ, Morton CC, Gusella JF (2012) Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 149(3):525–537. https://doi.org/10.1016/j.cell.2012.03.028
Trillingsgaard A, Østergaard JR (2004) Autism in Angelman syndrome. Autism 8(2):163–174. https://doi.org/10.1177/1362361304042720
Úbeda F, Gardner A (2010) A model for genomic imprinting in the social brain: juveniles. Evolution 64(9):2587–2600. https://doi.org/10.1111/j.1558-5646.2010.01015.x
Úbeda F, Gardner A (2011) A model for genomic imprinting in the social brain: Adults. Evolution 65(2):462–475. https://doi.org/10.1111/j.1558-5646.2010.01115.x
Urraca N, Cleary J, Brewer V, Pivnick EK, Mcvicar K, Thibert RL, Schanen NC, Esmer C, Lamport D, Reiter LT (2013) The interstitial duplication 15q11.2–q13 syndrome includes autism, mild facial anomalies and a characteristic EEG signature. Autism Res 6(4):268–279. https://doi.org/10.1002/aur.1284
Veltman MWM, Craig EE, Bolton PF (2005) Autism spectrum disorders in Prader-Willi and Angelman syndromes: a systematic review. Psychiatr Genet 15(4):243–254. https://doi.org/10.1097/00041444-200512000-00006
Veltman MWM, Thompson RJ, Roberts SE, Thomas NS, Whittington J, Bolton PF (2004) Prader-Willi syndrome: a study comparing deletion and uniparental disomy cases with reference to autism spectrum disorders. Eur Child Adolesc Psychiatry 13(1):42–50. https://doi.org/10.1007/s00787-004-0354-6
Victor AK, Donaldson M, Johnson D, Miller W, Reiter LT (2021) Molecular changes in Prader-Willi syndrome neurons reveals clues about increased autism susceptibility. Front Mol Neurosci 14:259. https://doi.org/10.3389/fnmol.2021.747855
Voineagu I, Wang X, Johnston P, Lowe JK, Tian Y, Horvath S, Mill J, Cantor RM, Blencowe BJ, Geschwind DH (2011) Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474(7351):380–386. https://doi.org/10.1038/nature10110
Vos T, Abajobir AA, Abbafati C, Abbas KM, Abate KH, Abd-Allah F, Abdulle AM, Abebo TA, Abera SF, Aboyans V, Abu-Raddad LJ, Ackerman IN, Adamu AA, Adetokunboh O, Afarideh M, Afshin A, Agarwal SK, Aggarwal R, Agrawal A, Agrawal S, Ahmad Kiadaliri A, Ahmadieh H, Murray CJL (2017) Global, regional, and national incidence, prevalence, and years lived with disability for 328 diseases and injuries for 195 countries, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. The Lancet 390(10100):1211–1259. https://doi.org/10.1016/S0140-6736(17)32154-2
Wang Y, Picard M, Gu Z (2016) Genetic evidence for elevated pathogenicity of mitochondrial DNA heteroplasmy in autism spectrum disorder. PLoS Genet 12(10):e1006391. https://doi.org/10.1371/journal.pgen.1006391
Waterhouse L (2013) Rethinking autism. Elsevier Inc., In Rethinking Autism. https://doi.org/10.1016/C2011-0-05166-9
Whalley HC, O’Connell G, Sussmann JE, Peel A, Stanfield AC, Hayiou-Thomas ME, Johnstone EC, Lawrie SM, McIntosh AM, Hall J (2011) Genetic variation in CNTNAP2 alters brain function during linguistic processing in healthy individuals. Am J Med Genet B Neuropsychiatr Genet 156(8):941–948. https://doi.org/10.1002/ajmg.b.31241
Williams TA, Mars AE, Buyske SG, Stenroos ES, Wang R, Factura-Santiago MF, Lambert GH, Johnson WG (2007) Risk of autistic disorder in affected offspring of mothers with a glutathione S-transferase P1 haplotype. Arch Pediatr Adolesc Med 161(4):356–361. https://doi.org/10.1001/archpedi.161.4.356
Wolf JB, Wade MJ (2009) What are maternal effects (and what are they not)? Philos Trans R Soc B Biol Sci 364(1520):1107–1115. https://doi.org/10.1098/rstb.2008.0238
Wray NR, Gratten J (2018) Sizing up whole-genome sequencing studies of common diseases news-and-views. Nat Genet 50(5):635–637. https://doi.org/10.1038/s41588-018-0113-0. (Nature Publishing Group)
Yazdi PG, Su H, Ghimbovschi S, Fan W, Coskun PE, Nalbandian A, Knoblach S, Resnick JL, Hoffman E, Wallace DC, Kimonis VE (2013) Differential gene expression reveals mitochondrial dysfunction in an imprinting center deletion mouse model of Prader-Willi syndrome. Clin Transl Sci 6(5):347–355. https://doi.org/10.1111/cts.12083
Yi JJ, Berrios J, Newbern JM, Snider WD, Philpot BD, Hahn KM, Zylka MJ (2015) An autism-linked mutation disables phosphorylation control of UBE3A. Cell 162(4):795–807. https://doi.org/10.1016/j.cell.2015.06.045
Yoo HJ, Park M, Kim SA (2017) Difference in mitochondrial DNA copy number in peripheral blood cells between probands with autism spectrum disorders and their unaffected siblings. World J Biol Psychiatry 18(2):151–156. https://doi.org/10.1080/15622975.2016.1234069
Yoon SH, Choi J, Lee WJ, Do JT (2020) Genetic and epigenetic etiology underlying autism spectrum disorder. J Clin Med 9(4):966. https://doi.org/10.3390/jcm9040966
Žigman T, PetkovićRamadža D, Šimić G, Barić I (2021) Inborn errors of metabolism associated with autism spectrum disorders: approaches to intervention. Front Neurosci 15:624. https://doi.org/10.3389/fnins.2021.673600. (Frontiers Media S.A)
Acknowledgements
The authors would like to thank an anonymous reviewer for very helpful comments.
Funding
Open Access funding provided by the IReL Consortium This work was supported by the Brain & Behavior Research Foundation 2016 NARSAD Young Investigator Grant no. 24875.
Author information
Authors and Affiliations
Contributions
Both authors contributed to the conception, design and material preparation for this review article. The manuscript was written jointly by both authors and both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
As this is a review article, no ethics approval is required.
Conflict of interests
The authors declare no competing interests.
Additional information
Communicated by Michal Witt
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ryan, N.M., Heron, E.A. Evidence for parent-of-origin effects in autism spectrum disorder: a narrative review. J Appl Genetics 64, 303–317 (2023). https://doi.org/10.1007/s13353-022-00742-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-022-00742-8