Abstract
Human T-cell leukaemia virus type 1 (HTLV-1) is the causative agent of two life-threatening diseases, adult T cell leukaemia/lymphoma (ATLL), and HTLV-1-associated myelopathy/tropical spastic (HAM/TSP). HTLV-1 protease (HTLV-1-PR) is an aspartic protease that represents a promising target for therapeutic purposes like human immunodeficiency virus-PR inhibitors (HIV-PR). Therefore, in this study, the human Fc fusion recombinant-PR (HTLV-1-PR:hFcγ1) was designed and expressed for two applications, finding a blocking substrate as a potential therapeutic or a potential subunit peptide vaccine. The PCR amplified DNA sequences encoding the HTLV-1-PR from the MT2-cell line using specific primers with restriction enzyme sites of Not1 and Xba1. The construct was then cloned to pTZ57R/T TA plasmid and, after confirming the PR sequence, subcloned into the pDR2ΔEF1α Fc-expression vector to create pDR2ΔEF1α.HTLV-1-PR:hFcγ1. The integrity of recombinant DNA was confirmed by sequencing to ensure that the engineered construct was in the frame. The recombinant fusion protein was then produced in the Chinese hamster ovary cell (CHO) system and was purified from its supernatant using HiTrap-rPA column affinity chromatography. Then, the immunofluorescence assay (IFA) co-localisation method showed that HTLV-1-PR:hFc recombinant fusion protein has appropriate folding as it binds to the anti-Fcγ antibody; the Fcγ1 tag participates to have HTLV-1-PR:hFcγ1 as a dimeric secretory protein. The development and production of HTLV-1-PR can be used to find a blocking substrate as a potential therapeutic molecule and apply it in an animal model to assess its immunogenicity and potential protection against HTLV-1 infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human T-cell lymphotropic virus type 1 (HTLV-1), as first known oncovirus in the Retroviridae family, is clinically associated with two life-threatening diseases: adult T cell leukaemia/lymphoma (ATLL) and HTLV-1-associated myelopathy/tropical spastic (HAM/TSP) [1]. Although HTLV-1 infection affects 10–20 million people with slowly grows globally, an effective vaccine or therapeutic agent is yet to be introduced [2]. Therefore, it was brought to the attention of the World Health Organization (WHO) to introduce the elimination planning for HTLV-1. The only promising strategy to reach such a goal is discovering medications and anti-retroviral therapies that optimistically change the perspectives of HTLV-1 infection and its associated diseases [3,4,5].
HTLV-1, like human immunodeficiency virus (HIV) in the Retroviridae family, shares three critical enzymes which are pivotal in its life cell cycle, reverse transcriptase (RT), integrase (IN), and protease (PR) [6]. These essential enzymes, which are exclusively expressed by the HTLV-1, particularly PR, are attractive candidates for anti-viral drug development, as has been done for HIV infection, and mainly improve the survival time and quality of life in those patients [2, 7]. Therefore, HTLV-1-PR is also an impressive target for making effective therapeutic medications for HTLV-1-associated diseases. Moreover, it can be used as a subunit vaccine to study HTLV-1-infected animal models. Of note, HTLV-1 and HIV PRs share only 28% sequence identity. However, their binding sites are more conserved (45%) [8].
HTLV-1-PR is an aspartic acid protease, a 28-kDa homodimer protein with each chain consisting of 125 amino acid residues. It is particular to cleave the primary protein into the structural proteins and enzymes necessary for producing virus particles [9,10,11]. Consequently, deactivating this vital enzyme is more likely to prevent HTLV-1 replication and dissemination strongly. It is necessary to study the structure and physiology of the protein by making it a recombinant protein to reach such a goal.
In 1988 the same program was designed to inhibit HIV replication by blocking the viral PR [12, 13]. The subsequent attempts resulted in manufacturing effective anti-HIV protease, a combination therapy regimen that improved the survival time and life quality of HIV-infected patients [14]. Despite sharing properties with HIV-PR, using several commercial HIV-1-PR inhibitors, such as ritonavir, amprenavir, saquinavir, indinavir, and nelfinavir, for HTLV-1-associated diseases treatment did not come the good results in HTLV-1-associated disease infections. It seems that in the active site of HTLV-1-PR, Met37 residue differs from Asp30 in HIV protease [15] and contains extra-amino acid in the C terminus [15,16,17].
Recombinant protein productions are widely used as critical components to characterise structural, functional, and distributional features [18]. The Fc moiety design is aimed at improving immunogenicity by increasing Fc-receptor-type I (FcγR-I, CD64) uptake on Ag-presenting cells (APCs) and inducing cross-presentation for potentiation of Th1 anti-viral responses [19]. Many theranostic applications of Fc-fusion proteins are illustrious with details [20, 21]. Production of recombinant PR bound to Fc has many advantages such as increased capacity, increased half-life, easy secretion of protein, stability, and particularly, inducing T cell-mediated immunity [21] as a potential fusion peptide vaccine candidate. Due to the increasing prevalence of HTLV-1 as a neglected virus that grows worldwide [22] and the lack of effective treatment or vaccine, designing a specific drug or vaccine is a priority for global health.
