Abstract
The global coronavirus disease 2019 (COVID-19) pandemic has had significant economic and social impacts on billions of people worldwide since severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was first reported in Wuhan, China, in November 2019. Although polymerase chain reaction (PCR)-based technology serves as a robust test to detect SARS-CoV-2 in patients with COVID-19, there is a high demand for cost-effective, rapid, comfortable, and accurate point-of-care diagnostic tests in medical facilities. This review introduces the SARS-CoV-2 viral structure and diagnostic biomarkers derived from viral components. A comprehensive introduction of a paper-based diagnostic platform, including detection mechanisms for various target biomarkers and a COVID-19 commercial kit is presented. Intrinsic limitations related to the poor performance of currently developed paper-based devices and unresolved issues are discussed. Furthermore, we provide insight into novel paper-based diagnostic platforms integrated with advanced technologies such as nanotechnology, aptamers, surface-enhanced Raman spectroscopy (SERS), and clustered regularly interspaced short palindromic repeats (CRISPR)-Cas. Finally, we discuss the prospects for the development of highly sensitive, accurate, cost-effective, and easy-to-use point-of-care COVID-19 diagnostic methods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was first reported in Wuhan, China, in November 2019, rapidly spreading worldwide and causing a global pandemic of coronavirus disease 2019 (COVID-19). This led to more than 500 million confirmed cases and more than 6.2 million deaths in more than 215 countries, as of April 28, 2022 [1]. The COVID-19 outbreak has had a significant economic and social impact on billions of people worldwide. Strong government sanctions, such as social distancing, quarantine, and lockdowns, have paralyzed businesses, severely weakening the global economy. This has caused many people to lose their jobs or has severely affected their household economy. Moreover, as the COVID-19 outbreak continues for an extended period, the medical community and staff, as well as the general public, are facing significant stressful situations (e.g., anxiety and depression) due to the intractable virus. Although effective vaccines and therapeutics for COVID-19 have been already developed and widely disseminated, transmission is still rapid and sustained owing to the emergence of the highly contagious variants of SARS-CoV-2 caused by mutations [2, 3].
Since the beginning of the COVID-19 pandemic, SARS-CoV-2 has accumulated mutations, and several types of SARS-CoV-2 variants identified by viral genome sequencing have been discovered in various global regions. Recent studies have shown that SARS-CoV-2 variants are more transmissible than the wild-type virus [4,5,6,7]. Additionally, the SARS-CoV-2 variants can bypass immune protection against exposure to the same virus conferred by previous vaccinations and infections [8, 9]. Fortunately, variant viruses do not continuously evolve to increase the risk of disease severity. The Omicron variant, which is currently dominant worldwide, is less severe than previous strains, especially compared to the Delta variant [10]. However, there is no guarantee that the subsequent dominant variants will not cause severe disease symptoms. In the future, devastating SARS-CoV-2 variants may emerge with higher viral transmissibility, disease severity, and vaccine-bypassing efficacy through the continued accumulation of mutations in the virus genome. To date, only a limited number of therapeutics, such as remdesivir and dexamethasone, have shown some effectiveness in reducing the mortality or severity of COVID-19. However, universally applicable therapeutics for asymptomatic and mild to moderate COVID-19 patients have not yet been developed. [11]. In the absence of universal antiviral therapeutics and reduced vaccine efficacy due to the immune escape of the variants, accurate, fast, and efficient diagnostic tools to identify patients with COVID-19 during the early stages of infection are vital for the control and further prevention of this disease.
Timely diagnosis, effective treatment, and future prevention are the most critical factors for successfully managing COVID-19 [12]. Among these, timely diagnosis plays an essential role in preventing and slowing the spread of the disease as the first line of defense. Early diagnosis of infected individuals enables immediate isolation of patients with highly contagious viruses (e.g., SARS-CoV-2), effectively controlling the spread of the disease. Moreover, early diagnosis allows physicians to provide immediate treatment, increasing the chances of cure and survival [13]. From the first outbreak of COVID-19 until now, the gold standard for confirming COVID-19 infection has been quantitative reverse transcription PCR (RT-qPCR) which amplifies small amounts of viral RNA in samples collected from an infected individual. However, this standard technique requires the analysis to be performed by trained experts and in a fully equipped laboratory, which inevitably increases the cost and time of testing. In a public health emergency where COVID-19 cases are increasing and COVID-19 testing capacity needs to be expanded, the introduction of more inexpensive, faster, easier-to-use diagnostic testing that analyzes samples at the point of use and screens for COVID-19 in a larger population is essential [14]. To compensate for the shortcomings of the current RT-qPCR assays, paper-based rapid diagnostic tests, especially lateral flow assays (LFA), have recently received widespread attention as an alternative to suppress the rapid spread and reinfection of COVID-19 [15]. Numerous LFA-based rapid diagnostic devices for diagnosing COVID-19 have been developed with great success, especially in resource-constrained environments and rapidly growing numbers of patients [16, 17]. So far, LFA-based rapid diagnostic technology has been used as an adjunct to RT-qPCR for COVID-19 confirmation due to its limited performance in terms of assay sensitivity and reproducibility [15]. However, extensive and intensive efforts are being made in academia and industry to improve the performance of LFA-based rapid inspection.
This review introduces the SARS-CoV-2 viral structure and diagnostic biomarkers derived from viral components. Then, a comprehensive introduction of paper-based diagnostic platforms, including device components, detection mechanisms for different target markers (e.g., nucleic acid, antigen, and antibody makers), and COVID-19 commercial kits, is presented. The intrinsic limitations related to the poor performance of the currently developed paper-based devices are also discussed. Furthermore, novel paper-based diagnostic platforms integrated with advanced technologies such as nanotechnology, aptamers, surface-enhanced Raman spectroscopy (SERS), and clustered regularly interspaced short palindromic repeats (CRISPR)-Cas are presented. These novel detection methods are promising for improving the diagnostic performance of previously reported paper-based tests. Finally, we discuss the prospects for the development of highly sensitive, accurate, cost-effective, and easy-to-use point-of-care COVID-19 diagnostic methods.
2 SARS-CoV-2 Viral Structure and Diagnostic Targets
The SARS-CoV-2 belongs to the genus β-coronavirus and comprises a single positive-strand RNA with a genome of ~ 30 kb. The SARS-CoV-2 genome codes ten genes that produce 26 proteins [18]. The genes are arranged in the sequence 5′ cap structure-replicase (open reading frame1/ab, ORF1/ab)-structural proteins with a [spike (S)-envelope (E)-membrane (M)-nucleocapsid (N)]-3′ poly (A) tail [19]. Among these genes, unique and specific nucleotide sequences representing SARS-CoV-2, such as genomic fingerprints, are targets for COVID-19 diagnosis. The genome sequence of SARS-CoV-2 was shared through the Global Initiative on Sharing All Influenza Data (GISAID) platform, on January 12, 2020 [20]. Subsequently, various primer and probe sets have been developed to amplify specific viral RNA sequences, and the WHO has posted these primer–probe sets [20,21,22,23], enabling the rapid development of COVID-19 nucleic acid amplification tests (NAATs). More than 100 RT-qPCR kits have been designed and prototyped and are the United States food and drug administration (FDA) emergency use authorization (EUA)-approved for COVID-19 diagnosis. These kits aim to amplify specific regions of viral genes, such as structural protein genes (N, E, S, and M) and confirmation genes (ORF1ab and RNA-dependent RNA polymerase (RdRp)) [20, 23, 24] (Fig. 1a). RT-qPCR offers high accuracy and very low analytical sensitivity; thus, it has been used as the gold standard for confirming COVID-19 infection.
