Abstract
Non-alcoholic fatty liver disease (NAFLD) is a multifactorial condition with a complex etiology. Its incidence is increasing globally in parallel with the obesity epidemic, and it is now considered the most common liver disease in Western countries. The precise mechanisms underlying the development and progression of NAFLD are complex and still poorly understood. The dysregulation of epigenetic and epitranscriptomic mechanisms is increasingly recognized to play pathogenic roles in multiple conditions, including chronic liver diseases. Here, we have performed a comprehensive analysis of the expression of epigenetic and epitranscriptomic genes in a total of 903 liver tissue samples corresponding to patients with normal liver, obese patients, and patients with non-alcoholic fatty liver (NAFL) and non-alcoholic steatohepatitis (NASH), advancing stages in NAFLD progression. We integrated ten transcriptomic datasets in an unbiased manner, enabling their robust analysis and comparison. We describe the complete landscape of epigenetic and epitranscriptomic genes’ expression along the course of the disease. We identify signatures of genes significantly dysregulated in association with disease progression, particularly with liver fibrosis development. Most of these epigenetic and epitranscriptomic effectors have not been previously described in human NAFLD, and their altered expression may have pathogenic implications. We also performed a comprehensive analysis of the expression of enzymes involved in the metabolism of the substrates and cofactors of epigenetic and epitranscriptomic effectors. This study provides novel information on NAFLD pathogenesis and may also guide the identification of drug targets to treat this condition and its progression towards hepatocellular carcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-alcoholic fatty liver disease (NAFLD) is a multifactorial condition with a complex etiology. Its incidence is increasing globally in parallel with the obesity epidemic, and it is now recognized as the most common liver disease in Western countries. The exact prevalence of NAFLD is difficult to determine as it is often asymptomatic, but recent estimates suggest that it affects more than 30% of the general population [117].
NAFLD encompasses a spectrum of disorders defined by the presence of abnormal lipid accumulation in hepatocytes with or without evidence of liver cell injury [5]. Most patients with NAFLD have non-alcoholic fatty liver (NAFL) defined as simple steatosis or isolated fat accumulation, whereas a subset of NAFL patients can progress to non-alcoholic steatohepatitis (NASH), characterized by steatosis with evidence of hepatocellular injury, inflammation, and fibrosis [5, 10]. The severity of fibrosis correlates with the risk of develo** complications like cirrhosis, liver failure, and hepatocellular carcinoma (HCC) [5, 64]. The progression of NAFLD to end-stage liver disease is depicted in Fig. 1A. The precise mechanisms underlying the development and progression of NAFLD are complex and still poorly understood, but a combination of dietary factors, a poor glycemic control, and genetic predisposition likely play a role [27]. Recent evidence suggests that NAFLD is linked with a range of extrahepatic manifestations such as insulin resistance, dyslipidemia, adipose tissue dysfunction, systemic inflammation, and gut microbiota dysbiosis [27]. In addition to these well-known factors related to obesity, type 2 diabetes, and cardiovascular disease, NAFLD is also associated with several other conditions, including chronic kidney disease and metabolic syndrome [7]. Early detection and treatment of NAFLD are essential to prevent its progression towards end-stage liver disease, which has significant implications for both patients and health care systems [116]. NAFL is diagnosed by the presence of steatosis on liver biopsy, whereas the diagnosis of NASH requires the presence of steatosis, necroinflammation, and fibrosis [69]. The most common cause of death in patients with NAFLD is cardiovascular disease, and the risk of death from NAFLD is increased in patients with NASH [69].
NAFLD progression, genetic variants in NAFLD, and the cross-talk between epigenetic and epitranscriptomic factors and metabolism. A Natural history of NAFLD to end-stage liver disease and hepatocellular carcinoma (HCC). B Genetic variants with a known pathogenic or protective role for NAFLD, epigenetic and epitranscriptomic genes known to be up- (↑) or downregulated (↓) in NAFLD. The DNA methylation status of genes potentially involved in NAFLD pathogenesis is also indicated. C Cross-talk between epigenetic and epitranscriptomic factors and metabolites acting as substrates or inhibitors of these reactions. D Major metabolic pathways involved in the synthesis and conversion of substrates and cofactors of epigenetic and epitranscriptomic enzymatic reactions. Abbreviations can be found in Suppl. Table 8
Epigenetics refer to the heritable modification of gene expression that does not involve changes in the underlying DNA sequence [33]. These modifications can occur in the form of DNA methylation, histone modification, and non-coding RNA-mediated gene regulation [33]. The field of epigenetics is rapidly evolving and holds great promise for the treatment of various diseases including NAFLD [20, 37, 44, 50]. Emerging evidence suggests that epigenetic alterations may contribute to the development of NAFLD by regulating genes involved in lipid metabolism, inflammation, and cell death [65, 97]. DNA methylation is a common epigenetic modification based on the addition of a methyl group to cytosine residues in CpG dinucleotides [74]. This modification can silence gene expression by preventing the binding of transcription factors to DNA and by promoting chromatin compaction. Several studies have found increased DNA methylation in the liver tissue of patients with NAFLD, particularly in genes involved in lipid metabolism and inflammation [42, 111, 122]. The covalent modification of histones is another epigenetic mechanism that can affect chromatin organization and gene expression. These modifications, which include reactions such as acetylation, methylation, and phosphorylation, can change the structure of chromatin, modulate the recruitment of transcriptional regulators, and influence gene expression. Several studies have found altered histone modification patterns in the liver tissue of patients with NAFLD, which suggests that this epigenetic mechanism may also play a role in the development of this disease [19, 54, 111]. Non-coding RNA-mediated gene silencing is another epigenetic mechanism that can regulate gene expression. MicroRNAs (miRNAs) are a type of non-coding RNA that can bind to complementary sequences in mRNA resulting in gene silencing. Several studies have found altered miRNA expression in the liver tissue of patients with NAFLD [111], suggesting that this epigenetic mechanism may play a role in develo** this disease. Overall, the evidence suggests that epigenetic modifications may contribute to the development of NAFLD.
Over the last couple of decades, considerable efforts have been made to characterize the highly dynamic and reversible RNA modifications. More than 150 modifications with regulatory potential have been identified to date and together define the field of epitranscriptomics. Among them, methylation reactions leading to the formation of N6-methyladenosine, N1-methyladenosine, N6,20-O-dimethyladenosine, 5-methylcytosine, and 5-hydroxymethylcytidine are the most extensively investigated [14, 108]. It has become evident that these RNA modifications significantly influence gene expression, and their dysregulation is increasingly being associated with different pathologies including NAFLD [123]. Interestingly, epigenetic and epitranscriptomic mechanisms also rely on reversible enzymatic reactions performed by the so-called epigenetic and epitranscriptomic “writers” and “erasers,” as well as protein-protein and protein-DNA/RNA interactions mediated by “readers,” processes that are amenable to pharmacological targeting [28, 29].
While some gene variants are well-known pathogenic or protective factors in NAFLD (Fig. 1B), the roles of specific epigenetic and epitranscriptomic factors in the progression of this disease remain poorly defined. There are some reports showing the dysregulation in the expression of certain epigenetic and epitranscriptomic factors in NAFL and NASH, as well as altered methylation patterns in key genes for liver disease progression in these conditions (Fig. 1B) [49, 86, 123]. On the other hand, fluctuations in the intracellular levels of specific intermediary metabolites, including those associated with dietary alterations and obesity, can impact on epigenetic and epitranscriptomic mechanisms for which these metabolites behave as substrates or inhibitors [13, 28, 41, 58, 65, 80]. Thus, there is a complex interplay between the intracellular pools of metabolites, related metabolic enzymes, and the epigenetic and epitranscriptomic machinery (Fig. 1C and D). Further studies are needed to confirm the role of epigenetics and epitranscriptomics in NAFLD pathogenesis and to identify potential epigenetic targets for the prevention and treatment of this chronic liver condition. In the present study, we have performed a comprehensive transcriptomic analysis of hepatic epigenetic and epitranscriptomic genes in different cohorts of patients. Our integrated data revealed marked changes in the expression of specific genes in association with the course of the disease. These findings may improve our understanding of NAFLD pathogenesis and pave the way for the identification of novel therapies [93].
Materials and methods
Functional selection of interrogated genes
Epigenetic and epitranscriptomic genes were selected from the literature [11] and Modomics [12], EpiFactors [70], and ChromoHub [61] databases to generate a manually curated list. Among the epigenetic genes, those whose function was related to the methylation and acetylation of DNA and histones, as well as the citrullination (or deimination) of histone arginine residues, were considered conventional epigenetic genes. All other epigenetic genes were classified as non-conventional epigenetic genes. Then, the function of each selected gene was confirmed by the availability in reliable databases (GeneCards, PubMed, and Uniprot) of experimental evidence demonstrating their purported biochemical activity. All the genes with no experimental evidence of functional activity were discarded. Reader and eraser functions were prioritized above reader or cofactor activity to classify the genes, although some of them have several functions as readers and erasers or writers [11]. On the other hand, genes encoding for metabolic enzymes involved in the metabolism of epigenetic cofactors were selected based on the proximity of function to the epigenetic and epitranscriptomic machinery.
Transcriptomic data preprocessing
Transcriptomic high-throughput data (RNA-seq) were downloaded from the NCBI Sequence Read Archive (SRA) in fastq format using SRA Toolkit version 3.0.0. The first step in the workflow involves quality control and preprocessing of the raw RNA-seq data. Adapter sequences and low-quality reads are removed using TrimGalore version 0.6.0 with Cutadapt version 1.18 [53, 71]. Subsequently, clean reads are aligned to the reference genome using a splice-aware aligner STAR version 020201 over genome version hg38 [26]. Aligned reads are then quantified at the gene level using HTseq version 0.11.0 [4]. EdgeR version 3.28.1 [90] for R software version 3.6.3 (hereafter referred to as R) requires raw read counts as input and performs normalization using the trimmed mean of M-values (TMM) method [91]. TMM normalization accounts for library size differences and composition biases, ensuring accurate comparisons between samples.
A “normal liver” was considered when a sample is collected from a healthy liver (without NAFLD, alcoholic hepatitis, virus infection, or cancer). However, the original sequencing studies may have not reported other non-hepatic diseases that could affect the liver.
Data integration
Once all samples were correctly normalized, ComBat [47] from sva R package version 3.44.0 [56] was used to remove the batch effect using sex and disease stage as covariates, the only variables present in all samples or inferable (sex). ComBat integrates and harmonizes high-dimensional biological data from multiple sources by removing batch effects, such as differences in experimental protocols while preserving true biological differences. The method consists on organizing data into a single matrix, identifying batch effects, fitting a linear model that includes both biological factors and batch effects, and using an empirical Bayes framework to estimate model parameters. ComBat then computes adjusted values for each feature, subtracting the estimated batch effect from the original data. The resulting dataset, with reduced batch effects, can be used for downstream analyses more accurately and reliably.