Taken together, an effective and powerful medication against HTLV-1-PR is yet to be introduced. In the present study, the profit gained from Fcγ1 fusion protein, the HTLV-1-PR:hFcγ1 recombinant fusion molecule, was designed and produced to find proper inhibitors in vitro and in vivo for the treatment of HTLV-1 infection and HTLV-1-associated diseases. Moreover, due to the unique property of hFcγ1 in inducing cross-presentation and Th1 responses, it may be a potential tool for planning and develo** a protective vaccine against HTLV-1 infection.
Material and Methods
In-Silico Molecular Modelling of HTLV-1-PR:hFcγ1
Three-dimensional models of the HTLV-1-PR:hFcγ1 sequences were constructed via homology modelling. Basic Local Alignment Search Tool (BLAST) homology sequence searches were performed to identify the template proteins by position-specific iterative (PSI-BLAST) (http://www.ncbi.nlm.nih.gov/BLAST/) [23]. For modelling, the HTLV-1-PR (Protein Data Bank [PDB] entry: 2B7F), HTLV-1 capsid protein (PDB: 1QRJ), and human IgG1 Fc-fragment (PDB entry: 4CDH) were selected as the templates of HTLV-1-PR and Fc domains, respectively. Multiple alignments were carried out on the selected sequences by ClustalX2 (protein weight matrix: BLOSUM series). Model building was performed by the MODELLER9v20 software using a model-ligand algorithm (http://salilab.org/modeller/) [24]. Finally, structure refinements and energy minimisation were done by the YASARA energy minimisation server and Chimera 1.14 [25]. The protein homodimer form was created by HOMCOS (http://homcos.pdbj.org) in the base of a human IgG1 heavy chain (1HZH). All models were validated using the ERRAT and PROCHECK software at the University of California, Los Angeles (UCLA, https://saves.mbi.ucla.edu/).
Construction of HTLV-1-PR:hFcγ1 Recombinant Fusion Molecule
The DNA fragment was isolated from the complete genome of the MT2-cell line. MT2-cell line was obtained from the National Cell Bank of Iran (Pasteur Institute of Iran, Tehran). It is amplified using proofreading pfu polymerase (Thermo Fisher Scientific, USA), with the relevant primers as follows: forward, 5′-ATGACAGTCCTT CCGATAGC-3′ and reverse, 5′-TTAGAGAGTTAGTGGCCCG-3′ to produce the HTLV-1-PR recombinant molecule (HTLV-1-PR:hFcγ1). The gradient PCR was performed to amplify protease with temperatures ranging from 44 to 56 °C.
Appropriate primer pairs assist with restriction enzyme digestion (GC^GGCCGC) Not1 and (T^CTAGA) Xba1 to enable ligation into the expression vector, the engineered-Fc eukaryotic high-expression vector, pDR2EF1α:hFcγ1 (kindly provided by Prof. David Blackbourn, University of Aberdeen, UK). This expression vector was initially generated to put desired DNA fragment in frame with hinge and CH2 and CH3 domains of human FcIgG1 regions. Therefore, the Chinese hamster ovary (CHO) Fc fusion protein expression system was chosen to express the construct of HTLV-1-PR as N-terminal fusion with the C-terminal Fc of the human IgG1 (γ1) chain for characterisation and functional studies.
Transformation and Cloning Setting
The PCR products harbouring the PR gene were assessed to streamline the cloning process and confirm the absence of unwanted base changes by sequencing analysis. First, TA was cloned into the pTZ57R/T vector (Thermo Fisher Scientific, USA), then was transformed by chemical method (CaCl2) into Escherichia coli Top10F′ cells and were grown onto Luria–Bertani (LB) agar medium, supplemented with 100 μg/mL ampicillin. The plates were then incubated at 37 °C overnight, and the white colonies were screened out based on recombinant (White) and plasmid-specific (Blue) screening.
The plasmid was extracted from the transformant white colonies using a PrimePrep Plasmid DNA Isolation Kit (Genet Bio, Korea) and then wholly sequenced in a bi-direction (Milligen, France). The integrity of recombinant DNA was confirmed by sequencing to ensure that the engineered construct was in-frame. Recombinant plasmids were cleaned up for transformation with integrative vectors (HiPure DNA Clean-Up, Roche, Germany) before transformation according to the manufacturer’s instructions.
