Abstract
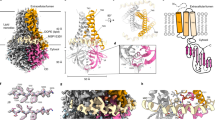
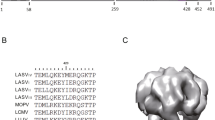
Lassa virus (LASV) is the most prevalent member of the arenavirus family and the causative agent of Lassa fever, a viral hemorrhagic fever. Although there are annual outbreaks in West Africa, and recently isolated cases worldwide, there are no current therapeutics or vaccines. As such, LASV poses a significant global public health threat. One of the key steps in LASV infection is delivering its genetic material by fusing its viral membrane with the host cell membrane. This process is facilitated by significant conformational changes within glycoprotein 2 (GP2), yielding distinct prefusion and postfusion structural states. However, structural information is missing to understand the changes that occur in the transmembrane domain (TM) during the fusion process. Previously, we showed that the TM undergoes pH-dependent structural changes that result in a helical extension. Here, we provide the 1H, 15N, and 13C assignment of the LASV TM backbone in the prefusion and postfusion states. We also provide the 1H, 15N, and 13C assignment of two mutants, G429P and D432P, which prevent this helical extension. These results will help understand the role the TM plays in membrane fusion and can lead to the design of therapeutics against LASV infection.


Similar content being viewed by others
Data availability
The chemical shift information for the wild type LASV TM and mutants was deposited in the BMRB data bank under the following entry numbers: Prefusion wild type TM – 52449; Postfusion wild type TM – 52450; Prefusion TM G429P – 52451; Postfusion TM G429P – 52452; Prefusion TM D432P – 52453; Postfusion TM D432P – 52454.
References
Agbonlahor DE, Akpede GO, Happi CT, Tomori O (2021) 52 years of Lassa Fever outbreaks in Nigeria, 1969–2020: an epidemiologic analysis of the temporal and spatial trends. Am J Trop Med Hyg 105:974–985. https://doi.org/10.4269/ajtmh.20-1160
Apellániz B, Edurne R, Pablo C et al (2014) Cholesterol-dependent membrane Fusion Induced by the gp41 membrane-proximal External region–transmembrane domain connection suggests a mechanism for broad HIV-1 neutralization. J Virol 88:13367–13377. https://doi.org/10.1128/jvi.02151-14
Bax A, Ikura M (1991) An efficient 3D NMR technique for correlating the proton and 15 N backbone amide resonances with the α-carbon of the preceding residue in uniformly 15 N/13 C enriched proteins. J Biomol Nmr 1:99–104. https://doi.org/10.1007/bf01874573
Chang D-K, Cheng S-F, Kantchev EAB et al (2008) Membrane interaction and structure of the transmembrane domain of influenza hemagglutinin and its fusion peptide complex. Bmc Biol 6:2. https://doi.org/10.1186/1741-7007-6-2
Clubb RT, Thanabal V, Wagner G (1992) A constant-time three-dimensional triple-resonance pulse scheme to correlate intraresidue 1HN, 15 N, and 13 C′ chemical shifts in 15 N 13 C-labelled proteins. J Magn Reson 1969 97:213–217. https://doi.org/10.1016/0022-2364(92)90252-3
Delaglio F, Grzesiek S, Vuister GW et al (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol Nmr 6:277–293. https://doi.org/10.1007/bf00197809
Grzesiek S, Bax A (1992) An efficient experiment for sequential backbone assignment of medium-sized isotopically enriched proteins. J Magnetic Reson 1969 99:201–207. https://doi.org/10.1016/0022-2364(92)90169-8
Hastie KM, Zandonatti MA, Kleinfelter LM et al (2017) Structural basis for antibody-mediated neutralization of Lassa virus. Science 356:923–928. https://doi.org/10.1126/science.aam7260
Holmes GP, McCormick JB, Trock SC et al (1990) Lassa Fever in the United States. New Engl J Med 323:1120–1123. https://doi.org/10.1056/nejm199010183231607
Hu B, Tina HC, Christoph T, Michael V (2019) Cholesterol binding to the Transmembrane Region of a Group 2 Hemagglutinin (HA) of Influenza Virus is essential for Virus Replication, affecting both Virus Assembly and HA Fusion Activity. J Virol 93. https://doi.org/10.1128/jvi.00555-19
Katz M, Weinstein J, Eilon-Ashkenazy M et al (2022) Structure and receptor recognition by the Lassa virus spike complex. Nature 1–6. https://doi.org/10.1038/s41586-022-04429-2