In addition, the adjusted expression matrix was transformed into a positive value matrix adding the minimum value of expression of genes that were negative in some of the samples. Some of the genes were not detected in any of the RNA-seq datasets of liver samples although they could be detected by microarray technology as far as there is no competence between the mRNA molecules. These genes were discarded for not being present in none of the samples of a complete dataset, which inclusion may result in a loss of statistical power and robustness.
Sample clustering and visualization
The robustness of the batch effect removal was tested using four different dimensional reduction techniques including principal component analysis (PCA), uniform manifold approximation and projection (UMAP), and t-distributed stochastic neighbor embedding analysis (t-SNE) with M3C version 1.18.0 [46] and discriminant analysis of principal components (DAPC) (adegenet version 2.1.7) [48]. UMAP excels in preserving the topological structure of the data by approximating the manifold on which it resides and projecting it onto a lower-dimensional space. Its primary advantage lies in its ability to maintain both local and global structures, making it particularly useful for analyzing complex data. On the other hand, t-SNE focuses on minimizing the divergence between two probability distributions: one representing pairwise similarities in high-dimensional space and the other in low-dimensional space. By employing a t-distribution to model similarities in the latter, t-SNE effectively addresses the crowding problem often encountered in other techniques [105]. Its strength lies in revealing intricate, nonlinear structures in the data and generating visually interpretable embeddings. Meanwhile, PCA is a linear dimensionality reduction technique that identifies the principal components or axes in high-dimensional space where data variance is maximized. By projecting data onto the first few principal components, PCA effectively reduces dimensionality while retaining as much original variance as possible. This technique is particularly suited for datasets with linear relationships and Gaussian-like distributions.
Finally, to complete these unsupervised approximations (clustering with unlabeled data), supervised DAPC was carried out. The number of principal components (PC) selected explained 95% of the variance. All the linear discriminants generated were taken because the number of eigenvalues was small enough to make the difference at this point. The explained intergroup variance of each gene of the signature was taken as a variable importance measure, to uncover the most informative genes (genes that explain more than 2% of the variance). The supervised mode of DAPC was also used to reconstruct the evolution of the disease with the whole transcriptome following the same parameter choice, checking if it was congruent with the development described in the bibliography. Using gene set variation analysis (GSVA) package version 1.44.2 [35] with the implemented single-sample gene set enrichment analysis (ssGSEA) method [55], the pathway activity score of a combination of two previously published gene signatures of NAFLD was computed [36, 92]. This activity score was comprised by the combined list of genes divided into two signatures: the first one, with the genes described to be upregulated in the published signatures, and the second one with those described to be downregulated with disease progression. Thereby, this analysis proved the biological relevance of integrated data.
Gene expression analyses
To evaluate transcriptomic differences of these epigenetic, epitranscriptomic, and metabolic genes, edgeR fits a generalized linear model (GLM) to the normalized count data, using the negative binomial distribution to account for both biological and technical variability. Low expression genes were filtered using the filterByExpr function implemented in edgeR with the default settings. Dispersion parameters are estimated using the empirical Bayes method, and statistical tests are performed using the likelihood ratio test or the quasi-likelihood F-test. Multiple testing correction is applied using the Benjamini-Hochberg procedure to control the false discovery rate (FDR).
Regarding the number of differentially expressed genes described throughout the present study, only statistically significant changes were reported for each comparison. To clarify the possible molecular and biological functions of differentially expressed genes, gene set enrichment analysis (GSEA) was conducted using clusterProfiler R package version 4.4.4 [118] with whole msigdb database [99]. Between 200 and 800 significant genes were taken, ordered by the log2 fold change, as input for pathway analyses. The influence of age, sex, and fibrotic stage was tested, and the presence of a potential confounding effect was ruled out.
Hierarchical clustering
Hierarchical clustering of conventional epigenetic genes using Euclidean distance in combination with Ward’s clustering method, with dist function in R over adjusted read counts, resulted in three gene sets or clusters. The most informative and changing cluster was selected for the signature definition from epigenetic and epitranscriptomic gene lists using GSVA. Gene signatures revealed a gradient of expression in both disease stages that, for simplicity of downstream comparisons, were subclassified using the same hierarchical clustering method in three subgroups (low, medium, and high expression levels). This approach resulted in three groups per signature in each disease stage (NAFL and NASH).
Statistical analyses
Genes were considered differentially expressed if the adjusted p-value (FDR method of Benjamini and Hochberg) was lower than 0.05. For exploratory pathway analyses, all genes with p < 0.05 were included. The identified categories were considered significant and up- or downregulated if p-value < 0.05. Gene expression correlation was computed using Pearson’s correlation methods, and significantly correlating gene pairs were plotted using corrplot package version 0.92.
For the statistical significance of more than two groups, the Kruskal-Wallis test was applied. Multiple comparisons were controlled by the false discovery rate (FDR) using the Benjamini and Hochberg correction (Q = 5%). Chi-square tests of homogeneity were used to analyze if the observed proportion of patients in a certain fibrosis stage was distributed as in the integrative dataset (used as expected proportions). Gene signature association with HCC patients’ survival was measured with the survival package in R v3.3-5 as described [104]. HCC gene expression and patients’ related information were obtained from data generated by the TCGA research network (https://www.cancer.gov/tcga). GraphPad Prism 9.0.2 software (GraphPad Prism, San Diego, CA, USA) was used for these statistical analyses and the corresponding boxplots. Data are presented as mean and standard error. Values of p < 0.05 were considered statistically significant.
Results
Integration of transcriptomic data
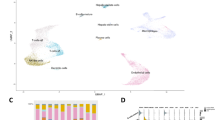
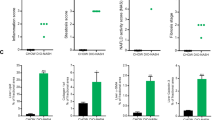
The transcriptomic data of 903 human liver samples were integrated from 10 publicly available datasets (Table 1), including samples from normal (n = 103, 11%), obese (n = 27, 3%), NAFL (n = 194, 21%), and NASH (n = 579, 64%) patients. The dimensionality reduction analyses using t-SNE, UMAP, and PCA techniques revealed a strong tendency for liver samples to cluster by disease stage (Fig. 2A). This finding suggests that the batch effect was successfully mitigated, enabling a more accurate representation of the underlying biological processes. In addition to the unsupervised dimensionality reduction techniques, a supervised DAPC conducted to test if samples would be grouped by independent datasets demonstrated that the batch effect was diminished even using the supervised mode over the integrated data (Fig. 2B). Supervised DAPC based on the similarity of the full transcriptional profile was able to reconstruct the natural history of NAFLD (Fig. 2C). This approach recapitulates at the transcriptional level what is described in the literature in the transition from normal liver, and livers from obese patients, towards NAFL and finally NASH stages. We also found that the activity scores using previously published NAFLD signatures [36, 92] in our integrated data clearly reflected the progression of the disease (Fig. 2D). The soundness of our approach was further supported by the analysis of specific genes within previously reported NAFLD gene signatures [36, 92] (Fig. 2E and Suppl. Fig. 1A), as well as genes functionally linked to NAFLD pathogenesis (Fig. 2F) [1, 2, 31, 36, 78, 92]. As mentioned above, liver fibrosis is a hallmark of NASH progression [10, 109], and consistently, we observed that the expression of a broad range of these genes significantly changed according to the fibrosis stage of liver tissues (F0 to F4) [10] in the NASH cohort (Suppl. Fig. 1B). Altogether, these analyses confirm the robustness of our strategy and support the accuracy of further gene expression studies performed in this integrated dataset.
Bioinformatic and biological validation of the integrated transcriptomic data from human liver gene expression datasets. A Dimensionality reduction analyses using t-SNE, UMAP, and PCA unsupervised techniques. B Supervised dimensionality reduction by DAPC represented as a scatterplot using disease stage as groups (left) and the source (published article, right). C Supervised DAPC representing the centroid and ellipses of 95% confidence interval joined by the maximum similarity path representing the natural history of the disease. D Pathway activity score performed using previously published NAFLD signatures [36, 92]. E Expression of genes previously reported in NAFLD gene signatures in our integrated transcriptomic datasets. F Expression of genes functionally linked to NAFLD pathogenesis in our integrated datasets. Transcriptomic data are expressed as the trimmed mean of M-values (TMM) and grouped according to liver disease classification (Normal, Obese, NAFL, and NASH). p-values were obtained from the Kruskal-Wallis test and adjusted by FDR Benjamini and Hochberg correction (E and F). Values of p < 0.05 were considered statistically significant
Differential gene expression of epigenetic and epitranscriptomic genes
Epigenetic factors comprising 20 families, a total of 419 genes, were selected for the analyses. Of these, 257 genes (11 families) were categorized as conventional epigenetic factors (Table 2). These are considered the most widely described genes that belong to three different categories of epigenetic writers: DNA methyltransferases (DNMTs), protein arginine-methyltransferases (PRMTs), protein lysine-methyltransferases (KMTs), histone acetyl-transferases (HATs); epigenetic erasers: DNA demethylases (TETs), histone-lysine demethylases (HDMs), histone deacetylases (HDACs), and histone deiminases (HDIs); and epigenetic readers: DNA methyl-binding proteins (MBPs), histone methyl readers (HMRs), and histone acetyl readers (HARs). Other 162 genes (9 families) were categorized as non-conventional epigenetic factors (Table 2). These included chromatin remodeling factors (ChrRs) and additional writers, erasers, and readers with other epigenetic activities such as histone-tyrosine phosphatases (HTPs), histone deubiquitylases (HDUs), or phosphorylated histone readers (PHRs) (Table 2). Regarding the epitranscriptomic factors, 6 families involving 137 genes were selected for further analyses, equally comprising the three different categories of epitranscriptomic writers: RNA methyltransferases (RNMTs), pseudo-uridine synthases (PUSs); epitranscriptomic erasers: RNA demethylases (RDMs), RNA hydroxylases (RNHLs); epitranscriptomic readers: methylated RNA readers (MRRs) and others with miscellaneous epitranscriptomic functions, such as RNA acetylation or RNA decap** (Table 2). The classification, function, target, and source of information for selected epigenetic and epitranscriptomic factors are described in Suppl. Table 1 and Suppl. Table 2, respectively. From 556 genes successfully validated as epigenetic and/or epitranscriptomic effectors, 506 were present in all 10 datasets interrogated. Genes not detected in the expression analysis were independently checked set by set without integration. None of them was present in most of the datasets, thereby concluding that their expression in the liver was residual or not present. Out of 419 selected epigenetic genes, 379 were detected, whereas 128 out of 137 epitranscriptomic genes were expressed in the selected transcriptomic datasets.
