Abstract
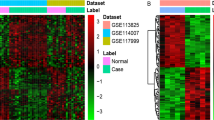
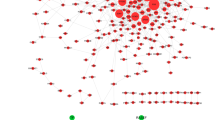
Osteoarthritis (OA) is a common degenerative joint disorder that adversely affects the quality of life of patients. Identification of novel diagnostic biomarkers is pivotal for the early detection and prevention of OA. Dataset GSE185059 was selected from Gene Expression Omnibus database to obtain differentially expressed lncRNAs (DE-lncRNAs), mRNAs (DE-mRNAs), and circRNAs (DE-circRNAs) between OA and normal samples. The Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses as well as protein–protein interaction (PPI) network construction of DE-mRNAs were conducted. Hub genes were identified from PPI networks and validated by RT-qPCR. starBase database was utilized for predicting miRNAs binding with hub genes, selected DE-lncRNAs and DE-circRNAs, respectively. The competing endogenous RNA (ceRNA) networks were constructed. A total of 818 DE-mRNAs, 191 DE-lncRNAs, and 2053 DE-circRNAs were identified. The DE-mRNAs were significantly enriched in several inflammation-related GO terms and KEGG pathways such as positive regulation of cell–cell adhesion, TNF-alpha signaling pathway and NF-kappa B signaling pathway. Thirteen hub genes were identified, which were CFTR, GART, SMAD2, NCK1, TJP1, UBE2D1, EFTUD2, PRKACB, IL10, SNRPG, CHD4, RPS24, and SRSF6. OA-related DE-lncRNA/circRNA-miRNA-hub gene networks were constructed. We identified 13 hub genes and constructed the ceRNA networks related to OA, providing a theoretical basis for further research.








Similar content being viewed by others
References
He, K., Huang, X., Shan, R., Yang, X., Song, R., **e, F., & Huang, G. (2021). Intra-articular injection of Lornoxicam and MicroRNA-140 co-loaded cationic liposomes enhanced the therapeutic treatment of experimental osteoarthritis. An Official Journal of the American Association of Pharmaceutical Scientists, 23(1), 9.
Tramś, E., Malesa, K., Pomianowski, S., & Kamiński, R. (2022). Role of platelets in osteoarthritis-updated systematic review and meta-analysis on the role of platelet-rich plasma in osteoarthritis. Cells, 11(7), 1080.
Wang, Y., Hussain, S. M., Gan, D., Lim, Y. Z., Estee, M. M., Heritier, S., Wluka, A. E., & Cicuttini, F. M. (2021). Topical corticosteroid for treatment of hand osteoarthritis: Study protocol for a randomised controlled trial. BMC Musculoskeletal Disorders, 22(1), 1036.
Franco-Trepat, E., Alonso-Pérez, A., Guillán-Fresco, M., Jorge-Mora, A., Crespo-Golmar, A., López-Fagúndez, M., Pazos-Pérez, A., Gualillo, O., Belén Bravo, S., & Gómez Bahamonde, R. (2022). Amitriptyline blocks innate immune responses mediated by toll-like receptor 4 and IL-1 receptor: Preclinical and clinical evidence in osteoarthritis and gout. British Journal of Pharmacology, 179(2), 270–286.
Kreitmaier, P., Suderman, M., Southam, L., Coutinho de Almeida, R., Hatzikotoulas, K., Meulenbelt, I., Steinberg, J., Relton, C. L., Wilkinson, J. M., & Zeggini, E. (2022). An epigenome-wide view of osteoarthritis in primary tissues. The American Journal of Human Genetics, 109(7), 1255–1271.
Qu, Z., Yang, F., Yan, Y., Huang, J., Zhao, J., Hong, J., Li, S., Jiang, G., Wang, W., & Yan, S. (2021). A Mendelian randomization study on the role of serum parathyroid hormone and 25-hydroxyvitamin D in osteoarthritis. Osteoarthritis Cartilage, 29(9), 1282–1290.
Udhaya, K. S., Rajan, B., Thirumal, K. D., Anu, P. V., Abunada, T., Younes, S., Okashah, S., Ethiraj, S., George Priya Doss, C., & Zayed, H. (2020). Involvement of essential signaling cascades and analysis of gene networks in diabesity. Genes (Basel), 11(11), 1256.
Zhou, J., Zou, D., Wan, R., Liu, J., Zhou, Q., Zhou, Z., Wang, W., Tao, C., & Liu, T. (2022). Gene expression microarray data identify hub genes involved in osteoarthritis. Frontiers in Genetics, 13, 870590.
Wang, W., Chen, Z., & Hua, Y. (2023). Bioinformatics prediction and experimental validation identify a novel cuproptosis-related gene signature in human synovial inflammation during osteoarthritis progression. Biomolecules, 13(1), 127.