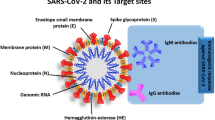
SARS-CoV-2 genome, structure comprising major proteins, and variation levels of biomarkers across the duration of the infection. a The SARS-CoV-2 genome codes ten genes, and the genes are arranged in the sequence 5′ cap structure-ORF1/ab-spike (S)-envelope (E)-membrane (M)-nucleocapsid (N)-3′ poly (A) tail. Primer–probe sets for SARS-CoV-2 RNA amplification developed by research groups around the world [US Centers for Disease Control and Prevention (US-CDC) – target sequences: 28,287 ~ 28,358, 28,681 ~ 28,752, 29,164 ~ 29,230; China CDC – target sequences: 13,342 ~ 13,460, 28,881 ~ 28,979; Charité–Universitätsmedizin Berlin in Germany (Charité) – target sequences: 15,431 ~ 15,530, 26,269 ~ 26,381; National Institute of Infectious Disease in Japan (Japan NIID) – target sequence: 29,125 ~ 29,282; and University of Hong Kong (HKU) – target sequences: 18,778 ~ 18,909, 29,145 ~ 29,254]. b SARS-CoV-2 is mainly composed of four major proteins: spike (S) (red), membrane (M) (orange), envelope (E) (green), and nucleocapsid (N) (purple) proteins. c Temporal dynamics of the viral load and antigen and antibody levels. Since the types and amounts of biomarkers present in a patient's body fluid differ depending on the stage of infection, it is critical to select an appropriate biomarker and a method that can effectively detect it for an accurate diagnosis of COVID-19
The SARS-CoV-2 genome encodes four major structural and functional proteins: the spike (S), membrane (M), envelope (E), and nucleocapsid (N) proteins (Fig. 1b) [11, 25, 26]. The S protein comprises two functional subunits, S1 and S2; S1 acts as a key protein for selective binding with the host cell receptor, angiotensin-converting enzyme 2 (ACE2), while S2 mediates membrane fusion [11, 27, 28]. The N protein is the most abundant protein in the virus and plays a crucial role in packaging and protecting viral genomic RNA. The N protein is highly immunogenic and rarely changes with disease progression [29,30,31,32]. The M protein is the most abundant structural protein that defines the shape of a virus, while the E protein is the smallest major structural protein involved in virus assembly and pathogenesis [28]. The S and N proteins are considered the most valuable antigenic biomarkers for diagnosing COVID-19 in the various detection methods for SARS-CoV-2 [33, 34].
In contrast, specific antibodies produced by the immune response to protect the body from SARS-CoV-2 infection could be another option for diagnosing COVID-19. Immunoglobulin M (IgM) is the first line of defense during viral infections appearing in the blood after SARS-CoV-2 infection, and increases rapidly, which is an indicator of early stage infection. IgG antibodies subsequently produced by the IgM antibodies are responsible for immunological memory and long-term immunity, which serves as the body's immune defense system to avoid reinfection with the same pathogen [35, 36]. IgA antibodies limit the entry of microorganisms and antigens into the susceptible mucosal barrier through respiratory mucosal epithelial protection and homeostasis regulation [37,38,39]. IgA antibody responses appear early and are characterized by intense and sustained maintenance [40]. Therefore, the detection of IgA antibodies is advantageous in the early infection stages.
Assay sensitivity is affected by the temporal profile of the viral load (or concentration of the biomarkers) across the duration of the infection. This is a critical factor in determining the diagnostic accuracy. Several studies have suggested the temporal dynamics of the viral load, antigen and antibody levels after SARS-CoV-2 infection [41,42,43,44]. In general, the viral load increased rapidly from the time of infection, peaked, and then decreased rapidly (within a few days) (Fig. 1c). After 10 days of infection, the viral load is reduced by a factor of 100 or more [41, 45]. Considering the virus structure, the quantities of viral RNA and antigens were consistent with the trend of the viral load. Therefore, the optimal time for detecting viral RNA and antigens is approximately 7 days, immediately after symptom onset. Conversely, viral RNA cannot be detected in the early or late phase of infection. Another study analyzed the serological response of COVID-19 patients to viral infections [42]. Approximately 10 days after symptom onset, the IgM response to SARS-CoV-2 is predominant over other immunoglobulins. After approximately 15 days, the IgM response decreased and disappeared. IgG antibodies are produced later than IgM production, 10–14 days after symptom onset; however, IgG antibodies persist and are detectable for a long time [43, 44]. IgM antibodies can mainly be used for the early detection of SARS-CoV-2 infection, whereas IgG antibodies may be more appropriately used to identify past infections.
The type and amount of biomarkers present in the patient's body fluid differed depending on the infection stage of the COVID-19 patient (Fig. 1c). To analyze these biomarkers, an appropriate sample preparation process according to the specimen and analysis method is also essential. In general, the upper respiratory tract sample is used in the case of RT-qPCR and antigen tests, and if possible, the lower respiratory tract sample is inspected simultaneously. Generally, the nasopharyngeal swab method is more sensitive than the nasal swab method for specimen collection in COVID-19 [46]. However, the nasopharyngeal swab method should be performed by a trained healthcare provider. The Centers for Disease Control and Prevention (CDC) noted that if both nasopharyngeal and oropharyngeal specimens can be collected, the sensitivity of the test can be maximized by combining the two specimens in one tube. On the other hand, the nasal swab method is more comfortable than the nasopharyngeal swab method and can be used easily by the general public; therefore, many diagnostic kits have been developed for nasal swab samples. Before the RT-qPCR test, RNA extraction from the sample is a crucial step to getting accurate amplification results [47, 48]. It should be carried out using proven equipment and reagents, and contamination should be prevented throughout the sampling and analysis process. In the case of the antibody tests, a small amount of blood should be collected through venipuncture or the fingerstick method. To reduce interference or increase sensitivity, any clots or erythrocyte sediments in the samples may be removed after the appropriate clotting or centrifuge method. It is essential for serum or plasma analysis kits, but these processes may be omitted for several whole blood or fingerstick kits [49, 50]. Therefore, for accurate diagnosis of COVID-19, it is crucial to select an appropriate biomarker and a method that can effectively detect these biomarkers according to the stage of infection.
3 Paper-Based Diagnostic Platform for COVID-19 Diagnosis
While the economically developed world has access to many advanced medical tools, trained personnel, and resources to perform diagnostic tests for maintaining health, these similar resources are unfortunately not as accessible in develo** countries. Rapid and accurate diagnosis is the first step toward improving healthcare conditions in develo** countries. In dire situations, such as the COVID-19 pandemic, the damage is inevitably more severe for countries marginalized from good medical services. In these countries, the supply of the COVID-19 vaccine is delayed and accurate diagnostic techniques for early diagnosis are extremely limited. Many disease-related deaths, including COVID-19, would have been preventable if the disease had been diagnosed earlier and followed by appropriate treatment. Moreover, large-scale diagnostic testing is critical to contain the COVID-19 epidemic, even in developed countries with well-established healthcare services [51]. Standard diagnostic techniques (i.e., RT-qPCR) are limited in the rapid screening of confirmed cases from a rapidly elevated number of suspected cases. For these outbreaks, it is crucial to have medical diagnostic platforms that can analyze samples on-site and provide immediate results [13].