As can be observed in Fig. 3 and Suppl. Fig. 2, among the 379 epigenetic genes examined (213 conventional, and 133 non-conventional, respectively), profound alterations in their expression were observed for many of them across the different disease stages. When comparing NAFL liver with normal liver, 39 genes (28 conventional and 11 non-conventional) were significantly changed, 12 upregulated (10 conventional and 2 non-conventional) and 27 downregulated (18 conventional and 9 non-conventional). Between NASH and normal liver, we found an increase in the number of differentially expressed genes, 108 of them (62 conventional and 46 non-conventional) were significantly altered, 67 upregulated (35 conventional and 35 non-conventional) and 41 downregulated (27 conventional and 14 non-conventional). The expression of some of these epigenetic effectors, such as DNMT1, SIRT1, SIRT3, PHF2, and ZBTB33, has been previously described to change in NAFL and NASH, confirming the robustness of our findings [18, 32, 79, 86]. We also identified other genes not previously reported to be dysregulated in NAFLD, such as HAT1, SMYD2, CBX5, CBX1, and MPHOSPH8 which were induced, and KAT8 which was downregulated (Suppl. Fig. 3A).
Focusing on epitranscriptomic genes, 128 genes were interrogated of which 13 were differentially expressed (2 upregulated and 11 downregulated) in NAFL, while in NASH 40 genes were differentially expressed (12 upregulated and 28 downregulated) when compared to normal liver samples (Fig. 4). Among these genes, YTHDF3, YTHDC2, RNMT, METTL5, IGFBP3, and TRMT10C were upregulated in both NAFL and NASH liver tissues, while IGFBP1 expression was downregulated, in agreement with previous reports [34] (Suppl. Fig. 3B).
Despite these changes in both types of genes, the total number of them that showed statistically significant up- or downregulation in the NAFL or NASH cohorts was proportionally low. Interestingly, upon closer examination, for both epigenetic and epitranscriptomic genes, there seemed to be subgroups of patients within the NAFL and NASH cohorts that presented similar gene expression patterns (Figs. 3 and 4).
Epigenetic and epitranscriptomic gene signatures
In view of this apparent existence of different subgroups of patients, we next aimed to define the gene signatures that could stratify them. To this end, the observed expression gradients were split using hierarchical clustering on Euclidean distance and three subgroups of patients with low (EpiG-low), medium (EpiG-med), and high (EpiG-high) expression in each signature were generated. Taking gene expression as input for the hierarchical clustering, the Euclidean distance was used as a similarity measure to define specific signatures that captured the differences observed among patients with NAFL or NASH. This signature comprised 188 and 193 genes for NAFL and NASH patients, respectively. To increase the robustness of the stratification, only genes expressed in both disease stages and upregulated in the third cluster (EpiG-high) were selected in the final epigenetic signature, which included a total of 156 conventional epigenetic genes (Fig. 5A and B, and Suppl. Table 3). To define the epitranscriptomic signature the same process was carried out, and three subgroups were defined: EpiT-low, EpiT-med, EpiT-high. In this case, the signature included 123 and 119 genes for NAFL and NASH patients, respectively. When only genes expressed in both disease stages, and upregulated in the third cluster (EpiT-high), were considered, a more robust signature with 119 genes was established (Fig. 6A and B, and Suppl. Table 3). The distribution of male and female NASH patients in the EpiG and EpiT-low, EpiT-med, and EpiT-high clusters was homogeneous (Supplementary Fig. 4).
Heatmap showing the classification of patients according to their epigenetic gene expression signature. A NAFL and B NASH patients. Samples were grouped using hierarchical clustering based on similar expression profiles. Clusters of gene expression are classified as of low (EpiG-low), medium (EpiG-med), and high (EpiG-high) expression. Fold change is established comparing with EpiG-low
Heatmap showing the classification of patients according to their epitranscriptomic gene expression signature. A NAFL and B NASH patients. Samples were grouped using hierarchical clustering based on similar expression profiles. Clusters of gene expression are classified as of low (EpiT-low), medium (EpiT-med), and high (EpiT-high) expression. Fold change is established comparing with EpiT-low
Next, we evaluated if the differential expression of epigenetic and epitranscriptomic genes could be related to the pathological characteristics of the tissues. Therefore, we tested the distribution of patients according to their fibrosis stage (F0–F4) in the different Epi-G and Epi-T subgroups. The proportion of patients in each fibrosis stage was calculated in the full integrative dataset according to the information on biopsy-proven fibrosis as reported by the authors. Thus, 11% of samples from NASH patients showed no fibrosis (F0), 19% of patients corresponded to F1 fibrosis stage, 26% had F2 fibrosis, 30% had F3 fibrosis, and 13% of patients were at F4 fibrosis stage. Then, we examined the distribution of F0–F4 cases across the groups of NASH patients classified in the different epigenetic (EpiG) and epitranscriptomic (EpiT) subgroups. Our analysis revealed an unbalanced distribution of patients classified by fibrosis stage across the EpiG groups, with patients classified as EpiG-high including significantly less F0 cases and the highest proportion of patients with advanced fibrosis (F4) (Fig. 7A–C). Although less marked, a similar observation was made when patients were classified according to EpiT category. F0 cases were very few, while F4 patients were more frequent in the EpiT-high subgroup compared to EpiT-low (Fig. 7D, E).
To explore the molecular landscape and biological characteristics of the NAFL and NASH liver tissue samples classified according to their EpiG and EpiT profiles, we performed Gene Ontology (GO) and gene set enrichment analysis (GSEAs) on the genes differentially expressed (DEGs) between these subgroups (Fig. 8A and B). In the NAFL stage, while the analysis of gene expression among the epigenetic subgroups uncovered relevant processes such as apoptotic signaling, events related to cell adhesion, cell death, or processes involving Notch, TGFβ, and other signaling pathways mediated by GTPases (Fig. 8A and Suppl. Table 4), it was the epitranscriptomic subgroups that exhibited a greater number of significant differences. EpiT-high vs EpiT-low subgroups differences encompassed crucial biological functions associated with the pathophysiology of the disease and its progression. Specifically, individuals classified as EpiT-high displayed enrichments in metabolic pathways (carbohydrates and lipids metabolism, cellular respiration), DNA damage and apoptosis-related mechanisms, and, remarkably, numerous inflammatory events (Fig. 8B and Suppl. Table 5). When considering the NASH stage, the comparison between epigenetic subgroups (EpiG-high vs EpiG-low) revealed greater number of differences than in the NAFL cohort. Notably, differentially expressed genes were primarily associated with important metabolic events (lipids, lipoproteins, bile acids, and mitochondrial metabolism), inflammatory and fibrogenic processes (collagen fibril organization and metabolism, response to TGFβ), and WNT/β-catenin signaling (Fig. 8C and Suppl. Table 6). The comparison between epitranscriptomic subgroups (EpiT-high vs EpiT-low) in NASH primarily revealed processes associated with collagen biosynthesis, response to TGFβ, and pathways involved in cell adhesion and migration, epithelial-mesenchymal transition phenomena, and angiogenesis (Fig. 8D and Suppl. Table 7).
Pathway analyses performed on the differentially expressed genes in the epigenetic and epitranscriptomic “high” and “low” signatures within NAFL and NASH liver samples. A Pathway analysis performed on differentially expressed genes between EpiG-low and EpiG-high clusters in NAFL patients, and B in NASH patients. C Pathway analysis performed on differentially expressed genes between EpiT-low and EpiT-high clusters in the NAFL patients, and D in NASH patients
In view of the key pathogenic role of fibrosis in the progression of NAFLD [5, 10, 30], we evaluated the potential correlation in the expression of epigenetic and epitranscriptomic genes with that of key genes involved in liver extracellular matrix synthesis. We selected the genes coding for fibrillar collagens I, III, and V; interfibrillar collagen VI; and collagen IV, which together with collagen VI is responsible for the capillarization of liver sinusoids, being all of them being upregulated in liver fibrosis [52]. As shown in Suppl. Fig. 5A, there were several epigenetic genes which showed strong correlations with the expression of these collagen genes, including writers, readers, and erasers such as CBX6, CHD3, DNMT1, EHMT2, HDAC7, MLLT3, PHF19, PRDM2, PRMT2, SMARCA4, TP53BP1, and ZBTB4, among others. Although less prominent, we also observed the correlation of collagen gene expression with that of certain epitranscriptomic genes such as ADARB1, APOBEC3G, HEMT1, IGFBP2, PRRC2A, TCOF1, and TRMT1 (Suppl. Fig. 5B). We also included in the analysis the ACTA2 and LRAT genes, coding for α-smooth muscle actin (αSMA) and lecithin-retinol acyltransferase, which are upregulated and downregulated, respectively, in activated liver extracellular matrix (ECM)-producing cells [9]. Most of the epigenetic and epitranscriptomic genes which expression correlated positively with that of collagen genes also correlated with that of ACTA2, but not, or very weakly, with LRAT gene expression (Suppl. Fig. 5A and 5B), further emphasizing their association with liver disease progression.
As previously mentioned, NASH is increasingly recognized as a risk factor for HCC development [5, 64]. Given the strong association between our epigenetic and epitranscriptomic gene signatures established in NAFLD patients with key molecular pathways related to disease progression described above, we explored whether these signatures could also be observed in peritumoral and tumoral tissues from NASH-associated HCC [85]. As shown in Suppl. Fig. 6A and 6B, subgroups of patients showing EpiG-low and EpiG-high signatures, as well as EpiT-low and EpiT-high signatures, were found in peritumoral and HCC tissues. In contrast with these observations, the expression of genes within the EpiG and EpiT signatures was uniformly altered in peritumoral tissues from patients develo** HCC of viral (HBV) [101] or alcoholic [67] etiologies, and these changes were more pronounced in tumoral tissues (Suppl. Fig. 7A-D).
These observations led us to explore if the expression of these epigenetic and epitranscriptomic gene signatures could be linked to the prognosis of HCC patients. To this end, we analyzed data generated by the TCGA network, and as depicted in Supplementary Fig. 8A, we found that HCC patients in the EpiG-high subgroup had significantly poorer survival than those in the EpiG-med and EpiG-low subgroups. However, this difference was not observed for patients in the three EpiT subclasses (Supplementary Fig. 8B).
Candidate epigenetic and epitranscriptomic biomarkers and therapeutic targets in NAFLD
Our current findings evidenced marked alterations in the expression of epigenetic and epitranscriptomic genes and their association with liver disease progression. Therefore, among these genes, there could be good candidates to be developed into biomarkers of disease severity and/or targets for pharmacological intervention. With this in mind, we implemented a supervised DAPC analysis for the selection of those genes that accounted for most of the intergroup variability (i.e., those that explained at least 2% of the variability between groups of samples). We performed these DAPC analyses in liver tissues from patients with NAFL and NASH, and for both the epigenetic (Fig. 9A and B) and epitranscriptomic (Fig. 9C and D) gene signatures.