Kay LE, Ikura M, Tschudin R, Bax A (1990) 3-Dimensional triple-resonance NMR-spectroscopy of isotopically enriched proteins. J Magnetic Reson 1969 89:496–514. https://doi.org/10.1016/0022-2364(90)90333-5
Keating PM, Pennington HN, Collins SD, Lee J (2023a) Purification and characterization of the Lassa virus transmembrane domain. Biochem Biophys Rep 33:101409. https://doi.org/10.1016/j.bbrep.2022.101409
Keating PM, Schifano NP, Wei X et al (2023b) pH-dependent conformational change within the Lassa virus transmembrane domain elicits efficient membrane fusion. Biochim Biophys Acta (BBA) - Biomembr 184233. https://doi.org/10.1016/j.bbamem.2023.184233
Lecompte E, Fichet-Calvet E, Daffis S et al (2006) Mastomys natalensis and Lassa Fever, West Africa. Emerg Infect Dis 12:1971–1974. https://doi.org/10.3201/eid1212.060812
Lee J, Nyenhuis DA, Nelson EA et al (2017) Structure of the Ebola virus envelope protein MPER/TM domain and its interaction with the fusion loop explains their fusion activity. Proc Natl Acad Sci 114:E7987–E7996. https://doi.org/10.1073/pnas.1708052114
Lee J, Kreutzberger AJB, Odongo L et al (2021) Ebola virus glycoprotein interacts with cholesterol to enhance membrane fusion and cell entry. Nat Struct Mol Biol 28:181–189. https://doi.org/10.1038/s41594-020-00548-4
Maciejewski MW, Schuyler AD, Gryk MR et al (2017) NMRbox: a resource for Biomolecular NMR computation. Biophys J 112:1529–1534. https://doi.org/10.1016/j.bpj.2017.03.011
McCormick JB, Webb PA, Krebs JW et al (1987) A prospective study of the Epidemiology and Ecology of Lassa Fever. J Infect Dis 155:437–444. https://doi.org/10.1093/infdis/155.3.437
Mehand MS, Al-Shorbaji F, Millett P, Murgue B (2018) The WHO R&D Blueprint: 2018 review of emerging infectious diseases requiring urgent research and development efforts. Antivir Res 159:63–67. https://doi.org/10.1016/j.antiviral.2018.09.009
Nunberg JH, York J (2012) The curious case of arenavirus entry, and its inhibition. Viruses 4:83–101. https://doi.org/10.3390/v4010083
Ogbu O, Ajuluchukwu E, Uneke CJ (2007) Lassa fever in west African sub-region: an overview. J Vector Dis 44:1–11
Pennington HN, Lee J (2022) Lassa virus glycoprotein complex review: insights into its unique fusion machinery. Bioscience Rep. https://doi.org/10.1042/bsr20211930
Pinto D, Fenwick C, Caillat C et al (2019) Structural basis for broad HIV-1 neutralization by the MPER-Specific Human broadly neutralizing antibody LN01. Cell Host Microbe 26:623–637e8. https://doi.org/10.1016/j.chom.2019.09.016
Richmond JK, Baglole DJ (2003) Lassa fever: epidemiology, clinical features, and social consequences. BMJ 327:1271. https://doi.org/10.1136/bmj.327.7426.1271
Salu OB, Amoo OS, Shaibu JO et al (2020) Monitoring of Lassa virus (LASV) infection in suspected and confirmed cases in Ondo State, Nigeria. Pan Afr Med J 36:253. https://doi.org/10.11604/pamj.2020.36.253.22104
Shulman A, Katz M, Cohen-Dvashi H et al (2019) Variations in Core Packing of GP2 from Old World mammarenaviruses in their Post-fusion conformations affect membrane-Fusion efficiencies. J Mol Biol 431:2095–2111. https://doi.org/10.1016/j.jmb.2019.04.012
Woonghee L, Marco T, John LM (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
World Health Organization (2022) Lassa fever – United Kingdom of Great Britain and Northern Ireland. https://www.who.int/emergencies/disease-outbreak-news/item/lassa-fever-united-kingdom-of-great-britain-and-northern-ireland. Accessed 11 May 2022
Yaro CA, Kogi E, Opara KN et al (2021) Infection pattern, case fatality rate and spread of Lassa virus in Nigeria. Bmc Infect Dis 21:149. https://doi.org/10.1186/s12879-021-05837-x
Acknowledgements
This work was supported by the University of Maryland startup funds. We would like to thank D. Zhang for assistance with the NMR experiments.
Author information
Authors and Affiliations
Contributions
PMK and JL designed the experiments; PMK prepared the samples and performed all the NMR analyses; PMK and JL wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Keating, P.M., Lee, J. Assignment of the Lassa virus transmembrane domain in the prefusion and postfusion states in detergent micelles. Biomol NMR Assign (2024). https://doi.org/10.1007/s12104-024-10184-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12104-024-10184-4