Paper-based analytical devices show great potential in delivering POC diagnostic systems to the develo** world because of their remarkable properties such as biocompatibility, porosity, ease of modification, flexibility, chemical inertness, eco-friendliness, and ease of storage and transportation [52, 53]. Moreover, various sample types can be applied to paper-based analysis equipment, and sample transfer is possible without requiring additional power owing to capillary force. Over the last three decades, paper-based POC tests have been developed for various biomedical applications and launched as both ‘over-the-counter’ products, such as glucose monitoring and pregnancy testing, and ‘professional market’ products that can diagnose infectious disease, cardiac markers, diabetes, lipidoses, hemopathies, and several cancers.
Paper-based POC tests have also played a crucial role in the current COVID-19 pandemic. Many researchers in academia and industry have made intensive efforts to develop a simple, convenient, fast, sensitive, and accurate technology that can detect SARS-CoV-2. The technology that can best meet the strong demand for practical POC diagnostic tests in COVID-19 control is paper-based POC testing. Paper-based POC diagnostic platforms range from simple one-dimensional platforms, such as dipstick and LFA, to complex three-dimensional platforms, such as microfluidic paper-based assay devices (μPAD) and electrochemical paper-based assay devices (ePAD). Among these, LFA is a highly mature paper-based diagnostic technology that researchers and manufacturers have invested the most effort and cost in develo** COVID-19 diagnostic kits. A typical LFA comprises a sample pad, a conjugate pad, a nitrocellulose membrane, and an absorbent pad. The sample flow begins on the sample pad and meets the signal molecules that have dried on the conjugate pad. All biomarkers, antigens, antibodies, and SARS-CoV-2 RNA used to detect SARS-CoV-2 can be applied to LFA.
3.1 LFA-Based Diagnostic Platform for Detecting Viral Antigens
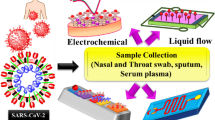
Unlike RT-qPCR, antigen-based diagnostics directly detect the presence of SARS-CoV-2 and related proteins in a sample taken from a nasopharyngeal swab or nasal passage without sample pretreatment and amplification. Therefore, it can provide a diagnosis of COVID-19 with faster and easier results at lower cost than RT-qPCR. Antigen-based diagnosis is based on immunoassay reactions that involve antigens and antibodies. The configuration of the general LFA diagnostic platforms for detecting SARS-CoV-2-specific antigens is shown in Fig. 2a. All proteins constituting SARS-CoV-2 can be targeted to diagnose COVID-19, but antigen tests for COVID-19 have been developed mainly targeting the S and N proteins [54,55,56]. To detect the SARS-CoV-2 antigens, a specific antibody pair that recognizes different regions of the target antigen is required. A capture antibody is immobilized on a nitrocellulose membrane to form a test line (first line), and another antibody is labeled with a signal molecule, mainly gold nanoparticles, and serves as the detection antibody. Additionally, the control line (second line) also serves to check whether the sample flowed through the nitrocellulose membrane, and additional Ig-types of antibodies that could not affect the test are used. When a nasopharyngeal swab sample of a patient with COVID-19 is loaded into the LFA device, the sample containing the target antigens flows along with the LFA strip by capillary force and first encounters the detection antibody. Target antigens are captured by both detection and capture antibodies to form a sandwich complex. After 15–20 min of sample loading, the appearance of color in the test and control lines is confirmed visually or by a portable analyzer.
Paper-based diagnostic platforms including device components, detection mechanisms for different target markers, a antigens, b antibodies, and c RNA. a To detect the SARS-CoV-2 antigens, a specific antibody pair is required. These capture and detection antibodies detect SARS-CoV-2-specific antigens (S and N proteins) while forming a sandwich complex. After 15–20 min of sample loading, the appearance of color in the test and control lines is confirmed visually or by a portable analyzer. b In serological tests (detecting IgM and IgG antibodies), the N (or S) proteins of SARS-CoV-2 are conjugated with gold nanoparticles and used as signal molecules to detect IgM and IgG antibodies. Anti-human IgM (or IgG) antibodies are immobilized on a nitrocellulose membrane to form test lines. When the sample contains the SARS-CoV-2-specific IgM or IgG antibodies, the antibodies are bound to the N (or S) protein-conjugated gold nanoparticles and finally bound to the test line, resulting in vivid color. c Isothermal amplification techniques combined with an LFA contribute to achieving POC tests for SARS-CoV-2 RNA detection. First, an isothermal amplification process is performed for target RNA amplification, and then an LFA reaction is performed so that the results can be easily checked
Although various paper-based antigen diagnostic tests have been developed, the sensitivity of the rapid antigen test is unclear and is lower than that of RT-qPCR. The limit of detection (LOD) of antigen tests is approximately 105 copies/mL, while that of RT-qPCR is as low as 102 copies/mL [57, 58]. False-negative results may occur when the concentration of the target antigen in the clinical specimen is below the analytical sensitivity of antigen tests. Several studies have been conducted to overcome these limitations. Liu et al. presented a novel nanozyme-based chemiluminescence paper assay for detecting SARS-CoV-2 S antigen. In this case, nanozyme (Co-Fe@hemin-peroxidase) and chemiluminescent immunoassays were integrated with LFA to achieve sensitivity (360 TCID50/mL) comparable to that of an ELISA method [59]. To improve the performance of the previous signal molecule, gold nanoparticles [60], latex beads [61], cellulose nanobeads [62], and fluorescent microparticles [30] have been introduced into LFA. These signaling molecules have higher signal intensities, resulting in an approximately tenfold improvement in sensitivity compared with previous gold nanoparticle-based LFAs.
Our group also proposed a novel rapid detection method for the SARS-CoV-2 S antigen. Using the cellular receptor for SARS-CoV-2, angiotensin-converting enzyme 2 (ACE2), the SARS-CoV-2 S1 antigen was successfully detected in clinical specimens of COVID-19 patients (Fig. 3a) [63, 64]. Furthermore, we developed SARS-CoV-2 N antigen-specific single-chain variable fragment crystallizable fragment (scFv-Fc) fusion antibodies using phage display technology and applied them to the LFA platform (Fig. 3b) [62]. This scFv-Fc-based rapid diagnostic test showed high specificity that could distinguish even the N protein of SARS-CoV. Baker et al. developed a glyconanoparticle consisting of multivalent gold nanoparticles bearing sialic acid derivatives [65]. They discovered that the N-acetyl neuraminic acid is bound to the S protein and developed LFAs that exploit this interaction as a detection mechanism. This glycoprotein-based LFA showed high selectivity for the SARS-CoV S protein.