Most relevant genes contributing to the stratification of patients in the low- and high- EpiG and EpiT clusters in NAFL and NASH patients. A Most relevant epigenetic genes contributing to the stratification of NAFL patients in the EpiG-low and EpiG-high clusters. B Most relevant epigenetic genes contributing to the stratification of NASH patients in the EpiG-low and EPIG-high clusters. C Most relevant epitranscriptomic genes contributing to the stratification of NAFL patients in the EpiT-low and EpiT-high clusters. D Most relevant epitranscriptomic genes contributing to the stratification of NASH patients in the EpiT-low and EpiT-high clusters. In the loading plot, the scale indicates the contribution to the separation between groups of each gene in a scale from 0 to 1. Selected genes explain at least 2% of the variability between low- and high- groups
Consistently, the expression of these largest contributors to the epigenetic and epitranscriptomic NASH signatures showed statistically significant upregulation across the EpiG and EpiT subclasses (Suppl. Fig. 9A and B). Interestingly, the expression of most of these epigenetic and epitranscriptomic genes was significantly dysregulated in NASH patients classified according to their fibrosis stage (F0–F4) (Suppl. Fig. 9C). Regarding the epigenetic gene signature, this analysis showed the upregulation of CHD3 and PRMT2 and the downregulation of TRIM24 expression across all fibrosis stages when compared to the F0 group. Also, DIDO1 and PRDM2 were upregulated, and ATAD2B and BRWD3 were downregulated in at least one fibrosis stage vs F0. For the epitranscriptomic genes, QTRT1 and TRMT112 expression was significantly upregulated across all fibrosis stages, while TRUB2 was upregulated in F1 to F3 compared to F0, and METTL3 and OSGEP were upregulated in F4 compared to F0 (Suppl. Fig. 9D).
Differential expression of metabolic genes
Finally, we also examined the expression of 89 genes encoding enzymes belonging to different metabolic pathways such as that of folates and one carbon (OCM) metabolism, tricarboxylic acid cycle (TCA), and the acetyl-CoA synthesis pathway (ACS). These enzymes are implicated in the synthesis and metabolism of cofactors involved in the activity and regulation of most epigenetic and epitranscriptomic reactions (Fig. 1C and D, and Suppl. Table 8) [13, 28, 58]. Unlike what was found for epigenetic and epitranscriptomic genes, a rather homogeneous expression pattern was found across samples within the NAFL and NASH groups (Fig. 10A). Significant changes were observed in the expression of 12 metabolic genes in NAFL when compared to normal liver samples (Fig. 10B), of which 10 were downregulated and 2 were upregulated. Regarding the expression of metabolic enzymes in NASH liver samples, these changes were much more pronounced. The expression of 35 genes involved in all the above-mentioned metabolic pathways was significantly altered when compared to normal liver tissues, with 23 genes being downregulated and 12 upregulated (Fig. 10C).
Transcriptomic analysis of genes encoding enzymes involved in the syhthesis and conversion of substrates and cofactors of epigenetic and epitranscriptomic enzymatic reactions in the integrated transcriptomic dataset. A Heatmap of the expression of metabolic genes according to liver disease classification. Expression fold change is relative to normal liver. B and C Expression levels of the indicated metabolic genes genes in NAFL and NASH patients relative to normal liver tissues. Values are expressed as the trimmed mean of M-values (TMM) and transformed to log2 fold change versus normal liver. ACS-S/M, short- and medium-chain acyl-coA synthetase; ACS-L/VL, long- and very long-chain acyl-coA synthetase; OCM, one-carbon metabolism; TCA, tricarboxylic acid cycle. p-values were obtained from the Kruskal-Wallis test and adjusted by FDR Benjamini and Hochberg correction
Discussion
Weight loss through nutritional intervention and physical activity can improve liver disease in NAFLD, but currently there are no approved therapies to treat this condition. The progression of fibrosis, the deployment of cirrhosis, and the advent of adverse liver-related complications such as decompensation and HCC development are still quite unpredictable [17, 30, 64, 103]. A better understanding of the molecular mechanisms underlying this complex disease is needed to predict and eventually treat these outcomes. Accumulating experimental and clinical evidence indicates that epigenetic and epitranscriptomic alterations occur during NAFLD development [2, 22, 49, 86, 106, 124].
In our study, we combined ten liver transcriptomic datasets including patients at different stages of NAFLD progression together with obese and normal individuals so they can be interrogated as a single transcriptome. After validating the robust integration of the different datasets, we performed a comprehensive study of the expression of epigenetic and epitranscriptomic modifiers, including 20 epigenetic and 6 epitranscriptomic gene families. Our analyses revealed significant variations in their expression patterns across NAFL and NASH patients in comparison with normal liver samples. As could be anticipated, there were more pronounced statistically significant differences in both families of modifiers in patients with NASH. Among the 379 analyzed samples, 108 epigenetic effectors and 40 epitranscriptomic genes exhibited differential expression. Some of these genes have been previously reported to show altered expression in both NAFL and NASH stages, including epigenetic genes such as DNMT1, SIRT1, and ZBTB33, and epitranscriptomic genes like IGFBP1. However, for most of these genes, their differential expression in NAFL or NASH had not been previously described. Among them, we found HAT1, which codes for a histone acetyl- and succinyltransferase recently reported to be induced in HCC with protumorigenic consequences [114], and SMYD2, a histone methyltransferase also induced in HCCs with poor prognosis and just described to be critical for the development of experimental steatosis in mice [107, 113]. CBX1 and MPHOSPH8, epigenetic readers binding methylated lysine residues, were also markedly upregulated in NASH tissues. Although their upregulation in chronic liver disease has not been reported, both have been involved in carcinogenesis, including HCC development in the case of CBX1 [81, 100]. Similarly, genes coding for epitranscriptomic readers like YTHDF3 and YTHDC2 and epitranscriptomic writers such as RNMT, METTL5, TRNMT10C, and PUS7L were upregulated in NASH tissues and are known to be involved as well in hepatocarcinogenesis [45, 60, 62, 88, 98, 110].
Perhaps most interestingly, we also observed that within the NAFL and NASH cohorts, distinct patterns of expression of both epigenetic and epitranscriptomic genes were apparent among patients. Through hierarchical clustering, we identified three subtypes of patients based on their high, intermediate, or low expression of these two classes of genes. Interestingly, correlation analyses with clinico-pathological information showed that the EpiT-high and the EpiG-high subtypes, but specially the latter, encompassed the highest proportion of patients with advanced fibrosis (F4, cirrhosis). A particularly strong correlation between the expression of epigenetic effectors and collagen genes was evidenced, indicating that these genes are likely involved in disease progression, as previously suggested [8, 75]. This notion was reinforced when we performed functional analyses of the differentially expressed genes between each EpiG and EpiT subtype within NAFL and NASH cohorts. At the NAFL stage, the EpiT-high subgroup presented more active pathways involved in harmful events such as DNA damage, cell death, and inflammatory activation and signaling. Moreover, those patients within the EpiT-high signature also showed marked downregulation in gene sets related to aerobic respiration, mitochondrial function, fatty acid metabolism, and response to oxidative stress and insulin. Although changes in the expression of specific epitranscriptomic and epigenetic genes may have important consequences for disease development, globally, it appears that epitranscriptomic mechanisms could play adaptive and eventually pathogenic roles early in NAFLD progression. This idea fits well with the very dynamic nature of epitranscriptomic modifications, such as N6-methyladenosine, and their key role in the acute regulation of metabolic genes [106, 121]. Moreover, in line with our findings, a recent study found no significant relationship between the expression of N6-methyladenosine regulators and liver fibrosis in NAFLD patients [22]. Growing evidence indicates that epigenetic changes, particularly histones and DNA methylation, can even persist after removal of harmful exposures such as food restriction or bad dietary habits, a phenomenon known as maladaptive epigenetic memory [21, 22, 57, 83]. Therefore, certain persistently dysregulated epigenetic genes could be responsible for this long-term epigenetic memory contributing to disease progression. In this regard, in the NASH stage, the comparison between EpiG-low and EpiG-high revealed a clear stratification of patients with strong alterations in many important biological pathways mostly associated with lipid and steroid metabolism, ribosomal processes, inflammation, and fibrogenesis.
The magnitude of gene expression variation between different biological conditions does not necessarily relate to a more prominent functional role for a specific gene. Nevertheless, given the numerous epigenetic and epitranscriptomic genes differentially expressed between the EpiG and EpiT subclasses of patients, we performed an unbiassed analysis to identify those that contributed most to this stratification. Among the upregulated epigenetic genes were the reader and chromatin remodeller CHD3 [3], the methylated lysine reader DIDO1, the arginine methyltransferase PRMT2, and the lysine methyltransferase PRDM2, not previously described in NASH. Noteworthy, PRMT2 was recently reported to be induced and contribute to HCC tumorigenesis [39], DIDO1 has been involved in numerous types of tumors [16], and PRDM2 is a tumor suppressor epigenetically repressed in HCC [120]. Conversely, the acetylated histone reader TRIM24 was downregulated along fibrosis progression. Interestingly, TRIM24 is known to repress hepatic lipid accumulation and fibrosis in the murine liver [43]; however, its expression in human NAFLD has not been reported before. Among epitranscriptomic genes, we found QTRT, a queuine tRNA ribosyltransferase known to be induced in lung cancer [66], and TRUB2, a pseudouridine synthase essential for mitochondrial protein synthesis [6]. In this set of genes, we also identified TRMT112 which codes for a methyltransferase known to be induced in different cancers including HCC [112]. Most interestingly, TRMT112 was recently described to form an N6-methyladenosine methyltransferase complex with METTL5, which we also found upregulated in NASH, that remodels fatty acid metabolism and promotes HCC tumorigenesis [84].
In this study, we have identified for the first time numerous epigenetic and epitranscriptomic genes dysregulated along the course of human NAFLD. Interestingly, being NAFLD primarily a metabolic condition, the pathophysiological roles described in the literature for most of these genes are related to carcinogenic processes, including HCC development [15, 29, 106, 123]. Indeed, NAFLD is an emerging risk factor for liver cancer, particularly in patients with type 2 diabetes [64]. While just changes in the expression of epigenetic and epitranscriptomic effectors are not likely to be tumorigenic per se, these alterations may indeed facilitate metabolic rewiring and pave the way for neoplastic transformation triggered by NAFLD-HCC driving mutations [85]. Supporting this notion, there is already experimental evidence showing that drug-mediated inhibition of epigenetic reprogramming can improve NASH and fibrosis progression [73] and prevent NASH-associated HCC development [51], illustrating the therapeutic potential of targeting these pathways. Importantly, when we analyzed the TCGA HCC cohort, we observed that patients within the EpiG-high subclass had significantly lower survival than those in the EpiG-med or EpiG-low subclasses. This finding suggests that the epigenetic genes comprised in a signature that defines advanced NAFLD stages may also be involved in HCC progression. Nevertheless, it would be interesting to test the predictive capacity of this EpiG signature in a selected cohort of NASH-related HCC patients.