Subcloning to pDR2EF1α:hFcγ1 Expression Vector
The product was digested with XbaI and NotI (Thermo Fisher Scientific, USA) to clone in the frame to SpeI- and NotI-digested pDR2ΔEF1α.Fc expression vector to create the appropriate recombinant plasmids (pDR2ΔEF1α:HTLV-1-PR:hFcγ1). Cloning accuracy was performed by enzymatic digestion processes and confirmed via the colony PCR method.
Transfection of HTLV-1-PR:hFcγ1 Recombinant Plasmid Into CHO Cells
The CHO cell expression system was chosen to express the HTLV-1-PR:hFcγ1 construct. The CHO cell line as primary mammalian expression host cells was obtained from the National Cell Bank of Iran (Pasteur Institute of Iran, Tehran, Iran).
The CHO cells were grown in flasks (Thermo Fisher Scientific, USA) at 37 °C, 5% CO2, and 50 rpm until 90% confluent in RPMI 1640 medium (Biosera, France) supplemented with 10% foetal bovine serum (FBS; Gibco, USA) and containing hygromycin B (InvivoGen, USA). According to the manufacturer’s instructions, the cells were transfected using a Lipofectamine 2000 kit Plus LTX (Invitrogen, USA).
The minimal concentration of hygromycin B required to kill untransfected cells was determined by performing a kill curve (dose–response) in which increasing levels of the anti-biotic (0, 100, 200, 400, 800, and 1000 μg/mL) were added to 12-well plates seeded with 10,000 cells per well, the previous day. Cells were passed three times over 12 days in the selective medium. This experiment was carried out twice to establish that hygromycin B at 400 μg/mL concentration killed the untransfected cells.
Optimal transfection efficiency for Lipofectamine 2000 was achieved using a 400 ng per well of engineered pDR2ΔEF1α HTLV-1-PR:hFcγ1 construct and 10 μL of Lipofectamine 2000 reagent in 2 mL of growth medium containing 6.0 × 105 cells at 90% confluency.
After transfection, cells (72 h post-addition of DNA) were cultivated in a selective medium containing 10 μg/mL hygromycin.
Identification of HTLV-1-PR:hFcγ1 Recombinant Protein by the Co-localization Assay
The cells containing the stable expression of HTLV-1-PR:hFcγ1 fusion protein were incubated with anti-Fcγ1 human antibody conjugated with fluorescein isothiocyanate (FITC; BioLegend, USA) to visualise secreted recombinant fusion protein in the sample via a fluorescence microscope (Nikon Eclipse E200, Japan).
Cell suspension of (4 × 106, 2 μL) were placed on immunofluorescence slides, dried overnight at 4 °C, and then fixed with methanol/acetone at − 20 °C. Cells were permeabilised with 0.2% (v/v) Triton-X 100 in PBS for 30 min and blocking was performed for 15 min with 3% (w/v) BSA (Sigma, USA) in PBS. The primary antibody was added at the optimal concentration, diluted in 3% (w/v) BSA in PBS, and incubated in a humidified chamber at 37 °C for 90 min. Next, slides were washed with 0.1% (w/v) BSA in PBS, and the secondary antibody was added at the optimal concentration of 3% (w/v) BSA and incubated in a humidified chamber for 60 min. Cells were washed in PBS in 0.1% (w/v) BSA and stained with ethidium bromide. After fixing the coverslip with a fluorescent mounting medium (Citifluor, UK) and sealing with nail varnish, samples were viewed by a fluorescence microscope.
HTLV-1-PR:hFcγ1 Protein Purification
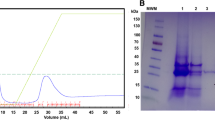
ÄKTA-purifier system (Amersham Biosciences, USA), an automated two-step purification method for Fc fusion proteins, was selected based on capture by affinity chromatography followed by “polishing” by gel filtration. Affinity chromatography on HiTrap recombinant protein A (GE Healthcare, USA) was selected due to its high selectivity for IgG1. Appropriate conditions for binding and elution buffers have been amended. Proteins were polished by gel filtration on HiLoad 16/60 Superdex 200 size exclusion columns.
Fluorescent Emission Assay
The biological activity of HTLV-1-PR:hFcγ1 recombinant protein was demonstrated and confirmed by using a synthetic fluorescent substrate. Fluorescent spectral scans were performed using a Synergy 4 Multi-Mode Microplate Reader (BioTek Instruments, USA). The λem and λex of HTLV-1 protease and substrate were recorded. The emission spectrum of the complex was measured at room temperature to determine the interaction between enzyme and substrate for evaluating the proper enzyme function.
Results
Molecular Modelling of the HTLV-1-PR:hFcγ1 Recombinant Protein
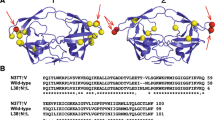
Twenty models at various refinement levels were generated using MODELER software (http://salilab.org/modeller/). Five models with the lowest molecular probability density function (molpdf) score were selected for structure refinements. Then, the most valid final model, which had an ERRAT score of 91.15%, was selected. The final model’s solid ribbon and solvent surface presentations were produced and depicted in Fig. 1A and B.