Wu, Z. Y., Du, G., & Lin, Y. C. (2021). Identifying hub genes and immune infiltration of osteoarthritis using comprehensive bioinformatics analysis. Journal of Orthopaedic Surgery and Research, 16(1), 630.
Feng, Z., & Lian, K. J. (2015). Identification of genes and pathways associated with osteoarthritis by bioinformatics analyses. European Review for Medical and Pharmacological Sciences, 19(5), 736–744.
Wu, Y., Lu, X., Shen, B., & Zeng, Y. (2019). The therapeutic potential and role of miRNA, lncRNA, and circRNA in osteoarthritis. Current Gene Therapy, 19(4), 255–263.
Salmena, L., Poliseno, L., Tay, Y., Kats, L., & Pandolfi, P. P. (2011). A ceRNA hypothesis: The Rosetta stone of a hidden RNA language? Cell, 146(3), 353–358.
Wu, T., & Du, Y. (2017). LncRNAs: From basic research to medical application. International Journal of Biological Sciences, 13(3), 295–307.
Wang, X. J., Gao, J., Yu, Q., Zhang, M., & Hu, W. D. (2022). Multi-Omics integration-based prioritisation of competing endogenous RNA regulation networks in small cell lung cancer: Molecular characteristics and drug candidates. Frontiers in Oncology, 12, 904865.
Barrett, T., Wilhite, S. E., Ledoux, P., Evangelista, C., Kim, I. F., Tomashevsky, M., Marshall, K. A., Phillippy, K. H., Sherman, P. M., Holko, M., Yefanov, A., Lee, H., Zhang, N., Robertson, C. L., Serova, N., Davis, S., & Soboleva, A. (2013). NCBI GEO: Archive for functional genomics data sets–update. Nucleic Acids Research, 41(D1), D991–D995.
Sherman, B. T., Hao, M., Qiu, J., Jiao, X., Baseler, M. W., Lane, H. C., Imamichi, T., & Chang, W. (2022). DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Research. https://doi.org/10.1093/nar/gkac194
da Huang, W., Sherman, B. T., & Lempicki, R. A. (2009). Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols, 4(1), 44–57.
Szklarczyk, D., Gable, A. L., Lyon, D., Junge, A., Wyder, S., Huerta-Cepas, J., Simonovic, M., Doncheva, N. T., Morris, J. H., Bork, P., Jensen, L. J., & Mering, C. V. (2019). STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Research, 47(D1), D607-d613.
Li, J. H., Liu, S., Zhou, H., Qu, L. H., & Yang, J. H. (2014). starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Research, 42(D1), D92–D97.
Bijlsma, J. W., Berenbaum, F., & Lafeber, F. P. (2011). Osteoarthritis: An update with relevance for clinical practice. Lancet, 377(9783), 2115–2126.
You, R., Liu, S., & Tan, J. (2022). Screening and identification of osteoarthritis related differential genes and construction of a risk prognosis model based on bioinformatics analysis. Annals of Translational Medicine, 10(8), 444.
de Rezende, M. U., & de Campos, G. C. (2013). Is osteoarthritis a mechanical or inflammatory disease? Revista Brasileira de Ortopedia, 48(6), 471–474.
Goldring, M. B., & Otero, M. (2011). Inflammation in osteoarthritis. Current Opinion in Rheumatology, 23(5), 471–478.
Ragni, E., De Luca, P., Valli, F., Zagra, L., & de Girolamo, L. (2023). Inflammatory treatment used to mimic osteoarthritis and patients’ synovial fluid have divergent molecular impact on chondrocytes in vitro. International Journal of Molecular Sciences, 24(3), 2625.
Sanchez-Lopez, E., Coras, R., Torres, A., Lane, N. E., & Guma, M. (2022). Synovial inflammation in osteoarthritis progression. Nature Reviews Rheumatology, 18(5), 258–275.
Choi, M. C., Jo, J., Park, J., Kang, H. K., & Park, Y. (2019). NF-κB signaling pathways in osteoarthritic cartilage destruction. Cells, 8(7), 734.
Yang, J., & Jiang, W. (2020). The role of SMAD2/3 in human embryonic stem cells. Front Cell Dev Biol, 8, 653.
He, Y., Fan, L., Aaron, N., Feng, Y., Fang, Q., Zhang, Y., Zhang, D., Wang, H., Ma, T., Sun, J., & Chen, J. (2021). Reduction of Smad2 caused by oxidative stress leads to necrotic death of hypertrophic chondrocytes associated with an endemic osteoarthritis. Rheumatology (Oxford), 61(1), 440–451.