Epigenetic and epitranscriptomic mechanisms are complex and intertwined, and fluctuations in the expression of these effectors are likely to influence the course of NAFLD progression. However, gene expression regulation at the chromatin and epitranscriptomic levels involves another layer of complexity linked to the intrinsic enzymatic nature of these processes. As previously mentioned, epigenetic and epitranscriptomic writers and erasers utilize a range of metabolites as substrates and cofactors (Fig. 1C). For chromatin regulatory mechanisms, this interaction has proven so relevant that epigenetic processes can even act as sensors of the activity of central metabolic pathways, including pyruvate metabolism and the TCA cycle, acetyl-CoA, NAD+, FAD, S-adenosylmethionine (SAM)/OCM, and energetic metabolism (AMP/ATP), and in turn can regulate the expression of metabolism-related genes [13, 21, 58]. Indeed, the intracellular, and even intranuclear, levels of metabolites that behave as substrates or inhibitors of epigenetic and epitranscriptomic reactions, such as SAM and S-adenosylhomocysteine (SAH) for methylation/demethylation; α-ketoglutarate (α-KG), fumarate, and succinate for demethylation; and acetyl-CoA and NAD for acetylation/deacetylation, strongly influence the homeostasis of these processes [13, 58]. Given the profound metabolic alterations occurring in NAFLD, changes in the levels of these metabolites are likely to happen, albeit their direct measurement in human liver tissues is quite challenging [40, 95]. Our transcriptomic analyses revealed profound alterations in the expression of genes coding for key enzymes in the metabolism of epigenetic and epitranscriptomic substrates and cofactors (Fig. 1D). These included acyl-CoA synthetases involved in fatty acid metabolism and acetyl-CoA synthesis such as ACSL4, which is also markedly induced in HCC [89], the key TCA cycle enzyme citrate synthase (CS), and the gluconeogenic rate-limiting enzyme PCK1, also relevant for acetyl-CoA availability [115]. Interestingly, we also observed a marked reduction in the expression of SDHA, consistent with observations in murine NASH models [63] and with the increased levels of succinate found in human NASH liver tissues [95]. Importantly, succinate can compete with α-KG and inhibit enzymes involved in histone, DNA, and RNA demethylation [13, 59]. Noteworthy, we found significant alterations in the expression of genes involved in folate metabolism and OCM. Low serum folate levels are consistently found in NAFLD patients; however, the expression of enzymes involved in folate metabolism has not been examined in the liver of these patients [24, 96]. We found induced and repressed levels of DHFR and FOLH1 expression, respectively, in NASH patients. Impaired folate metabolism may affect the synthesis of SAM from homocysteine, and enzymes involved in the metabolism of this non-proteinogenic amino acid were also significantly affected in these patients [96]. Expression of BHMT, an enzyme that converts homocysteine back to methionine using betaine as a methyl donor, was significantly repressed in NASH, as was the expression of CHDH, the enzyme that synthesizes betaine from choline [80]. Choline deficiency is well known to promote NASH in murine models, and choline metabolism is impaired at different levels in NASH patients [24]. Interestingly, BHMT knockout in mice results in liver SAM depletion along with NAFLD and HCC development [102]. We confirmed the downregulation of GNMT and MAT1A expression in NASH [68, 77], key enzymes in the consumption and synthesis of SAM in the liver, respectively, and also found a marked reduction in ACHY expression, which codes for the enzyme in charge of metabolizing SAH into adenosine and homocysteine [80]. Experimental studies in genetically modified mice and diet-induced NAFLD suggest that alterations in GNMT, MAT1A, and ACHY expression can contribute to NASH development in humans [72, 80, 87]. These transcriptional alterations in OCM-related genes can significantly modify on the hepatocellular SAM/SAH ratio, and therefore impact on numerous epigenetic and epitranscriptomic methylation reactions [13].
The changes in metabolic gene expression summarized above may affect the levels of key metabolites in epigenetic and epitranscriptomic reactions; however, there is little information available on the actual levels of these molecules in healthy liver and NAFLD. Emerging technologies such as spatial metabolomics will be crucial in providing this information [94]. Nevertheless, recent molecular and genetic studies reveal that the function of many epigenetic enzymes extends beyond their catalytic activity [76]. This “epigenetic moonlighting” must be taken into account when evaluating the contribution of these effectors to the pathogenesis of NAFLD, and also at the time of designing potential therapeutic interventions which perhaps would need also to look beyond their enzymatic inhibition [23].
In this study, we have provided a comprehensive overview of the expression of epigenetic and epitranscriptomic genes in NAFLD. Of course, it will be important to validate these transcriptional changes at the protein level, and to identify the key target genes downstream these epi-regulators. Likewise, it will be very interesting to understand the upstream mechanisms that control the expression of epigenetic and epitranscriptomic genes in the context of NAFLD. These are likely to be multifarious, and involve signaling pathways associated with the prevalent pro-inflammatory and lipotoxic environment that characterizes NAFLD. Interestingly, the presence of the PNPLA3 I148M variant, strongly associated with NASH progression [27], has been recently reported to trigger potent pro-inflammatory signaling in cultured hepatocytes [82]. Nonetheless, our work may help in the elucidation of the pathogenic mechanisms of this complex disease, and also for the identification of pathways contributing to HCC development in this condition.
Abbreviations
- NAFLD:
-
Non-alcoholic fatty liver disease
- NAFL:
-
Non-alcoholic fatty liver
- NASH:
-
Non-alcoholic steatohepatitis
- HCC:
-
Hepatocellular carcinoma
- SRA:
-
Sequence Read Archive
- TMM:
-
Trimmed mean of M-values
- PCA:
-
Principal component analysis
- UMAP:
-
Uniform manifold approximation and projection
- t-SNE:
-
t-distributed stochastic neighbor embedding analysis
- PC:
-
Principal components
- GSVA:
-
Gene set variation analysis
- ssGSEA:
-
Single-sample gene set enrichment analysis
- GLM:
-
Generalized linear model
- FDR:
-
False discovery rate
- GSEA:
-
Gene set enrichment analysis
- EpiG:
-
Epigenetic
- EpiT:
-
Epitranscriptomic
- GO:
-
Gene Ontology
- DEGs:
-
Differentially expressed genes
- ECM:
-
Extracellular matrix
- DAPC:
-
Discriminant analysis of principal components
- OCM:
-
One carbon metabolism
- TCA:
-
Tricarboxylic acid cycle
References
Ægidius HM, Veidal SS, Feigh M, Hallenborg P, Puglia M, Pers TH, Vrang N, Jelsing J, Kornum BR, Blagoev B, Rigbolt KTG (2020) Multi-omics characterization of a diet-induced obese model of non-alcoholic steatohepatitis. Sci Rep 10:1148. https://doi.org/10.1038/s41598-020-58059-7
Ahrens M, Ammerpohl O, Von Schönfels W, Kolarova J, Bens S, Itzel T, Teufel A, Herrmann A, Brosch M, Hinrichsen H, Erhart W, Egberts J, Sipos B, Schreiber S, Häsler R, Stickel F, Becker T, Krawczak M, Röcken C et al (2013) DNA methylation analysis in nonalcoholic fatty liver disease suggests distinct disease-specific and remodeling signatures after bariatric surgery. Cell Metab 18:296–302. https://doi.org/10.1016/j.cmet.2013.07.004
Alendar A, Berns A (2021) Sentinels of chromatin: chromodomain helicase DNA-binding proteins in development and disease. Genes Dev 35:1403–1430. https://doi.org/10.1101/GAD.348897.121
Anders S, Pyl PT, Huber W (2015) HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169. https://doi.org/10.1093/bioinformatics/btu638
Anstee QM, Reeves HL, Kotsiliti E, Govaere O, Heikenwalder M (2019) From NASH to HCC: current concepts and future challenges. Nat Rev Gastroenterol Hepatol 16:411–428. https://doi.org/10.1038/s41575-019-0145-7
Antonicka H, Choquet K, Lin Z, Gingras A, Kleinman CL, Shoubridge EA (2017) A pseudouridine synthase module is essential for mitochondrial protein synthesis and cell viability. EMBO Rep 18:28–38. https://doi.org/10.15252/embr.201643391
Armstrong MJ, Adams LA, Canbay A, Syn WK (2014) Extrahepatic complications of nonalcoholic fatty liver disease. Hepatology 59:1174–1197. https://doi.org/10.1002/hep.26717
Barcena-Varela M, Colyn L, Fernandez-Barrena MG (2019) Epigenetic mechanisms in hepatic stellate cell activation during liver fibrosis and carcinogenesis. Int J Mol Sci 20. https://doi.org/10.3390/ijms20102507
Barcena-Varela M, Paish H, Alvarez L, Uriarte I, Latasa MU, Santamaria E, Recalde M, Garate M, Claveria A, Colyn L, Arechederra M, Iraburu MJ, Milkiewicz M, Milkiewicz P, Sangro B, Robinson SM, French J, Pardo-Saganta A, Oyarzabal J et al (2020) Epigenetic mechanisms and metabolic reprogramming in fibrogenesis: dual targeting of G9a and DNMT1 for the inhibition of liver fibrosis. Gut. https://doi.org/10.1136/gutjnl-2019-320205