Bioinformatic analysis and homology modelling of HTLV-1-PR:hFcγ1 fusion protein. A Solid ribbon view of the modelled HTLV-1-PR:hFcγ1 dimer, positions of the disulphide bonds (Cys264-Cys324, Cys370-Cys428), and N-glycosylation sites: Asn168 and Asn300. B Solvent surface with glycosylation sites of HTLV-1-PR:hFcγ1 dimer
Bioinformatic Data for Protease Homology and Domains
The results revealed that the intact HTLV-1-PR:hFcγ1 fusion protein is a glycosylated homodimer with a theoretical pI of 8.30 and an apparent molecular weight of 50 kDa. The number of residues is 450 amino acids, whilst extinction coefficients are in units of M−1 cm−1, at 280 nm measured in water is 55,515 Abs 0.1% (= 1 g/L) 1.116, assuming that all pairs of Cys residues form cysteine. The N-terminal of the sequence considered is P (Pro).
The estimated half-life is > 20 h in mammalian reticulocytes in vitro, and > 20 h in yeast, in vivo. The instability index (II) is computed as 43.72, classifying the protein as unstable.
Moreover, N-linked glycosylation is one of the predominant post-translational modifications involved in several biological functions in eukaryotes. To predict N-glycosylation sites of the HTLV-1-PR:hFcγ1 sequence, the online server was used (https://services.healthtech.dtu.dk/service.php?NetNGlyc-1.0). Four potential N-glycosylation sites were predicted at Asn63, Asn168, Asn187, and Asn300, the last one located in the Fc fragment and the others in the extracellular region of recombinant protein (Fig. 2). These sites make proteins susceptible to highly glycosylation.
N-glycosylation analysis. The predicted N-glycosylation sites of the HTLV-1-PR:hFcγ1 fusion protein sequence were determined using the NetNGlyc 1.0 Server (https://services.healthtech.dtu.dk/service.php?NetNGlyc-1.0). The HTLV-1-PR:hFcγ1 contains four potential N-glycosylation sites
Cloning and Plasmid Sequencing
Confirmatory analysis of extracted plasmid by sequencing meets the appropriate quality, and PCR introduced no errors. Transformed colonies were screened by colony PCR using sequencing primers to check the proper integration of DNA fragments (Fig. 3).
Constructing and Characterization of High-Expression Eukaryotic pDR2DEF1α.PR:hFcγ1
The product was digested with XbaI and NotI (Fig. 4A) to be cloned in-frame with XbaI- and NotI-digested pDR2EF1a.hFcγ1 expression vector for making the appropriate expression vector (pDR2DEF1a.HTLV-1-PR:hFcγ1; Fig. 4B). The resulting vectors were sequenced at the Fc and protease junction site; the results indicated that the ligation had generated an in-frame construct. Figure 5 shows the nucleotide sequence of HTLV-1-PR:hFcγ1 in detail. After final confirmation, the DNA sequence of HTLV-1-PR:hFcγ1 recombinant protein was registered in the NCBI database (GenBank accession no. OM970797; https://www.ncbi.nlm.nih.gov/Genbank/update.html).
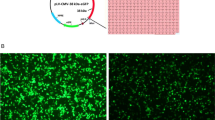
A Schematic image of pDR2DEF1a.HTLV-1-PR:hFcγ1 eukaryotic expression vector. B Agarose gel electrophoresis of the HTLV-1-PR product. Double-digestion expression vector, pDR2DEF1a.HTLV-1, and plasmid with HTLV-1. Lane L: 1-kb DNA size marker (Thermo Fisher Scientific, USA). (A) Undigested pTZ57R/T plasmid: a band of 3000 bp with incorporated sequences of two restriction sites, (B) digested plasmid: the HTLV-1-PR weight is in the 500 kb, (C) undigested expression vector, and (D) digested vector
The nucleotide sequences of recombinant HTLV-1-PR:Fc fused into the C-terminal of Fcγ1 (hinge region, Cγ2 and Cγ3). The ExPASy Translate tool (http://www.expasy.org/tools/dna) was used to translate the entire cDNA from a compiled sequence of HTLV-1-PR:Fc construct in pDR2ΔEF1α.hFc vector into the corresponding amino acid sequence (XbaI, HTLV-I PR, NotI, Linker, Fcγ1)
Confirmation of HTLV-1-PR:hFcγ1 Recombinant Protein Expression
Immunofluorescence staining of CHO cells showed the HTLV-1-PR:hFcγ1 recombinant fusion expression. The red signal shows CHO cells stained with ethidium bromide, and the green signal displays HTLV-1-PR:hFcγ1 recombinant fusion stained with FITC antibody (Fig. 6)
Fluorescent Emission Assay
The excitation wavelength was set to 400 nm for emission scans, the emission wavelengths were scanned from 400 to 750 nm in 1-nm increments, and the λem max was 450 nm. The emission spectra of enzyme and substrate are as follows: λem substrate: 540 nm, λex substrate: 350 nm, λem enzyme: 460 nm, and λex enzyme: 400 nm. Fig. 7 shows that the HTLV-1-PR and substrate fluorescence emission was slightly lower than the protein substrate composition.