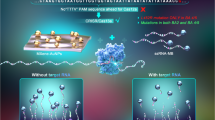
a Cellular receptor (ACE2)-based LFA for detecting SARS-CoV-2 S1 antigen, reproduced with permission from [63], copyright 2021 Elsevier. b Development of scFv-Fc-based LFA for detection of the SARS-CoV-2 N protein. Highly sensitive and specific scFv-Fc fusion proteins are rapidly screened by phage display technology, reproduced with permission from [62], copyright 2021 Elsevier. c Configuration of detecting system to quantify LFA results with the photon-counting approach and representative results for IgG antibody detection (concentrations range: from 1000 to 0.1 ng/mL), reproduced with permission from [74], copyright 2020 AIP. d LFA strip to detect anti-SARS-CoV-2 IgA antibody and the simple and universal smartphone reader to detect the optical signal from LFA, reproduced with permission from [75], copyright 2021 Elsevier. e Lateral Flow Strip Membranes (LFSM)-based on highly specific and sensitive detection of SARS-CoV-2. The LFSM assay allows simultaneous detection of the multiple regions of SARS-CoV-2 RNA in a sing test, reproduced with permission from [77, 76], copyright 2020 ACS. f Principle of reverse transcription-enzymatic recombinase amplification (RT-ERA). The RT-ERA has the capability of ultrasensitive, field-deployable, and simultaneous dual-gene detection of SARS-CoV-2 RNA, reproduced with permission from [92]. Typical LFA tests use gold nanoparticles as signal reporters; however, their sensitivities are not very high. Extensive efforts have been made to increase diagnostic sensitivity by improving the performance of signaling molecules in various studies. Liu et al. reported an advanced LFA based on gold nanoparticles for enhanced specific binding and thermal contrast amplification (TCA) for signal amplification. With TCA, the gold nanoparticles captured in the test line were excited by laser irradiation and exhibited a substantial photothermal effect, enabling the detection of subvisual positives. They successfully detected SARS-CoV-2 receptor-binding domain (RBD) antigen as low as 28.6 aM in a human nasopharyngeal swab [93]. Selenium nanoparticles exhibit favorable biocompatibility and are readily conjugated with biological molecules without losing their activity [94]. A POC selenium nanoparticle-based LFA was developed to detect SARS-CoV-2 IgM and IgG [72]. In this study, the authors made a new selenium nanoparticle-based LFA kit and visually detected anti-SARS-CoV-2 IgG and IgM antibodies in human serum within 10 min. Furthermore, the authors performed a clinical evaluation using a sample of 90 patients with COVID-19 and 263 uninfected negative controls, demonstrating a sensitivity and specificity of 93.33% and 97.34%, respectively. This selenium nanoparticle-based LFA showed superior detection limits compared to the gold nanoparticle-based LFA in IgM antibody detection and did not show cross-reactivity with influenza A, influenza B, anti-nuclear antibodies, and rheumatoid factor. On the other hand, in recent years, quantum dots (QDs) have been widely used as fluorescent signal reporters in LFA because of their excellent optical properties, such as quantifiable fluorescence intensity, broad excitation, and high light stability [95,96,13]. However, there is still an aperture that needs to be addressed to develop an ideal and universal POC diagnostic platform. Smartphones, which offer imaging, filtering, and image/data processing, can be powerful tools to compensate for current POC diagnostic platforms. Because many people always carry their smartphones, they can effectively serve as handheld readers that rapidly and accurately check for infections and contribute significantly to disease control and surveillance. In addition, it is expected that diagnosis performance will be improved through the introduction of artificial intelligence (AI). Recently, the potential application of AI-based COVID-19 diagnosis has been extensively explored in the field of lung detection imaging, such as computed tomography (CT) imaging [132, 133], chest radiographs (X-ray) imaging [134, 135]. By learning from tremendous amounts of diagnostic results through deep learning techniques and using them to present accurate current or future results, more effective infectious disease management is possible.
References
Johns Hopkins University (JHU) and Center for Systems Science and Engineering (CSSE), Coronavirus Resource Center., cited April 28, 2022. https://coronavirus.jhu.edu/map.html
Forni, G., Mantovani, A., Forni, G., Mantovani, A., Moretta, L., Rappuoli, R., et al.: COVID-19 vaccines: where we stand and challenges ahead. Cell Death Differ. 28, 626–639 (2021)
Kyriakidis, N.C., López-Cortés, A., González, E.V., Grimaldos, A.B., Prado, E.O.: SARS-CoV-2 vaccines strategies: a comprehensive review of phase 3 candidates. npj Vaccines 6, 28 (2021)
Li, L., Liu, Y., Tang, X., He, D.: The disease severity and clinical outcomes of the SARS-CoV-2 variants of concern. Front. Public Health 9, 775224 (2021)
Tegally, H., Wilkinson, E., Giovanetti, M., Iranzadeh, A., Fonseca, V., Giandhari, J., et al.: Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 592, 438–443 (2021)
Campbell, F., Archer, B., Laurenson-Schafer, H., **nai, Y., Konings, F., Batra, N., et al.: Increased transmissibility and global spread of SARS-CoV-2 variants of concern as at June 2021. Eurosurveillance 26, 2100509 (2021)
Faria, N.R., Mellan, T.A., Whittaker, C., Claro, I.M., Candido, Dd.S., Mishra, S., et al.: Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 372, 815–821 (2021)
Corey, L., Beyrer, C., Cohen, M.S., Michael, N.L., Bedford, T., Rolland, M.: SARS-CoV-2 variants in patients with immunosuppression. N. Engl. J. Med. 385, 562–566 (2021)
Thomas, E., Delabat, S., Carattini, Y.L., Andrews, D.M.: SARS-CoV-2 and variant diagnostic testing approaches in the United States. Viruses 13, 2492 (2021)
Nyberg, T., Ferguson, N.M., Nash, S.G., Webster, H.H., Flaxman, S., Andrews, N., et al.: Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: a cohort study. The Lancet 399, 1303–1312 (2022)
Habli, Z., Saleh, S., Zaraket, H., Khraiche, M.L.: COVID-19 in-vitro diagnostics: State-of-the-art and challenges for rapid, scalable, and high-accuracy screening. Front. Bioeng. Biotechnol. 8, 605702 (2021)
Carter, L.J., Garner, L.V., Smoot, J.W., Li, Y., Zhou, Q., Saveson, C.J., et al.: Assay techniques and test development for COVID-19 diagnosis. ACS Cent. Sci. 6, 591–605 (2020)