Bedossa P (2017) Pathology of non-alcoholic fatty liver disease. Liver Int 37:85–89
Biswas S, Rao CM (2018) Epigenetic tools (the writers, the readers and the erasers) and their implications in cancer therapy. Eur J Pharmacol 837:8–24. https://doi.org/10.1016/j.ejphar.2018.08.021
Boccaletto P, Stefaniak F, Ray A, Cappannini A, Mukherjee S, Purta E, Kurkowska M, Shirvanizadeh N, Destefanis E, Groza P, Avşar G, Romitelli A, Pir P, Dassi E, Conticello SG, Aguilo F, Bujnicki JM (2022) MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Res 50:D231–D235. https://doi.org/10.1093/nar/gkab1083
Boon R (2021) Metabolic fuel for epigenetic: nuclear production meets local consumption. Front Genet 12:1–15. https://doi.org/10.3389/fgene.2021.768996
Boulias K, Greer EL (2023) Biological roles of adenine methylation in RNA. Nat Rev Genet 24:143–160. https://doi.org/10.1038/s41576-022-00534-0
Braghini MR, Lo Re O, Romito I, Fernandez-Barrena MG, Barbaro B, Pomella S, Rota R, Vinciguerra M, Avila MA, Alisi A (2022) Epigenetic remodelling in human hepatocellular carcinoma. J Exp Clin Cancer Res 41:1–21. https://doi.org/10.1186/s13046-022-02297-2
Braig S, Bosserhoff AK (2013) Death inducer-obliterator 1 (Dido1) is a BMP target gene and promotes BMP-induced melanoma progression. Oncogene 32:837–848. https://doi.org/10.1038/onc.2012.115
Brennan PN, Elsharkawy AM, Kendall TJ, Loomba R, Mann DA, Fallowfield JA (2023) Antifibrotic therapy in nonalcoholic steatohepatitis: time for a human-centric approach. Nat Rev Gastroenterol Hepatol. https://doi.org/10.1038/s41575-023-00796-x
Bricambert J, Alves-Guerra MC, Esteves P, Prip-Buus C, Bertrand-Michel J, Guillou H, Chang CJ, Vander Wal MN, Canonne-Hergaux F, Mathurin P, Raverdy V, Pattou F, Girard J, Postic C, Dentin R (2018) The histone demethylase Phf2 acts as a molecular checkpoint to prevent NAFLD progression during obesity. Nat Commun 9. https://doi.org/10.1038/s41467-018-04361-y
Cai C, Yu H, Huang G, Du X, Yu X, Zhou Y, Shen W (2018) Histone modifications in fatty acid synthase modulated by carbohydrate responsive element binding protein are associated with non-alcoholic fatty liver disease. Int J Mol Med 42:1215–1228. https://doi.org/10.3892/ijmm.2018.3702
Cai Q, Gan C, Tang C, Wu H, Gao J (2021) Mechanism and therapeutic opportunities of histone modifications in chronic liver disease. Front Pharmacol 12:1–20. https://doi.org/10.3389/fphar.2021.784591
Carlberg C, Velleuer E (2022) Nutrition and epigenetic programming. Curr Opin Clin Nutr Metab Care:259–265. https://doi.org/10.1097/mco.0000000000000900
Cheng W, Li M, Zhang L, Zhou C, Yu S, Peng X, Zhang W, Zhang W (2022) New roles of N6-methyladenosine methylation system regulating the occurrence of non-alcoholic fatty liver disease with N6-methyladenosine-modified MYC. Front Pharmacol 13:1–15. https://doi.org/10.3389/fphar.2022.973116
Conery AR, Rocnik JL, Trojer P (2022) Small molecule targeting of chromatin writers in cancer. Nat Chem Biol 18:124–133. https://doi.org/10.1038/s41589-021-00920-5
Da Silva RP, Eudy BJ, Deminice R (2020) One-carbon metabolism in fatty liver disease and fibrosis: one-carbon to rule them all. J Nutr 150:994–1003. https://doi.org/10.1093/jn/nxaa032
de Oliveira DT, de Paiva NCN, Carneiro CM, Guerra-Sá R (2022) Dynamic changes in hepatic DNA methylation during the development of nonalcoholic fatty liver disease induced by a high-sugar diet. J Physiol Biochem 78:763–775. https://doi.org/10.1007/s13105-022-00900-w
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21. https://doi.org/10.1093/bioinformatics/bts635
Eslam M, George J (2020) Genetic contributions to NAFLD: leveraging shared genetics to uncover systems biology. Nat Rev Gastroenterol Hepatol 17:40–52. https://doi.org/10.1038/s41575-019-0212-0
Esteve-Puig R, Bueno-Costa A, Esteller M (2020) Writers, readers and erasers of RNA modifications in cancer. Cancer Lett 474:127–137. https://doi.org/10.1016/j.canlet.2020.01.021
Fernández-Barrena MG, Arechederra M, Colyn L, Berasain C, Avila MA (2020) Epigenetics in hepatocellular carcinoma development and therapy: the tip of the iceberg. JHEP Reports 2:100167. https://doi.org/10.1016/j.jhepr.2020.100167
Friedman SL, Pinzani M (2022) Hepatic fibrosis 2022: unmet needs and a blueprint for the future. Hepatology 75:473–488. https://doi.org/10.1002/hep.32285
Fujita K, Nozaki Y, Wada K, Yoneda M, Fujimoto Y, Fujitake M, Endo H, Takahashi H, Inamori M, Kobayashi N, Kirikoshi H, Kubota K, Saito S, Nakajima A (2009) Dysfunctional very-low-density lipoprotein synthesis and release is a key factor in nonalcoholic steatohepatitis pathogenesis. Hepatology 50:772–780. https://doi.org/10.1002/hep.23094
Ge J, Bai Y, Tang B, Wei D, Yan M (2021) The gene signature associated with hepatocellular carcinoma in patients with nonalcoholic fatty liver disease. J Oncol 2021. https://doi.org/10.1155/2021/6630535
Goldberg AD, Allis CD, Bernstein E (2007) Epigenetics: a landscape takes shape. Cell 128:635–638. https://doi.org/10.1016/j.cell.2007.02.006
Hagemann CA, Legart C, Møllerhøj MB, Madsen MR, Hansen HH, Kønig MJ, Helgstrand F, Hjørne FP, Toxværd A, Langhoff JL, Kielgast UL, Gluud LL, Ægidius H, Rigbolt KTG, Vilsbøll T, Jelsing J, Knop FK (2022) A liver secretome gene signature-based approach for determining circulating biomarkers of NAFLD severity. PloS One 17:1–19. https://doi.org/10.1371/journal.pone.0275901
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinformatics 14. https://doi.org/10.1186/1471-2105-14-7
Hasin-Brumshtein Y, Sakaram S, Khatri P, He YD, Sweeney TE (2022) A robust gene expression signature for NASH in liver expression data. Sci Rep 12:2571. https://doi.org/10.1038/s41598-022-06512-0
Heerboth S, Lapinska K, Snyder N, Leary M, Rollinson S, Sarkar S (2014) Use of epigenetic drugs in disease: an overview. Genet Epigenet 1:9–19. https://doi.org/10.4137/GeG.s12270
Hlady RA, Robertson KD (2023) Epigenetic memory of environmental exposures as a mediator of liver disease. Hepatology. https://doi.org/10.1097/HEP.0000000000000414
Hu G, Yan C, **e P, Cao Y, Shao J, Ge J (2020) PRMT2 accelerates tumorigenesis of hepatocellular carcinoma by activating Bcl2 via histone H3R8 methylation. Exp Cell Res 394:112152. https://doi.org/10.1016/j.yexcr.2020.112152
Hughey CC, Puchalska P, Crawford PA (2022) Integrating the contributions of mitochondrial oxidative metabolism to lipotoxicity and inflammation in NAFLD pathogenesis. Biochim Biophys Acta - Mol Cell Biol Lipids 1867:159209. https://doi.org/10.1016/j.bbalip.2022.159209
Huo M, Zhang J, Huang W, Wang Y (2021) Interplay among metabolism, epigenetic modifications, and gene expression in cancer. Front Cell Dev Biol 9:1–20. https://doi.org/10.3389/fcell.2021.793428
Hyun J, Jung Y (2020) Dna methylation in nonalcoholic fatty liver disease. Int J Mol Sci 21:1–26. https://doi.org/10.3390/ijms21218138
Jiang S, Minter LC, Stratton SA, Yang P, Abbas HA, Akdemir ZC, Pant V, Post S, Gagea M, Lee RG, Lozano G, Barton MC (2015) TRIM24 suppresses development of spontaneous hepatic lipid accumulation and hepatocellular carcinoma in mice. J Hepatol 62:371–379. https://doi.org/10.1016/j.jhep.2014.09.026
** N, George TL, Otterson GA, Verschraegen C, Wen H, Carbone D, Herman J, Bertino EM, He K (2021) Advances in epigenetic therapeutics with focus on solid tumors. Clin Epigenetics 13:1–27. https://doi.org/10.1186/s13148-021-01069-7
** Z, Song M, Wang J, Zhu W, Sun D, Liu H, Shi G (2022) Integrative multiomics evaluation reveals the importance of pseudouridine synthases in hepatocellular carcinoma. Front Genet 13:1–20. https://doi.org/10.3389/fgene.2022.944681
John CR, Watson D, Russ D, Goldmann K, Ehrenstein M, Pitzalis C, Lewis M, Barnes M (2020) M3C: Monte Carlo reference-based consensus clustering. Sci Rep 10:1–14. https://doi.org/10.1038/s41598-020-58766-1
Johnson WE, Li C, Rabinovic A (2007) Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8:118–127. https://doi.org/10.1093/biostatistics/kxj037
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Jonas W, Schürmann A (2021) Genetic and epigenetic factors determining NAFLD risk. Mol Metab 50:101111. https://doi.org/10.1016/j.molmet.2020.101111
Jones PA, Ohtani H, Chakravarthy A, De Carvalho DD (2019) Epigenetic therapy in immune-oncology. Nat Rev Cancer 19:151–161. https://doi.org/10.1038/s41568-019-0109-9
Jühling F, Hamdane N, Crouchet E, Li S, El Saghire H, Mukherji A, Fujiwara N, Oudot MA, Thumann C, Saviano A, Roca Suarez AA, Goto K, Masia R, Sojoodi M, Arora G, Aikata H, Ono A, Tabrizian P, Schwartz M et al (2021) Targeting clinical epigenetic reprogramming for chemoprevention of metabolic and viral hepatocellular carcinoma. Gut 70:157–169. https://doi.org/10.1136/gutjnl-2019-318918
Karsdal MA, Nielsen SH, Leeming DJ, Langholm LL, Nielsen MJ, Manon-Jensen T, Siebuhr A, Gudmann NS, Rønnow S, Sand JM, Daniels SJ, Mortensen JH, Schuppan D (2017) The good and the bad collagens of fibrosis – their role in signaling and organ function. Adv Drug Deliv Rev 121:43–56. https://doi.org/10.1016/j.addr.2017.07.014