Discussion
HTLV-1 protease was expressed as a recombinant Fc fusion protein in CHO cells. The construction of fusion proteins enables the production of recombinant proteins for structural, functional, therapeutic, and vaccine research. In this study, recombinant HTLV-1-PR:hFcγ1 was generated as N-terminal HTLV-1-PR fused to the C-terminal fragment of human Fcγ1, including the hinge region. Fcγ1 makes the HTLV-1-PR release from the engineered-CHO cells as a homodimer secretory protein, purifying it from the supernatant. Moreover, without a high concentration of cell debris and proteins in purification steps, the supernatant can be introduced HiTrap-rPA column FF as a powerful affinity chromatography technique to reach highly purified protein. For producing eukaryotic proteins in a secretory type, an Fcγ fusion construct is a popular approach, which confers remarkable biological and pharmacological properties upon the fusion protein. These include facilitated detection and purification of the target protein, decreased proteolysis of the target protein, and increased expression and secretion [26], in addition increasing the circulation serum half-life of protein for in vivo and therapeutic studies [21, 27].
A positive IFA test using anti-IgG1 indicated that the engineered homodimeric protein was appropriately folded; actually cloning, sub-cloning, and proper folding of bioengineered protein have been successful, as were confirmed by direct IFA assay. Of importance, HTLV-1-PR cleavage of a fluorescence-specific substrate revealed biological activity without impairment by the Fcγ1 tag. The efficacy of recombinant proteins for functional studies or therapeutic applications is affected by proper folding, stability, and post-translational modifications such as glycosylation and phosphorylation [26].
Altogether, these experiments indicated that engineered CHO correctly performed post-translational modifications and that HTLV-1-PR was enzymatically active. The appropriate activities of engineered HTLV-1-PR:hFcγ1 make it suitable for finding proper inhibitory agents for treating HTLV-1-associated diseases. Furthermore, the Fcγ1 tag of this viral protein can bind to FcγR-I (CD64), thus inducing cross-presentation and Th1 anti-viral immune responses. Targeting Fc receptor γ1 by Fcγ fragment strongly augments cross-presentation to initiate CTL responses, representing a critical step for anti-viral immune responses [28].
Therefore, it can also be tested in a suitable animal vaccination model, which is one of the limitations of this study. Some HTLV-1 glycoproteins such as ENV, Tax, and Pol are recognised by CTL for a protective immune response and pressure HTLV-1-infected cells. However, these responses could not eliminate HTLV-1 infection in vivo. Consequently, they might represent attractive components for HTLV-1 vaccine production [29]. In most attempts, the subunit vaccines for HTLV-1 infection failed to elicit a protective and long-lasting immune response even in CMI-inducing adjuvants [30]. Thus, in the present study, it was suggested that according to the Fc tag properties, the HTLV-1-PR Fc fusion recombinant protein could act in favour of increasing the immunogenicity and more understanding of its structure.
Regarding clinically used HIV-PR inhibitors as anti-retroviral drugs, notable differences in HTLV-1-PR have to be susceptible to those anti-viral agents. Accordingly, anti-HIV protease as a therapeutic agent invalidates further research [31]. Hence, the present study intended to introduce the Fc tag of the IgG1 antibody as a cell-mediated inducer by selective targeting of the APCs [21, 25, 32,33,34,35].
Recently, to address the novel SARS-CoV-2 pandemic, various vaccine formats and platforms have been developed [31, 36,37,38]. Amongst them, the Fc fusion protein with the receptor binding domain fragment of the virus demonstrated significant immunogenicity and clinical development for the induction and augmentation of neutralising IgG titres [39, 40]. Nevertheless, from some studies of FDA-approved Fc fusion drugs [25, 29, 35] and suitable vaccines against infectious diseases such as Ebola [41], HIV [42], and influenza [43], there have been no efforts to construct Fc fusion vaccines against HTLV-1.
Conclusion
Taken together, introducing HTLV-1-PR:hFcγ1 is an essential molecule for understanding its structural and physiological properties. Furthermore, proper expression and folding and maintenance of its enzymatic activity make it a suitable target for develo** inhibitors and exploring potent plant materials that suppress the HTLV-1-PR as a critical point of intervention in the viral life cycle. On the other hand, introducing the covalent binding of Fcγ1 in the C-terminal of this engineered recombinant protein as an effective vaccine can selectively target the FcγR-1 of APCs and induce appropriate CTL anti-HTLV-1 responses via cross-presentation.