Pérez-López, B., Mir, M.: Commercialized diagnostic technologies to combat SARS-CoV2: advantages and disadvantages. Talanta 225, 121898 (2021)
Somborac Bačura, A., Dorotić, M., Grošić, L., Džimbeg, M., Dodig, S.: Current status of the lateral flow immunoassay for the detection of SARS-CoV-2 in nasopharyngeal swabs. Biochem. Med. (Zagreb) 31, 020601 (2021)
Kevadiya, B.D., Machhi, J., Herskovitz, J., Oleynikov, M.D., Blomberg, W.R., Bajwa, N., et al.: Diagnostics for SARS-CoV-2 infections. Nat. Mater. 20, 593–605 (2021)
Montesinos, I., Gruson, D., Kabamba, B., Dahma, H., Van den Wijngaert, S., Reza, S., et al.: Evaluation of two automated and three rapid lateral flow immunoassays for the detection of anti-SARS-CoV-2 antibodies. J. Clin. Virol. 128, 104413 (2020)
Silveira, M.F., Barros, A.J.D., Horta, B.L., Pellanda, L.C., Victora, G.D., Dellagostin, O.A., et al.: Population-based surveys of antibodies against SARS-CoV-2 in Southern Brazil. Nat. Med. 26, 1196–1199 (2020)
Chen, L., Liu, W., Zhang, Q., Xu, K., Ye, G., Wu, W., et al.: RNA based mNGS approach identifies a novel human coronavirus from two individual pneumonia cases in 2019 Wuhan outbreak. Emerg. Microb. Infect. 9, 313–319 (2020)
Kilic, T., Weissleder, R., Lee, H.: Molecular and immunological diagnostic tests of COVID-19: current status and challenges. iScience 23, 101406 (2020)
Jung, Y., Park, G.-S., Moon, J.H., Ku, K., Beak, S.-H., Lee, C.-S., et al.: Comparative analysis of primer–probe sets for RT-qPCR of COVID-19 causative virus (SARS-CoV-2). ACS Infect. Dis. 6, 2513–2523 (2020)
WHO. Coronavirus disease (COVID-19) technical guidance: Laboratory testing for 2019-nCoV in humans cited April 8 2020. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/laboratory-guidance/>
Chu, D.K.W., Pan, Y., Cheng, S.M.S., Hui, K.P.Y., Krishnan, P., Liu, Y., et al.: Molecular diagnosis of a novel coronavirus (2019-nCoV) causing an outbreak of pneumonia. Clin. Chem. 66, 549–555 (2020)
Corman, V.M., Landt, O., Kaiser, M., Molenkamp, R., Meijer, A., Chu, D.K., et al.: Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 25, 2000045 (2020)
Chan, J.F.-W., Yip, C.C.-Y., To, K.K.-W., Tang, T.H.-C., Wong, S.C.-Y., Leung, K.-H., et al.: Improved molecular diagnosis of COVID-19 by the novel, highly sensitive and specific COVID-19-RdRp/Hel real-time reverse transcription-PCR assay validated In Vitro and with clinical specimens. J. Clin. Microbiol. 58, e00310-00320 (2020)
Bong, J.-H., Kim, T.-H., Jung, J., Lee, S.J., Sung, J.S., Lee, C.K., et al.: Pig sera-derived anti-SARS-CoV-2 antibodies in surface plasmon resonance biosensors. BioChip J. 14, 358–368 (2020)
Zou, L., Ruan, F., Huang, M., Liang, L., Huang, H., Hong, Z., et al.: SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N. Engl. J. Med. 382, 1177–1179 (2020)
Jung, J., Bong, J.-H., Kim, T.-H., Sung, J.S., Lee, C., Kang, M.-J., et al.: Isolation of antibodies against the spike protein of SARS-CoV from pig serum for competitive immunoassay. BioChip J. 15, 396–405 (2021)
Wang, Q., Zhang, Y., Wu, L., Niu, S., Song, C., Zhang, Z., et al.: Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 181, 894-904.e899 (2020)
Yamaoka, Y., Miyakawa, K., Jeremiah, S.S., Funabashi, R., Okudela, K., Kikuchi, S., et al.: Highly specific monoclonal antibodies and epitope identification against SARS-CoV-2 nucleocapsid protein for antigen detection tests. Cell Reports Medicine 2, 100311 (2021)
Diao, B., Wen, K., Zhang, J., Chen, J., Han, C., Chen, Y., et al.: Accuracy of a nucleocapsid protein antigen rapid test in the diagnosis of SARS-CoV-2 infection. Clin. Microbiol. Infect. 27(289), e281-289.e284 (2021)
Dutta, N.K., Mazumdar, K., Gordy, J.T., Dutch, R.E.: The nucleocapsid protein of SARS-CoV-2: a target for vaccine development. J. Virol. 94, e00647-e1620 (2020)
Bai, Z., Cao, Y., Liu, W., Li, J.: The SARS-CoV-2 nucleocapsid protein and its role in viral structure, biological functions, and a potential target for drug or vaccine mitigation. Viruses 13, 1115 (2021)
Che, X.-y, Qiu, L.-w, Pan, Y.-x, Wen, K., Hao, W., Zhang, L.-y, et al.: Sensitive and specific monoclonal antibody-based capture enzyme immunoassay for detection of nucleocapsid antigen in sera from patients with severe acute respiratory syndrome. J. Clin. Microbiol. 42, 2629–2635 (2004)
Woo, P.C.Y., Lau, S.K.P., Wong, B.H.L., Tsoi, H.-w, Fung, A.M.Y., Kao, R.Y.T., et al.: Differential sensitivities of severe acute respiratory syndrome (SARS) coronavirus spike polypeptide enzyme-linked immunosorbent assay (ELISA) and SARS coronavirus nucleocapsid protein ELISA for serodiagnosis of SARS coronavirus pneumonia. J. Clin. Microbiol. 43, 3054–3058 (2005)
To, K.K.-W., Tsang, O.T.-Y., Yip, C.C.-Y., Chan, K.-H., Wu, T.-C., Chan, J.M.-C., et al.: Consistent detection of 2019 novel coronavirus in saliva. Clin. Infect. Dis. 71, 841–843 (2020)
Kohmer, N., Westhaus, S., Rühl, C., Ciesek, S., Rabenau, H.F.: Clinical performance of different SARS-CoV-2 IgG antibody tests. J. Med. Virol. 92, 2243–2247 (2020)
Fagarasan, S., Honjo, T.: Intestinal IgA synthesis: regulation of front-line body defences. Nat. Rev. Immunol. 3, 63–72 (2003)
Bidgood, S.R., Tam, J.C.H., McEwan, W.A., Mallery, D.L., James, L.C.: Translocalized IgA mediates neutralization and stimulates innate immunity inside infected cells. Proc. Natl. Acad. Sci. 111, 13463–13468 (2014)
Moor, K., Diard, M., Sellin, M.E., Felmy, B., Wotzka, S.Y., Toska, A., et al.: High-avidity IgA protects the intestine by enchaining growing bacteria. Nature 544, 498–502 (2017)
Infantino, M., Manfredi, M., Grossi, V., Lari, B., Fabbri, S., Benucci, M., et al.: Closing the serological gap in the diagnostic testing for COVID-19: The value of anti-SARS-CoV-2 IgA antibodies. J. Med. Virol. 93, 1436–1442 (2021)
Zhou, P., Yang, X.-L., Wang, X.-G., Hu, B., Zhang, L., Zhang, W., et al.: A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020)
Tan W, Lu Y, Zhang J, Wang J, Dan Y, Tan Z, et al. Viral kinetics and antibody responses in patients with COVID-19. medRxiv 2020.2003.2024.20042382 (2020)
an observational cohort study: To KK-W, Tsang OT-Y, Leung W-S, Tam AR, Wu T-C, Lung DC, et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2. Lancet. Infect. Dis 20, 565–574 (2020)
Thevarajan, I., Nguyen, T.H.O., Koutsakos, M., Druce, J., Caly, L., van de Sandt, C.E., et al.: Breadth of concomitant immune responses prior to patient recovery: a case report of non-severe COVID-19. Nat. Med. 26, 453–455 (2020)