Kechin A, Boyarskikh U, Kel A, Filipenko M (2017) CutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J Comput Biol 24:1138–1143. https://doi.org/10.1089/cmb.2017.0096
Kim JH, Jung DY, Nagappan A, Jung MH (2018) Histone H3K9 demethylase JMJD2B induces hepatic steatosis through upregulation of PPARγ2. Sci Rep 8:2–12. https://doi.org/10.1038/s41598-018-31953-x
Lee E, Chuang HY, Kim JW, Ideker T, Lee D (2008) Inferring pathway activity toward precise disease classification. PLoS Comput Biol 4. https://doi.org/10.1371/journal.pcbi.1000217
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD (2012) The SVA package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28:882–883. https://doi.org/10.1093/bioinformatics/bts034
Leung A, Trac C, Du J, Natarajan R, Schones DE (2016) Persistent chromatin modifications induced by high fat diet. J Biol Chem 291:10446–10455. https://doi.org/10.1074/jbc.M115.711028
Li X, Egervari G, Wang Y, Berger SL, Lu Z (2018) Regulation of chromatin and gene expression by metabolic enzymes and metabolites. Nat Rev Mol Cell Biol 19:563–578. https://doi.org/10.1038/s41580-018-0029-7
Liu J, Huang H, Zhang M, Qing G, Liu H (2023) Intertwined regulation between RNA m6A modification and cancer metabolism. Cell Insight 2:100075. https://doi.org/10.1016/j.cellin.2022.100075
Liu J, Sun G, Pan S, Qin M, Ouyang R, Li Z, Huang J (2020) The Cancer Genome Atlas (TCGA) based m6A methylation-related genes predict prognosis in hepatocellular carcinoma. Bioengineered 11:759–768. https://doi.org/10.1080/21655979.2020.1787764
Liu L, Zhen XT, Denton E, Marsden BD, Schapira M (2012) ChromoHub: a data hub for navigators of chromatin-mediated signalling. Bioinformatics 28:2205–2206. https://doi.org/10.1093/bioinformatics/bts340
Liu T, Sun L, Li Z-z, Yang K, Chen J-m, Han X-y, Qi L-m, Zhou X-g, Wang P (2023) The m6A/m5C/m1A regulator genes signature reveals the prognosis and is related with immune microenvironment for hepatocellular carcinoma. BMC Gastroenterol 23:1–12. https://doi.org/10.1186/s12876-023-02776-6
Liu X, **e L, Du K, Liu C, Zhang N, Gu C, Wang Y, Abdelmalek MF, Dong W, Liu X, Niu C, Yang C, Diehl AM, Wu J (2020) Succinate-GPR-91 receptor signalling is responsible for nonalcoholic steatohepatitis-associated fibrosis: effects of DHA supplementation. Liver Int 40:830–843. https://doi.org/10.1111/liv.14370
Llovet JM, Willoughby CE, Singal AG, Greten TF, Heikenwälder M, El-Serag HB, Finn RS, Friedman SL (2023) Nonalcoholic steatohepatitis-related hepatocellular carcinoma: pathogenesis and treatment. Nat Rev Gastroenterol Hepatol. https://doi.org/10.1038/s41575-023-00754-7
Longo R, Ferrari A, Zocchi M, Crestani M (2017) Of mice and humans through the looking glass: “reflections” on epigenetics of lipid metabolism. Mol Aspects Med 54:16–27. https://doi.org/10.1016/j.mam.2017.01.005
Ma Q, He J (2020) Enhanced expression of queuine tRNA-ribosyltransferase 1 (QTRT1) predicts poor prognosis in lung adenocarcinoma. Ann Transl Med 8:1658–1658. https://doi.org/10.21037/atm-20-7424
Makowska Z, Boldanova T, Adametz D, Quagliata L, Vogt JE, Dill MT, Matter MS, Roth V, Terracciano L, Heim MH (2016) Gene expression analysis of biopsy samples reveals critical limitations of transcriptome-based molecular classifications of hepatocellular carcinoma. J Pathol Clin Res 2:80–92. https://doi.org/10.1002/cjp2.37
Männistö V, Kaminska D, Kärjä V, Tiainen M, de Mello VD, Hanhineva K, Soininen P, Ala-Korpela M, Pihlajamäki J (2019) Total liver phosphatidylcholine content associates with non-alcoholic steatohepatitis and glycine N-methyltransferase expression. Liver Int 39:1895–1905. https://doi.org/10.1111/liv.14174
Mantovani A, Scorletti E, Mosca A, Alisi A, Byrne CD, Targher G (2020) Complications, morbidity and mortality of nonalcoholic fatty liver disease. Metabolism 111:154170. https://doi.org/10.1016/j.metabol.2020.154170
Marakulina D, Vorontsov IE, Kulakovskiy IV, Lennartsson A, Drabløs F, Medvedeva YA (2023) EpiFactors 2022: expansion and enhancement of a curated database of human epigenetic factors and complexes. Nucleic Acids Res 51:D564–D570. https://doi.org/10.1093/nar/gkac989
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17:10. https://doi.org/10.14806/ej.17.1.200
Mato JM, Alonso C, Noureddin M, Lu SC (2019) Biomarkers and subtypes of deranged lipid metabolism in non-alcoholic fatty liver disease. World J Gastroenterol 25:3009–3020. https://doi.org/10.3748/wjg.v25.i24.3009
Middleton SA, Rajpal N, Cutler L, Mander P, Rioja I, Prinjha RK, Rajpal D, Agarwal P, Kumar V (2018) BET inhibition improves NASH and liver fibrosis. Sci Rep 8:1–13. https://doi.org/10.1038/s41598-018-35653-4
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38:23–38. https://doi.org/10.1038/npp.2012.112
Moran-Salvador E, Mann J (2017) Epigenetics and liver fibrosis. Cell Mol Gastroenterol Hepatol 4:125–134. https://doi.org/10.1016/j.jcmgh.2017.04.007
Morgan MAJ, Shilatifard A (2023) Epigenetic moonlighting: catalytic-independent functions of histone modifiers in regulating transcription. Sci Adv 9:eadg6593. https://doi.org/10.1126/sciadv.adg6593
Moylan CA, Pang H, Dellinger A, Suzuki A, Garrett ME, Guy CD, Murphy SK, Ashley-Koch AE, Choi SS, Michelotti GA, Hampton DD, Chen Y, Tillmann HL, Hauser MA, Abdelmalek MF, Diehl AM (2014) Hepatic gene expression profiles differentiate presymptomatic patients with mild versus severe nonalcoholic fatty liver disease. Hepatology 59:471–482. https://doi.org/10.1002/hep.26661
Murphy SK, Yang H, Moylan CA, Pang H, Dellinger A, Abdelmalek MF, Garrett ME, Ashley-Koch A, Suzuki A, Tillmann HL, Hauser MA, Diehl AM (2013) Relationship between methylome and transcriptome in patients with nonalcoholic fatty liver disease. Gastroenterology 145:1076–1087. https://doi.org/10.1053/j.gastro.2013.07.047
Nassir F, Ibdah JA (2016) Sirtuins and nonalcoholic fatty liver disease. World J Gastroenterol 22:10084–10092. https://doi.org/10.3748/wjg.v22.i46.10084
Pajares MA, Pérez-Sala D (2018) Mammalian sulfur amino acid metabolism: a nexus between redox regulation, nutrition, epigenetics, and detoxification. Antioxid Redox Signal 29:408–452. https://doi.org/10.1089/ars.2017.7237
Pan C, Luo N, Guo K, Wang W, Li L, Fan N, Tian Y (2022) Members of the chromobox family have prognostic value in hepatocellular carcinoma. Front Genet 13:1–13. https://doi.org/10.3389/fgene.2022.887925
Park J, Zhao Y, Zhang F, Zhang S, Kwong AC, Zhang Y, Hoffmann H-H, Bushweller L, Wu X, Ashbrook AW, Stefanovic B, Chen S, Branch AD, Mason CE, Jung JU, Rice CM, Wu X (2023) IL-6/STAT3 axis dictates the PNPLA3-mediated susceptibility to non-alcoholic fatty liver disease. J Hepatol 78:45–56. https://doi.org/10.1016/j.jhep.2022.08.022
Paulus T, Wernecke L, Lundie A, Friedrich J, Verrel J, Rawish T, Weissbach A, Frings C, Beste C, Bäumer T, Münchau A (2023) The role of the left inferior parietal cortex in Gilles de la Tourette syndrome—an rTMS study. Biomedicines 11:1–9. https://doi.org/10.3390/biomedicines11030980
Peng H, Chen B, Wei W, Guo S, Han H, Yang C, Ma J, Wang L, Peng S, Kuang M, Lin S (2022) N6-methyladenosine (m6A) in 18S rRNA promotes fatty acid metabolism and oncogenic transformation. Nat Metab 4:1041–1054. https://doi.org/10.1038/s42255-022-00622-9
Pinyol R, Torrecilla S, Wang H, Montironi C, Piqué-Gili M, Torres-Martin M, Wei-Qiang L, Willoughby CE, Ramadori P, Andreu-Oller C, Taik P, Lee YA, Moeini A, Peix J, Faure-Dupuy S, Riedl T, Schuehle S, Oliveira CP, Alves VA et al (2021) Molecular characterisation of hepatocellular carcinoma in patients with non-alcoholic steatohepatitis. J Hepatol 75:865–878. https://doi.org/10.1016/j.jhep.2021.04.049
Pirola CJ, Sookoian S (2022) Epigenetics factors in nonalcoholic fatty liver disease. Expert Rev Gastroenterol Hepatol 16:521–536. https://doi.org/10.1080/17474124.2020.1765772
Pogribny IP, Dreval K, Kindrat I, Melnyk S, Jimenez L, De Conti A, Tryndyak V, Pogribna M, Ortega JF, James SJ, Rusyn I, Beland FA (2018) Epigenetically mediated inhibition of S-adenosylhomocysteine hydrolase and the associated dysregulation of 1-carbon metabolism in nonalcoholic steatohepatitis and hepatocellular carcinoma. FASEB J 32:1591–1601. https://doi.org/10.1096/fj.201700866R
Qin S, Liu G, ** H, Chen X, He J, **ao J, Qin Y, Mao Y, Zhao L (2022) The comprehensive expression and functional analysis of m6A modification “readers” in hepatocellular carcinoma. Aging 14:6269–6298. https://doi.org/10.18632/aging.204217
Quan J, Bode AM, Luo X (2021) ACSL family: the regulatory mechanisms and therapeutic implications in cancer. Eur J Pharmacol 909:174397. https://doi.org/10.1016/j.ejphar.2021.174397
Robinson MD, McCarthy DJ, Smyth GK (2009) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11:R25. https://doi.org/10.1186/gb-2010-11-3-r25
Ryaboshapkina M, Hammar M (2017) Human hepatic gene expression signature of non-alcoholic fatty liver disease progression, a meta-analysis. Sci Rep 7:1–12. https://doi.org/10.1038/s41598-017-10930-w
Sangro P, de la Torre AM, Sangro B, D’Avola D (2023) Metabolic dysfunction-associated fatty liver disease (MAFLD): an update of the recent advances in pharmacological treatment. J Physiol Biochem. https://doi.org/10.1007/s13105-023-00954-4