Data Availability
All data that support the findings of this study are included in the manuscript and are available from the corresponding author upon reasonable request.
References
Ghezeldasht, S. A., Shirdel, A., Assarehzadegan, M. A., Hassannia, T., Rahimi, H., Miri, R., & Rezaee, S. R. (2013). Human T lymphotropic virus type I (HTLV-I) oncogenesis: Molecular aspects of virus and host interactions in pathogenesis of adult T cell leukemia/lymphoma (ATL). Iranian Journal of Basic Medical Sciences, 16(3), 179.
Martin, J. L., Maldonado, J. O., Mueller, J. D., Zhang, W., & Mansky, L. M. (2016). Molecular studies of HTLV-1 replication: An update. Viruses, 8(2), 31.
Marino-Merlo, F., Balestrieri, E., Matteucci, C., Mastino, A., Grelli, S., & Macchi, B. (2020). Antiretroviral therapy in HTLV-1 infection: An updated overview. Pathogens, 9(5), 342.
Boostani, R., Lotfinejad, N., Zemorshidi, F., Vahidi, Z., Rezaee, S. A., Farid, R., & Rafatpanah, H. (2021). Planning and management to control and eliminate HTLV-1 infection in Iran. Iranian Journal of Basic Medical Sciences, 24(3), 264.
Martin, F., Tagaya, Y., & Gallo, R. (2018). Time to eradicate HTLV-1: An open letter to WHO. The Lancet, 391(10133), 1893–1894.
Boross, P., Bagossi, P., Weber, I. T., & Tozser, J. (2009). Drug targets in human T-lymphotropic virus type 1 (HTLV-1) infection. Infectious Disorders-Drug Targets (Formerly Current Drug Targets-Infectious Disorders), 9(2), 159–171.
Sierra, S., & Walter, H. (2012). Targets for inhibition of HIV replication: Entry, enzyme action, release and maturation. Intervirology, 55(2), 84–97. https://doi.org/10.1159/000331995
Sperka, T., Miklossy, G., Tie, Y., Bagossi, P., Zahuczky, G., Boross, P., Matuz, K., Harrison, R. W., Weber, I. T., & Tözsér, J. (2007). Bovine leukemia virus protease: Comparison with human T-lymphotropic virus and human immunodeficiency virus proteases. Journal of General Virology, 88(7), 2052–2063.
Kheirabadi, M., Maleki, J., Soufian, S., & Hosseini, S. (2016). Design of new potent HTLV-1 protease inhibitors: In silico study. Molecular Biology Research Communications, 5(1), 19.
Nguyen, J.-T., Kato, K., Hidaka, K., Kumada, H.-O., Kimura, T., & Kiso, Y. (2011). Design and synthesis of several small-size HTLV-I protease inhibitors with different hydrophilicity profiles. Bioorganic and medicinal chemistry letters, 21(8), 2425–2429.
Mesters, J. R., Tan, J., & Hilgenfeld, R. (2006). Viral enzymes. Current Opinion in Structural Biology, 16(6), 776–786.
Kohl, N. E., Emini, E. A., Schleif, W. A., Davis, L. J., Heimbach, J. C., Dixon, R. A., Scolnick, E. M., & Sigal, I. S. (1988). Active human immunodeficiency virus protease is required for viral infectivity. Proceedings of the National Academy of Sciences, 85(13), 4686. https://doi.org/10.1073/pnas.85.13.4686
Larder, B. A. (1995). Viral resistance and the selection of anti-retroviral combinations. Journal of Acquired Immune Deficiency Syndromes and Human Retrovirology, 10(Suppl 1), S28-33.
Mahdi, M., Mótyán, J. A., Szojka, Z. I., Golda, M., Miczi, M., & Tőzsér, J. (2020). Analysis of the efficacy of HIV protease inhibitors against SARS-CoV-2’s main protease. Virology Journal, 17(1), 1–8.
Selvaraj, C., Singh, P., & Singh, S. K. (2014). Molecular modeling studies and comparative analysis on structurally similar HTLV and HIV protease using HIV-PR inhibitors. Journal of Receptors and Signal Transduction, 34(5), 361–371. https://doi.org/10.3109/10799893.2014.898659
Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W., & Lipman, D. J. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25(17), 3389–3402. https://doi.org/10.1093/nar/25.17.3389
Selvaraj, C., Singh, P., & Singh, S. K. (2014). Molecular insights on analogs of HIV PR inhibitors toward HTLV-1 PR through QM/MM interactions and molecular dynamics studies: Comparative structure analysis of wild and mutant HTLV-1 PR. Journal of Molecular Recognition, 27(12), 696–706.