Pan, Y., Zhang, D., Yang, P., Poon, L.L.M., Wang, Q.: Viral load of SARS-CoV-2 in clinical samples. Lancet. Infect. Dis 20, 411–412 (2020)
Patriquin, G., LeBlanc, J.J., Williams, C., Hatchette, T.F., Ross, J., Barrett, L., et al.: Comparison between nasal and nasopharyngeal swabs for SARS-CoV-2 rapid antigen detection in an asymptomatic population, and direct confirmation by RT-PCR from the residual buffer. Microbiology Spectrum 10, e02455-e12421 (2022)
Ambrosi, C., Prezioso, C., Checconi, P., Scribano, D., Sarshar, M., Capannari, M., et al.: SARS-CoV-2: comparative analysis of different RNA extraction methods. J. Virol. Methods 287, 114008 (2021)
Wozniak, A., Cerda, A., Ibarra-Henríquez, C., Sebastian, V., Armijo, G., Lamig, L., et al.: A simple RNA preparation method for SARS-CoV-2 detection by RT-qPCR. Sci. Rep. 10, 16608 (2020)
Kovac, M., Risch, L., Thiel, S., Weber, M., Grossmann, K., Wohlwend, N., et al.: EDTA-anticoagulated whole blood for SARS-CoV-2 antibody testing by electrochemiluminescence immunoassay (ECLIA) and enzyme-linked immunosorbent assay (ELISA). Diagnostics 10, 593 (2020)
Chen, C., Hu, H., Li, X., Zheng, Z., Wang, Z., Wang, X., et al.: Rapid detection of anti-SARS-CoV-2 antibody using a selenium nanoparticle-based lateral flow immunoassay. IEEE Trans Nanobiosci. 21, 37–43 (2022)
Tromberg, B.J., Schwetz, T.A., Pérez-Stable, E.J., Hodes, R.J., Woychik, R.P., Bright, R.A., et al.: Rapid scaling up of Covid-19 diagnostic testing in the United State—the NIH RADx initiative. N. Engl. J. Med. 383, 1071–1077 (2020)
Bastian, L.A., Nanda, K., Hasselblad, V., Simel, D.L.: Diagnostic efficiency of home pregnancy test kits. A meta-analysis. Arch. Family Med. 7, 465–469 (1998)
Ozer, T., McMahon, C., Henry, C.S.: Advances in paper-based analytical devices. Annu. Rev. Anal. Chem. 13, 85–109 (2020)
Bong, J.-H., Kim, T.-H., Jung, J., Lee, S.J., Sung, J.S., Lee, C.K., et al.: Competitive immunoassay of SARS-CoV-2 using pig sera-derived anti-SARS-CoV-2 antibodies. BioChip J. 15, 100–108 (2021)
Liu, W., Kou, G., Dong, Y., Zheng, Y., Ding, Y., Ni, W., et al.: Clinical application of chemiluminescence microparticle immunoassay for SARS-CoV-2 infection diagnosis. J. Clin. Virol. 130, 104576 (2020)
Yakoh, A., Pimpitak, U., Rengpipat, S., Hirankarn, N., Chailapakul, O., Chaiyo, S.: Paper-based electrochemical biosensor for diagnosing COVID-19: detection of SARS-CoV-2 antibodies and antigen. Biosens. Bioelectron. 176, 112912 (2021)
Nash B, Badea A, Reddy A, Bosch M, Salcedo N, Gomez AR, et al. The impact of high frequency rapid viral antigen screening on COVID-19 spread and outcomes: a validation and modeling study. medRxiv 2020.2009.2001.20184713 (2020)
Vogels, C.B.F., Brito, A.F., Wyllie, A.L., Fauver, J.R., Ott, I.M., Kalinich, C.C., et al.: Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT–qPCR primer–probe sets. Nat. Microbiol. 5, 1299–1305 (2020)
Liu, D., Ju, C., Han, C., Shi, R., Chen, X., Duan, D., et al.: Nanozyme chemiluminescence paper test for rapid and sensitive detection of SARS-CoV-2 antigen. Biosens. Bioelectron. 173, 112817 (2021)
Mertens, P., De Vos, N., Martiny, D., Jassoy, C., Mirazimi, A., Cuypers, L., et al.: Development and potential usefulness of the COVID-19 Ag Respi-Strip diagnostic assay in a pandemic context. Front. Med. 7, 225 (2020)
Grant, B.D., Anderson, C.E., Williford, J.R., Alonzo, L.F., Glukhova, V.A., Boyle, D.S., et al.: SARS-CoV-2 coronavirus nucleocapsid antigen-detecting half-strip lateral flow assay toward the development of point of care tests using commercially available reagents. Anal. Chem. 92, 11305–11309 (2020)
Kim, H.-Y., Lee, J.-H., Kim, M.J., Park, S.C., Choi, M., Lee, W., et al.: Development of a SARS-CoV-2-specific biosensor for antigen detection using scFv-Fc fusion proteins. Biosens. Bioelectron. 175, 112868 (2021)
Lee, J.-H., Choi, M., Jung, Y., Lee, S.K., Lee, C.-S., Kim, J., et al.: A novel rapid detection for SARS-CoV-2 spike 1 antigens using human angiotensin converting enzyme 2 (ACE2). Biosens. Bioelectron. 171, 112715 (2021)
Lee, J.-H., Lee, Y., Lee, S.K., Kim, J., Lee, C.-S., Kim, N.H., et al.: Versatile role of ACE2-based biosensors for detection of SARS-CoV-2 variants and neutralizing antibodies. Biosens. Bioelectron. 203, 114034 (2022)
Baker, A.N., Richards, S.-J., Guy, C.S., Congdon, T.R., Hasan, M., Zwetsloot, A.J., et al.: The SARS-COV-2 spike protein binds sialic acids and enables rapid detection in a lateral flow point of care diagnostic device. ACS Cent. Sci. 6, 2046–2052 (2020)
Barnes CO, Jette CA, Abernathy ME, Dam K-MA, Esswein SR, Gristick HB, et al. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature, 588, 682–687 (2020)
Gaebler, C., Wang, Z., Lorenzi, J.C.C., Muecksch, F., Finkin, S., Tokuyama, M., et al.: Evolution of antibody immunity to SARS-CoV-2. Nature 591, 639–644 (2021)
Hansen, J., Baum, A., Pascal, K.E., Russo, V., Giordano, S., Wloga, E., et al.: Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 369, 1010–1014 (2020)
Huang, C., Wen, T., Shi, F.-J., Zeng, X.-Y., Jiao, Y.-J.: Rapid detection of IgM antibodies against the SARS-CoV-2 virus via colloidal gold nanoparticle-based lateral-flow assay. ACS Omega 5, 12550–12556 (2020)
Zeng, L., Li, Y., Liu, J., Guo, L., Wang, Z., Xu, X., et al.: Rapid, ultrasensitive and highly specific biosensor for the diagnosis of SARS-CoV-2 in clinical blood samples. Mater. Chem. Front. 4, 2000–2005 (2020)
Liu, C., Mao, B., Martinez, V., Chen, X., Li, Y., He, L., et al.: A facile assay for rapid detection of COVID-19 antibodies. RSC Adv. 10, 28041–28048 (2020)
Wang, Z., Zheng, Z., Hu, H., Zhou, Q., Liu, W., Li, X., et al.: A point-of-care selenium nanoparticle-based test for the combined detection of anti-SARS-CoV-2 IgM and IgG in human serum and blood. Lab Chip 20, 4255–4261 (2020)
Li, Z., Yi, Y., Luo, X., **ong, N., Liu, Y., Li, S., et al.: Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J. Med. Virol. 92, 1518–1524 (2020)
Peng, T., Liu, X., Adams, L.G., Agarwal, G., Akey, B., Cirillo, J., et al.: Enhancing sensitivity of lateral flow assay with application to SARS-CoV-2. Appl. Phys. Lett. 117, 120601 (2020)
Roda, A., Cavalera, S., Di Nardo, F., Calabria, D., Rosati, S., Simoni, P., et al.: Dual lateral flow optical/chemiluminescence immunosensors for the rapid detection of salivary and serum IgA in patients with COVID-19 disease. Biosens. Bioelectron. 172, 112765 (2021)
Yu, S., Nimse, S.B., Kim, J., Song, K.-S., Kim, T.: Development of a lateral flow strip membrane assay for rapid and sensitive detection of the SARS-CoV-2. Anal. Chem. 92, 14139–14144 (2020)