Ge, Q., Shi, Z., Zou, K. A., Ying, J., Chen, J., Yuan, W., Wang, W., **ao, L., Lin, X., Chen, D., Feng, X. H., Wang, P. E., Tong, P., & **, H. (2023). Protein phosphatase PPM1A inhibition attenuates osteoarthritis via regulating TGF-β/Smad2 signaling in chondrocytes. JCI Insight. https://doi.org/10.1172/jci.insight.166688
Li, G., **u, L., Li, X., Ma, L., & Zhou, J. (2022). miR-155 inhibits chondrocyte pyroptosis in knee osteoarthritis by targeting SMAD2 and inhibiting the NLRP3/Caspase-1 pathway. Journal of Orthopaedic Surgery and Research, 17(1), 48.
Shen, P., Yang, Y., Liu, G., Chen, W., Chen, J., Wang, Q., Gao, H., Fan, S., Shen, S., & Zhao, X. (2020). CircCDK14 protects against Osteoarthritis by sponging miR-125a-5p and promoting the expression of Smad2. Theranostics, 10(20), 9113–9131.
Li, J., Liu, M., Li, X., Shi, H., & Sun, S. (2021). Long noncoding RNA ZFAS1 suppresses chondrocytes apoptosis via miR-302d-3p/SMAD2 in osteoarthritis. Bioscience, Biotechnology, and Biochemistry, 85(4), 842–850.
Wang, X., Wang, L. T., & Yu, B. (2022). UBE2D1 and COX7C as potential biomarkers of diabetes-related sepsis. BioMed Research International, 2022, 9463717.
Mao, D., Wu, M., Wei, J., Zhou, X., Yang, L., & Chen, F. (2021). MicroRNA-101a-3p could be involved in the pathogenesis of temporomandibular joint osteoarthritis by mediating UBE2D1 and FZD4. Journal of Oral Pathology and Medicine, 50(2), 236–243.
Wang, Y., Guo, T., Liu, Q., & **e, X. (2020). CircRAD18 accelerates the progression of acute myeloid Leukemia by modulation of miR-206/PRKACB axis. Cancer Management and Research, 12, 10887–10896.
Ham, O., Lee, C. Y., Song, B. W., Lee, S. Y., Kim, R., Park, J. H., Lee, J., Seo, H. H., Lee, C. Y., Chung, Y. A., Maeng, L. S., Lee, M. Y., Kim, J., Hwang, J., Woo, D. K., & Chang, W. (2014). Upregulation of miR-23b enhances the autologous therapeutic potential for degenerative arthritis by targeting PRKACB in synovial fluid-derived mesenchymal stem cells from patients. Molecules and Cells, 37(6), 449–456.
Zhao, C. (2021). Identifying the hub gene and immune infiltration of osteoarthritis by bioinformatical methods. Clinical Rheumatology, 40(3), 1027–1037.
Yu, L. K., Zhang, J., Sun, Z. Y., Ruan, C. L., Li, H., & Ruan, X. J. (2021). Coculture with interleukin-10 overexpressed chondrocytes: A cell therapy model to ameliorate the post-traumatic osteoarthritis development. Journal of Biological Regulators and Homeostatic Agents, 35(2), 593–603.
Li, H., Gao, L., Kang, X., Wang, X., Yu, Y., Zhang, Y., & Chen, H. (2023). RPS24 is associated with a poor prognosis and immune infiltration in hepatocellular carcinoma. International Journal of Molecular Sciences, 24(1), 806.
Mi, B., Liu, G., Zhou, W., Lv, H., Liu, Y., & Liu, J. (2018). Identification of genes and pathways in the synovia of women with osteoarthritis by bioinformatics analysis. Molecular Medicine Reports, 17(3), 4467–4473.
Kong, H., Sun, M. L., Zhang, X. A., & Wang, X. Q. (2021). Crosstalk among circRNA/lncRNA, miRNA, and mRNA in osteoarthritis. Frontiers in Cell and Development Biology, 9, 774370.
Zhang, P., Sun, J., Liang, C., Gu, B., Xu, Y., Lu, H., Cao, B., & Xu, H. (2020). lncRNA IGHCγ1 acts as a ceRNA to regulate macrophage inflammation via the miR-6891-3p/TLR4 axis in osteoarthritis. Mediators of Inflammation, 2020, 9743037.
Liu, C., Cheng, P., Liang, J., Zhao, X., & Du, W. (2021). Circular RNA circ_0128846 promotes the progression of osteoarthritis by regulating miR-127-5p/NAMPT axis. Journal of Orthopaedic Surgery and Research, 16(1), 307.
Acknowledgements
Not applicable.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare.
Ethical Approval
This study was approved by the medical Ethics Committee of Changzhi People’s Hospital (approval number: IEC202302003). Written informed consent was obtained from each participant.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, W., Wei, C. & Wang, L. Identification of Key lncRNAs, circRNAs, and mRNAs in Osteoarthritis via Bioinformatics Analysis. Mol Biotechnol 66, 1660–1672 (2024). https://doi.org/10.1007/s12033-023-00790-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-023-00790-3