Gopal, G. J., & Kumar, A. (2013). Strategies for the production of recombinant protein in Escherichia coli. The Protein Journal, 32(6), 419–425.
Alleva, D. G., Delpero, A. R., Scully, M. M., Murikipudi, S., Ragupathy, R., Greaves, E. K., Sathiyaseelan, T., Haworth, J. R., Shah, N. J., Rao, V., Nagre, S., Lancaster, T. M., Webb, S. S., Jasa, A. I., Ronca, S. E., Green, F. M., Elyard, H. A., Yee, J., Klein, J., … Zion, T. C. (2021). Development of an IgG-Fc fusion COVID-19 subunit vaccine, AKS-452. Vaccine, 39(45), 6601–6613. https://doi.org/10.1016/j.vaccine.2021.09.077
Mahmuda, A., Bande, F., Al-Zihiry, K. J. K., Abdulhaleem, N., Abd Majid, R., Hamat, R. A., Abdullah, W. O., & Unyah, Z. (2017). Monoclonal antibodies: A review of therapeutic applications and future prospects. Tropical Journal of Pharmaceutical Research, 16(3), 713–722.
Soleimanpour, S., Hassannia, T., Motiee, M., Amini, A. A., & Rezaee, S. (2017). Fcγ1 fragment of IgG1 as a powerful affinity tag in recombinant Fc-fusion proteins: Immunological, biochemical and therapeutic properties. Critical Reviews in Biotechnology, 37(3), 371–392.
Rosadas, C., Malik, B., Taylor, G. P., & Puccioni-Sohler, M. (2018). Estimation of HTLV-1 vertical transmission cases in Brazil per annum. PLoS Neglected Tropical Diseases, 12(11), e0006913. https://doi.org/10.1371/journal.pntd.0006913
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F., & Higgins, D. G. (1997). The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 25(24), 4876–4882. https://doi.org/10.1093/nar/25.24.4876
Krieger, E., Joo, K., Lee, J., Lee, J., Raman, S., Thompson, J., Tyka, M., Baker, D., & Karplus, K. (2009). Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: Four approaches that performed well in CASP8. Proteins, 77 Suppl 9(Suppl 9), 114–122. https://doi.org/10.1002/prot.22570
Czajkowsky, D. M., Hu, J., Shao, Z., & Pleass, R. J. (2012). Fc-fusion proteins: New developments and future perspectives. EMBO Molecular Medicine, 4(10), 1015–1028.
Langone, J. J. (1982). Use of labeled protein A in quantitative immunochemical analysis of antigens and antibodies. Journal of Immunological Methods, 51(1), 3–22.
Roopenian, D. C., & Akilesh, S. (2007). FcRn: The neonatal Fc receptor comes of age. Nature Reviews Immunology, 7(9), 715–725.
Guermonprez, P., Valladeau, J., Zitvogel, L., Théry, C., & Amigorena, S. (2002). Antigen presentation and T cell stimulation by dendritic cells. Annual Review of Immunology, 20(1), 621–667.
Kuo, C.-W.S., Mirsaliotis, A., & Brighty, D. W. (2011). Antibodies to the envelope glycoprotein of human T cell leukemia virus type 1 robustly activate cell-mediated cytotoxic responses and directly neutralise viral infectivity at multiple steps of the entry process. The Journal of Immunology, 187(1), 361–371.
Levin, D., Golding, B., Strome, S. E., & Sauna, Z. E. (2015). Fc fusion as a platform technology: Potential for modulating immunogenicity. Trends in Biotechnology, 33(1), 27–34.
Zhu, F.-C., Guan, X.-H., Li, Y.-H., Huang, J.-Y., Jiang, T., Hou, L.-H., Li, J.-X., Yang, B.-F., Wang, L., & Wang, W.-J. (2020). Immunogenicity and safety of a recombinant adenovirus type-5-vectored COVID-19 vaccine in healthy adults aged 18 years or older: A randomised, double-blind, placebo-controlled, phase 2 trial. The Lancet, 396(10249), 479–488.
Soleimanpour, S., Farsiani, H., Mosavat, A., Ghazvini, K., Eydgahi, M. R. A., Sankian, M., Sadeghian, H., Meshkat, Z., & Rezaee, S. A. (2015). APC targeting enhances immunogenicity of a novel multistage Fc-fusion tuberculosis vaccine in mice. Applied Microbiology and Biotechnology, 99(24), 10467–10480.