Zhu, X., Wang, X., Han, L., Chen, T., Wang, L., Li, H., et al.: Multiplex reverse transcription loop-mediated isothermal amplification combined with nanoparticle-based lateral flow biosensor for the diagnosis of COVID-19. Biosens. Bioelectron. 166, 112437 (2020)
Shelite, T.R., Uscanga-Palomeque, A.C., Castellanos-Gonzalez, A., Melby, P.C., Travi, B.L.: Isothermal recombinase polymerase amplification-lateral flow detection of SARS-CoV-2, the etiological agent of COVID-19. J. Virol. Methods 296, 114227 (2021)
**a, S., Chen, X.: Single-copy sensitive, field-deployable, and simultaneous dual-gene detection of SARS-CoV-2 RNA via modified RT–RPA. Cell Discov. 6, 37 (2020)
Kasetsirikul, S., Umer, M., Soda, N., Sreejith, K.R., Shiddiky, M.J.A., Nguyen, N.-T.: Detection of the SARS-CoV-2 humanized antibody with paper-based ELISA. Analyst 145, 7680–7686 (2020)
Gong, F., Wei, H.-x, Qi, J., Ma, H., Liu, L., Weng, J., et al.: Pulling-force spinning top for serum separation combined with paper-based microfluidic devices in COVID-19 ELISA diagnosis. ACS Sensors 6, 2709–2719 (2021)
Clarke, O.J.R., Goodall, B.L., Hui, H.P., Vats, N., Brosseau, C.L.: Development of a SERS-based rapid vertical flow assay for point-of-care diagnostics. Anal. Chem. 89, 1405–1410 (2017)
Noviana, E., Ozer, T., Carrell, C.S., Link, J.S., McMahon, C., Jang, I., et al.: Microfluidic paper-based analytical devices: from design to applications. Chem. Rev. 121, 11835–11885 (2021)
**ong, X., Zhang, J., Wang, Z., Liu, C., **ao, W., Han, J., et al.: Simultaneous multiplexed detection of protein and metal ions by a colorimetric microfluidic paper-based analytical device. BioChip J. 14, 429–437 (2020)
Ozer, T., Henry, C.S.: Paper-based analytical devices for virus detection: recent strategies for current and future pandemics. TrAC, Trends Anal. Chem. 144, 116424 (2021)
Selvakumar, B., Kathiravan, A.: Sensory materials for microfluidic paper based analytical devices—a review. Talanta 235, 122733 (2021)
Kim, S., Hao, Y., Miller, E.A., Tay, D.M.Y., Yee, E., Kongsuphol, P., et al.: Vertical flow cellulose-based assays for SARS-CoV-2 antibody detection in human serum. ACS Sensors 6, 1891–1898 (2021)
Garneret, P., Coz, E., Martin, E., Manuguerra, J.-C., Brient-Litzler, E., Enouf, V., et al.: Performing point-of-care molecular testing for SARS-CoV-2 with RNA extraction and isothermal amplification. PLoS ONE 16, e0243712 (2021)
Weiß, L.J.K., Lubins, G., Music, E., Rinklin, P., Banzet, M., Peng, H., et al.: Single-impact electrochemistry in paper-based microfluidics. ACS Sensors 7, 884–892 (2022)
Li, X., Qin, Z., Fu, H., Li, T., Peng, R., Li, Z., et al.: Enhancing the performance of paper-based electrochemical impedance spectroscopy nanobiosensors: an experimental approach. Biosens. Bioelectron. 177, 112672 (2021)
Liu, Y., Zhan, L., Qin, Z., Sackrison, J., Bischof, J.C.: Ultrasensitive and highly specific lateral flow assays for point-of-care diagnosis. ACS Nano 15, 3593–3611 (2021)
Hsiao, W.W.-W., Le, T.-N., Pham, D.M., Ko, H.-H., Chang, H.-C., Lee, C.-C., et al.: Recent advances in novel lateral flow technologies for detection of COVID-19. Biosensors 11, 295 (2021)
Liu, Y., Zhan, L., Shen, J.W., Baro, B., Alemany, A., Sackrison, J., et al.: fM–aM detection of the SARS-CoV-2 antigen by advanced lateral flow immunoassay based on gold nanospheres. ACS Appl. Nano Mater. 4, 13826–13837 (2021)
Wang, Z., **g, J., Ren, Y., Guo, Y., Tao, N., Zhou, Q., et al.: Preparation and application of selenium nanoparticles in a lateral flow immunoassay for clenbuterol detection. Mater. Lett. 234, 212–215 (2019)
Kim, S.-K., Sung, H., Hwang, S.-H., Kim, M.-N.: A new quantum dot-based lateral flow immunoassay for the rapid detection of Influenza viruses. BioChip J. 16, 175–182 (2022)
You, P.-Y., Li, F.-C., Liu, M.-H., Chan, Y.-H.: Colorimetric and fluorescent dual-mode immunoassay based on plasmon-enhanced fluorescence of polymer dots for detection of PSA in whole blood. ACS Appl. Mater. Interfaces. 11, 9841–9849 (2019)
Qie, Z., Liu, Q., Yan, W., Gao, Z., Meng, W., **ao, R., et al.: Universal and ultrasensitive immunochromatographic assay by using an antigen as a bifunctional element and antialbumin antibody on a test line. Anal. Chem. 91, 9530–9537 (2019)
Hu, J., Zhang, Z.-L., Wen, C.-Y., Tang, M., Wu, L.-L., Liu, C., et al.: Sensitive and quantitative detection of C-reaction protein based on immunofluorescent nanospheres coupled with lateral flow test strip. Anal. Chem. 88, 6577–6584 (2016)
Wang, C., Yang, X., Gu, B., Liu, H., Zhou, Z., Shi, L., et al.: Sensitive and simultaneous detection of SARS-CoV-2-specific IgM/IgG using lateral flow immunoassay based on dual-mode quantum dot nanobeads. Anal. Chem. 92, 15542–15549 (2020)
Posthuma-Trumpie, G.A., Korf, J., van Amerongen, A.: Lateral flow (immuno)assay: its strengths, weaknesses, opportunities and threats. A literature survey. Anal. Bioanal. Chem. 393, 569–582 (2009)
Gao, X., Xu, L.-P., Zhou, S.-F., Liu, G., Zhang, X.: Recent advances in nanoparticles-based lateral flow biosensors. Am. J. Biomed. Sci. 6, 41–57 (2014)
Li, Z., Wang, Y., Wang, J., Tang, Z., Pounds, J.G., Lin, Y.: Rapid and sensitive detection of protein biomarker using a portable fluorescence biosensor based on quantum dots and a lateral flow test strip. Anal. Chem. 82, 7008–7014 (2010)
Chen, Z., Zhang, Z., Zhai, X., Li, Y., Lin, L., Zhao, H., et al.: Rapid and sensitive detection of anti-SARS-CoV-2 IgG, using lanthanide-doped nanoparticles-based lateral flow immunoassay. Anal. Chem. 92, 7226–7231 (2020)
Tuerk, C., Gold, L.: Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990)
Kim, S.M., Kim, J., Noh, S., Sohn, H., Lee, T.: Recent development of aptasensor for Influenza virus detection. BioChip J. 14, 327–339 (2020)
Zou, X., Wu, J., Gu, J., Shen, L., Mao, L.: Application of aptamers in virus detection and antiviral therapy. Front. Microbiol. 10, 1462 (2019)
Chen, Z., Wu, Q., Chen, J., Ni, X., Dai, J.: A DNA aptamer based method for detection of SARS-CoV-2 nucleocapsid protein. Virol. Sin. 35, 351–354 (2020)
Li, J., Zhang, Z., Gu, J., Stacey, H.D., Ang, J.C., Capretta, A., et al.: Diverse high-affinity DNA aptamers for wild-type and B.1.1.7 SARS-CoV-2 spike proteins from a pre-structured DNA library. Nucl. Acids Res. 49, 7267–7279 (2021)
Yang, G., Li, Z., Mohammed, I., Zhao, L., Wei, W., **ao, H., et al.: Identification of SARS-CoV-2-against aptamer with high neutralization activity by blocking the RBD domain of spike protein 1. Signal Transduct. Target. Ther. 6, 227 (2021)