Santos A, Delgado TC, Marques V, Moncayo CR, Alonso C, Vidal-Puig A, Hall Z, Martínez-Chantar ML, Rodrigues CMP (2023) Spatial metabolomics and its application in the liver. Hepatology. https://doi.org/10.1097/hep.0000000000000341
Schofield Z, Reed MAC, Newsome PN, Adams DH, Günther UL, Lalor PF (2017) Changes in human hepatic metabolism in steatosis and cirrhosis. World J Gastroenterol 23:2685–2695. https://doi.org/10.3748/wjg.v23.i15.2685
Sid V, Siow YL, Karmin O (2017) Role of folate in nonalcoholic fatty liver disease. Can J Physiol Pharmacol 95:1141–1148. https://doi.org/10.1139/cjpp-2016-0681
Sinton MC, Hay DC, Drake AJ (2019) Metabolic control of gene transcription in non-alcoholic fatty liver disease: the role of the epigenome. Clin Epigenetics 11:104. https://doi.org/10.1186/s13148-019-0702-5
Stefanska B, Cheishvili D, Suderman M, Arakelian A, Huang J, Hallett M, Han ZG, Al-Mahtab M, Akbar SMF, Khan WA, Raqib R, Tanvir I, Khan HA, Rabbani SA, Szyf M (2014) Genome-wide study of hypomethylated and induced genes in patients with liver cancer unravels novel anticancer targets. Clin Cancer Res 20:3118–3132. https://doi.org/10.1158/1078-0432.CCR-13-0283
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Sun L, Kokura K, Izumi V, Koomen JM, Seto E, Chen J, Fang J (2015) MPP 8 and SIRT 1 crosstalk in E-cadherin gene silencing and epithelial–mesenchymal transition. EMBO Rep 16:689–699. https://doi.org/10.15252/embr.201439792
Sung WK, Zheng H, Li S, Chen R, Liu X, Li Y, Lee NP, Lee WH, Ariyaratne PN, Tennakoon C, Mulawadi FH, Wong KF, Liu AM, Poon RT, Fan ST, Chan KL, Gong Z, Hu Y, Lin Z et al (2012) Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat Genet 44:765–769. https://doi.org/10.1038/ng.2295
Teng YW, Mehedint MG, Garrow TA, Zeisel SH (2011) Deletion of betaine-homocysteine S-methyltransferase in mice perturbs choline and 1-carbon metabolism, resulting in fatty liver and hepatocellular carcinomas. J Biol Chem 286:36258–36267. https://doi.org/10.1074/jbc.M111.265348
Terracciani F, Falcomatà A, Gallo P, Picardi A, Vespasiani-Gentilucci U (2022) Prognostication in NAFLD: physiological bases, clinical indicators, and newer biomarkers. J Physiol Biochem. https://doi.org/10.1007/s13105-022-00934-0
Therneau TM, Grambsch PM (2000) Modeling survival data: extending the Cox model. Springer, New York, New York, NY
van der Maaten LJP, Hinton GE (2008) Visualizing high-dimensional data using t-SNE. J Mach Learn Res 9:2579–2605
Wang S, Gao S, Ye W, Li Y, Luan J, Lv X (2023) The emerging importance role of m6A modification in liver disease. Biomed Pharmacother 162:114669. https://doi.org/10.1016/j.biopha.2023.114669
Wang Z, Zhu S, Jia Y, Wang Y, Kubota N, Fujiwara N, Gordillo R, Lewis C, Zhu M, Sharma T, Li L, Zeng Q, Lin Y-H, Hsieh M-H, Gopal P, Wang T, Hoare M, Campbell P, Hoshida Y, Zhu H (2023) Positive selection of somatically mutated clones identifies adaptive pathways in metabolic liver disease. Cell 186:1968–1984.e20. https://doi.org/10.1016/j.cell.2023.03.014
Wiener D, Schwartz S (2021) The epitranscriptome beyond m6A. Nat Rev Genet 22:119–131. https://doi.org/10.1038/s41576-020-00295-8
Wiering L, Subramanian P, Hammerich L (2023) Hepatic stellate cells: dictating outcome in nonalcoholic fatty liver disease. Cell Mol Gastroenterol Hepatol 15:1277–1292. https://doi.org/10.1016/j.jcmgh.2023.02.010
**a P, Zhang H, Lu H, Xu K, Jiang X, Jiang Y, Gongye X, Chen Z, Liu J, Chen X, Ma W, Zhang Z, Yuan Y (2023) METTL5 stabilizes c-Myc by facilitating USP5 translation to reprogram glucose metabolism and promote hepatocellular carcinoma progression. Cancer Commun 43:338–364. https://doi.org/10.1002/cac2.12403
Xu F, Guo W (2020) The progress of epigenetics in the development and progression of non-alcoholic fatty liver disease. Liver Res 4:118–123. https://doi.org/10.1016/j.livres.2020.08.003
Xu H, Jiang C, Yao F, Liang H, Yan H, Chen D, Wu Y, Zhong L (2022) Pan-cancer analysis reveals the relation between TRMT112 and tumor microenvironment. J Oncol 2022. https://doi.org/10.1155/2022/1445932
Xu K, Ding J, Zhou L, Li D, Luo J, Wang W, Shang M, Lin B, Zhou L, Zheng S (2023) SMYD2 promotes hepatocellular carcinoma progression by reprogramming glutamine metabolism via c-Myc/GLS1 axis. Cells 12:1–15. https://doi.org/10.3390/cells12010025
Yang G, Yuan Y, Yuan H, Wang J, Yun H, Geng Y, Zhao M, Li L, Weng Y, Liu Z, Feng J, Bu Y, Liu L, Wang B, Zhang X (2021) Histone acetyltransferase 1 is a succinyltransferase for histones and non-histones and promotes tumorigenesis. EMBO Rep 22:1–18. https://doi.org/10.15252/embr.202050967
Ye Q, Liu Y, Zhang G, Deng H, Wang X, Tuo L, Chen C, Pan X, Wu K, Fan J, Pan Q, Wang K, Huang A, Tang N (2023) Deficiency of gluconeogenic enzyme PCK1 promotes metabolic-associated fatty liver disease through PI3K/AKT/PDGF axis activation in male mice. Nat Commun 14:1–19. https://doi.org/10.1038/s41467-023-37142-3
Younossi ZM (2019) Non-alcoholic fatty liver disease – a global public health perspective. J Hepatol 70:531–544. https://doi.org/10.1016/j.jhep.2018.10.033
Younossi ZM, Golabi P, Paik JM, Henry A, Van Dongen C, Henry L (2023) The global epidemiology of nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH): a systematic review. Hepatology 77:1335–1347. https://doi.org/10.1097/HEP.0000000000000004
Yu G, Wang LG, Han Y, He QY (2012) ClusterProfiler: an R package for comparing biological themes among gene clusters. Omi A J Integr Biol 16:284–287. https://doi.org/10.1089/omi.2011.0118
Zeybel M, Hardy T, Robinson SM, Fox C, Anstee QM, Ness T, Masson S, Mathers JC, French J, White S, Mann J (2015) Differential DNA methylation of genes involved in fibrosis progression in non-alcoholic fatty liver disease and alcoholic liver disease. Clin Epigenetics 7:1–11. https://doi.org/10.1186/s13148-015-0056-6
Zhang C, Li H, Wang Y, Liu W, Zhang Q, Zhang T, Zhang X, Han B, Zhou G (2010) Epigenetic inactivation of the tumor suppressor gene RIZ1 in hepatocellular carcinoma involves both DNA methylation and histone modifications. J Hepatol 53:889–895. https://doi.org/10.1016/j.jhep.2010.05.012
Zhang N, Tian X, Yan T, Wang H, Zhang D, Lin C, Liu Q, Jiang S (2023) Insights into the role of nucleotide methylation in metabolic-associated fatty liver disease. Front Immunol 14:1148722. https://doi.org/10.3389/fimmu.2023.1148722
Zhang X, Asllanaj E, Amiri M, Portilla-Fernandez E, Bramer WM, Nano J, Voortman T, Pan Q, Ghanbari M (2021) Deciphering the role of epigenetic modifications in fatty liver disease: a systematic review. Eur J Clin Invest 51:1–16. https://doi.org/10.1111/eci.13479
Zhao Z, Meng J, Su R, Zhang J, Chen J, Ma X, **a Q (2020) Epitranscriptomics in liver disease: basic concepts and therapeutic potential. J Hepatol 73:664–679. https://doi.org/10.1016/j.jhep.2020.04.009
Zhu Y, Zhang H, Jiang P, ** to diagnose non-alcoholic steatohepatitis. Biomedicines 11. https://doi.org/10.3390/biomedicines11030970
Acknowledgements
The generous support of Mr Eduardo Avila and the kind sponsorship of Ms Olga Alzueta are acknowledged.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was supported by grants from the Scientific Foundation of the Spanish Association Against Cancer (AECC) LABAE20011GARC (MGFB). Grants from Ministerio de Ciencia Innovación y Universidades MICINN-Agencia Estatal de Investigación integrado en el Plan Estatal de Investigación Científica y Técnica y Innovación, cofinanciado con Fondos FEDER PID2019-104878RB-100/AEI/10.13039/501100011033 (MAA), PID2020-117116RB-I00 (MGFB) and Proyectos en colaboración público-privada 2021 CPP2021-008411 (MGFB and MAA). FIMA AC pre-doctoral fellowships (ACC and EAV), AECC investigador fellowship INVES223049AREC (MA), Sara Borrell Contract CD22/00109 From Spanish Ministry of Health (ALP), and Ramón y Cajal Program contract RYC2018-024475-1 (MGFB) from the Spanish Ministry of Science and Innovation. Grants from Fundación Echébano, Fundación Mario Losantos, and Fundación M Torres (MAA, MA, CB, MGFB).
Author information
Authors and Affiliations
Contributions
Data search, analysis, and integration: JMH, ALP, ACC, MUL, MA, IU, BCU, EAV, BS, AI, CB; concept and design: JMH, ALP, MGFB, MAA; supervision: MGFB, MAA; writing of article: JMH, ALP, MGFB, MAA. The authors declare that all data were generated in-house and that no paper mill was used.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Points
-Epigenetic and epitranscriptomic genes’ expression is markedly altered in NAFLD.
-Epigenetic and epitranscriptomic signatures were associated with advanced NAFLD.
-Enzymes acting on epigenetic and epitranscriptomic cofactors are deranged in NAFLD.
-New epigenetic and epitranscriptomic effectors may constitute drug targets in NAFLD.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Herranz, J.M., López-Pascual, A., Clavería-Cabello, A. et al. Comprehensive analysis of epigenetic and epitranscriptomic genes’ expression in human NAFLD. J Physiol Biochem 79, 901–924 (2023). https://doi.org/10.1007/s13105-023-00976-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13105-023-00976-y