Akbarin, M. M., Rafatpanah, H., Soleimanpour, S., Amini, A. A., Arian, A., Mosavat, A., & Rezaee, S. A. (2022). TAX and HBZ: hFc Ɣ 1 proteins as targets for passive immunotherapy. Iranian Journal of Basic Medical Sciences, 25(5), 586–596. https://doi.org/10.22038/ijbms.2022.64787.14266
Shafifar, M., Mozhgani, S. H., Pashabayg, KR., Mosavat, A., Karbalaei, M., Norouzi, M., Rezaee, S. A. (2022). Selective APC-targeting of a novel Fc-fusion multi-immunodominant recombinant protein ((t)Tax-(t)Env:mFcγ2a) for HTLV-1 vaccine development. Life Science, 120920. https://doi.org/10.1016/j.lfs.2022.120920
Esser-Skala, W., Segl, M., Wohlschlager, T., Reisinger, V., Holzmann, J., & Huber, C. G. (2020). Exploring sample preparation and data evaluation strategies for enhanced identification of host cell proteins in drug products of therapeutic antibodies and Fc-fusion proteins. Analytical and Bioanalytical Chemistry, 412(24), 6583–6593.
Bos, R., Rutten, L., van der Lubbe, J. E., Bakkers, M. J., Hardenberg, G., Wegmann, F., Zuijdgeest, D., de Wilde, A. H., Koornneef, A., & Verwilligen, A. (2020). Ad26 vector-based COVID-19 vaccine encoding a prefusion-stabilised SARS-CoV-2 Spike immunogen induces potent humoral and cellular immune responses. npj Vaccines, 5(1), 1–11.
Ramasamy, M. N., Minassian, A. M., Ewer, K. J., Flaxman, A. L., Folegatti, P. M., Owens, D. R., Voysey, M., Aley, P. K., Angus, B., & Babbage, G. (2020). Safety and immunogenicity of ChAdOx1 nCoV-19 vaccine administered in a prime-boost regimen in young and old adults (COV002): A single-blind, randomised, controlled, phase 2/3 trial. The Lancet, 396(10267), 1979–1993.
Thompson, M. G., Burgess, J. L., Naleway, A. L., Tyner, H. L., Yoon, S. K., Meece, J., Olsho, L. E., Caban-Martinez, A. J., Fowlkes, A., & Lutrick, K. (2021). Interim estimates of vaccine effectiveness of BNT162b2 and mRNA-1273 COVID-19 vaccines in preventing SARS-CoV-2 infection among health care personnel, first responders, and other essential and frontline workers—Eight US locations, December 2020–March 2021. Morbidity and Mortality Weekly Report, 70(13), 495.
Chen, W.-H., Strych, U., Hotez, P. J., & Bottazzi, M. E. (2020). The SARS-CoV-2 vaccine pipeline: An overview. Current tropical medicine reports, 7(2), 61–64.
Lurie, N., Saville, M., Hatchett, R., & Halton, J. (2020). Develo** COVID-19 vaccines at pandemic speed. New England Journal of Medicine, 382(21), 1969–1973.
Konduru, K., Shurtleff, A., Bavari, S., Kaplan, G. (2013). Evaluation of ebolavirus glycoprotein Fc fusion protein as a subunit vaccine (P4417). Am Assoc Immnol,
Silva, M. T. (2010). Neutrophils and macrophages work in concert as inducers and effectors of adaptive immunity against extracellular and intracellular microbial pathogens. Journal of Leukocyte Biology, 87(5), 805–813.
Loureiro, S., Ren, J., Phapugrangkul, P., Colaco, C. A., Bailey, C. R., Shelton, H., Molesti, E., Temperton, N. J., Barclay, W. S., & Jones, I. M. (2011). Adjuvant-free immunisation with hemagglutinin-Fc fusion proteins as an approach to influenza vaccines. Journal of Virology, 85(6), 3010–3014.
Acknowledgements
The results described in this paper were part of the student thesis. This study was subjected to PhD thesis in Medical Immunology by Sanaz Ahmadi Ghezeldasht.
Funding
This study was financially supported by the Vice-Chancellor for Research and Technology, Mashhad University of Medical Sciences, Mashhad, Iran, under Grant MUMS 971409 and Vice-Chancellor for Research and Technology of ACECR-Razavi Khorasan, Mashhad, under Grant ACECR 3089–20.
Author information
Authors and Affiliations
Contributions
S. A. R. and A. M.: planned, data analysed, supervised, revised, and finalised the manuscript. S. A. G., M. M. H., and N. V.: performed the experiments and manuscript drafting. H. R. and S. A. S.: research advisors and preparing experimental facilities and kits. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Research Involving Human Participants and/or Animals
This work does not contain any studies with human participants or animals.
Informed Consent
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ahmadi Ghezeldasht, S., Momen Heravi, M., Valizadeh, N. et al. Development of a Novel HTLV-1 Protease: Human Fcγ1 Recombinant Fusion Molecule in the CHO Eukaryotic Expression System. Appl Biochem Biotechnol 195, 1862–1876 (2023). https://doi.org/10.1007/s12010-022-04259-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-04259-y