Valero, J., Civit, L., Dupont, D.M., Selnihhin, D., Reinert, L.S., Idorn, M., et al.: A serum-stable RNA aptamer specific for SARS-CoV-2 neutralizes viral entry. Proc. Natl. Acad. Sci. 118, e2112942118 (2021)
Zhang, L., Fang, X., Liu, X., Ou, H., Zhang, H., Wang, J., et al.: Discovery of sandwich type COVID-19 nucleocapsid protein DNA aptamers. Chem. Commun. 56, 10235–10238 (2020)
Kacherovsky, N., Yang, L.F., Dang, H.V., Cheng, E.L., Cardle, I.I., Walls, A.C., et al.: Discovery and characterization of spike N-terminal domain-binding aptamers for rapid SARS-CoV-2 detection. Angew. Chem. Int. Ed. 60, 21211–21215 (2021)
Wu, Z., He, D., Xu, E., Jiao, A., Chughtai, M.F.J., **, Z.: Rapid detection of β-conglutin with a novel lateral flow aptasensor assisted by immunomagnetic enrichment and enzyme signal amplification. Food Chem. 269, 375–379 (2018)
Cheng, N., Liu, Y., Mukama, O., Han, X., Huang, H., Li, S., et al.: A signal-enhanced and sensitive lateral flow aptasensor for the rapid detection of PDGF-BB. RSC Adv. 10, 18601–18607 (2020)
Liu, J., Qin, Q., Zhang, X., Li, C., Yu, Y., Huang, X., et al.: Development of a novel lateral flow biosensor combined with aptamer-based isolation: Application for rapid detection of grouper nervous necrosis virus. Front. Microbiol. 11, 886 (2020)
Du, Y., Liu, D., Wang, M., Guo, F., Lin, J.S.: Preparation of DNA aptamer and development of lateral flow aptasensor combining recombinase polymerase amplification for detection of erythromycin. Biosens. Bioelectron. 181, 113157 (2021)
Ahmad, Au., Liang, H., Ali, S., Abbas, Q., Farid, A., Ali, A., et al.: Cheap, reliable, reusable, thermally and chemically stable fluorinated hexagonal boron nitride nanosheets coated Au nanoparticles substrate for surface enhanced Raman spectroscopy. Sens. Actuators B 304, 127394 (2020)
Yadav, S., Sadique, M.A., Ranjan, P., Kumar, N., Singhal, A., Srivastava, A.K., et al.: SERS based lateral flow immunoassay for point-of-care detection of SARS-CoV-2 in clinical samples. ACS Appl. Bio Mater. 4, 2974–2995 (2021)
Perumal, J., Wang, Y., Attia, A.B.E., Dinish, U.S., Olivo, M.: Towards a point-of-care SERS sensor for biomedical and agri-food analysis applications: a review of recent advancements. Nanoscale 13, 553–580 (2021)
Quarin, S., Strobbia, P.: Recent advances towards point-of-care applications of surface-enhanced raman scattering sensing. Front. Chem. 9, 714113 (2021)
Liu, H., Dai, E., **ao, R., Zhou, Z., Zhang, M., Bai, Z., et al.: Development of a SERS-based lateral flow immunoassay for rapid and ultra-sensitive detection of anti-SARS-CoV-2 IgM/IgG in clinical samples. Sens. Actuators, B Chem. 329, 129196 (2021)
Srivastav, S., Dankov, A., Adanalic, M., Grzeschik, R., Tran, V., Pagel-Wieder, S., et al.: Rapid and sensitive SERS-based lateral flow test for SARS-CoV2-specific IgM/IgG antibodies. Anal. Chem. 93, 12391–12399 (2021)
Chen, S., Meng, L., Wang, L., Huang, X., Ali, S., Chen, X., et al.: SERS-based lateral flow immunoassay for sensitive and simultaneous detection of anti-SARS-CoV-2 IgM and IgG antibodies by using gap-enhanced Raman nanotags. Sens. Actuators B Chem. 348, 130706 (2021)
Serebrennikova, K.V., Byzova, N.A., Zherdev, A.V., Khlebtsov, N.G., Khlebtsov, B.N., Biketov, S.F., et al.: Lateral flow immunoassay of SARS-CoV-2 antigen with SERS-based registration: development and comparison with traditional immunoassays. Biosensors 11, 510 (2021)
Rahimi, H., Salehiabar, M., Barsbay, M., Ghaffarlou, M., Kavetskyy, T., Sharafi, A., et al.: CRISPR systems for COVID-19 diagnosis. ACS Sensors 6, 1430–1445 (2021)
Zhu, X., Wang, X., Li, S., Luo, W., Zhang, X., Wang, C., et al.: Rapid, ultrasensitive, and highly specific diagnosis of COVID-19 by CRISPR-based detection. ACS Sensors 6, 881–888 (2021)
Patchsung, M., Jantarug, K., Pattama, A., Aphicho, K., Suraritdechachai, S., Meesawat, P., et al.: Clinical validation of a Cas13-based assay for the detection of SARS-CoV-2 RNA. Nat. Biomed. Eng. 4, 1140–1149 (2020)
Zhou, Y., Wu, Y., Ding, L., Huang, X., **ong, Y.: Point-of-care COVID-19 diagnostics powered by lateral flow assay. TrAC, Trends Anal. Chem. 145, 116452 (2021)
Broughton, J.P., Deng, X., Yu, G., Fasching, C.L., Servellita, V., Singh, J., et al.: CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 38, 870–874 (2020)
Arizti-Sanz, J., Freije, C.A., Stanton, A.C., Petros, B.A., Boehm, C.K., Siddiqui, S., et al.: Streamlined inactivation, amplification, and Cas13-based detection of SARS-CoV-2. Nat. Commun. 11, 5921 (2020)
Ali, Z., Sánchez, E., Tehseen, M., Mahas, A., Marsic, T., Aman, R., et al.: Bio-SCAN: A CRISPR/dCas9-based lateral flow assay for rapid, specific, and sensitive detection of SARS-CoV-2. ACS Synth. Biol. 11, 406–419 (2022)
Ardakani, A.A., Kanafi, A.R., Acharya, U.R., Khadem, N., Mohammadi, A.: Application of deep learning technique to manage COVID-19 in routine clinical practice using CT images: results of 10 convolutional neural networks. Comput. Biol. Med. 121, 103795 (2020)
Wu, X., Hui, H., Niu, M., Li, L., Wang, L., He, B., et al.: Deep learning-based multi-view fusion model for screening 2019 novel coronavirus pneumonia: A multicentre study. Eur. J. Radiol. 128, 109041 (2020)
Loey, M., Smarandache, F., M. Khalifa NE.: Within the lack of chest COVID-19 X-ray dataset: a novel detection model based on GAN and deep transfer learning. Symmetry 12, 651 (2020)
Apostolopoulos, I.D., Mpesiana, T.A.: Covid-19: Automatic detection from X-ray images utilizing transfer learning with convolutional neural networks. Phys. Eng. Sci. Med. 43, 635–640 (2020)
Acknowledgments
This work was supported by the National Research Council of Science and Technology (NST) and the National Research Foundation (NRF), funded by the Ministry of Science and ICT, Republic of Korea (grant numbers: CRC‐16‐01‐KRICT and NRF-2020M3E9A1043749), and by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (Grant Number: HI20C0363).
Funding
National Research Council of Science and Technology (NST),CRC‐16‐01‐KRICT,Jonghwan Lee,National Research Foundation,NRF-2020M3E9A1043749,Jonghwan Lee,Korea Health Industry Development Institute,HI20C0363,Jonghwan Lee
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, S., Lee, JH. Current Advances in Paper-Based Biosensor Technologies for Rapid COVID-19 Diagnosis. BioChip J 16, 376–396 (2022). https://doi.org/10.1007/s13206-022-00078-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-022-00078-9