Abstract
Managing the nutritional status of strawberry plants is critical for optimizing yield. This study evaluated the potential of hyperspectral imaging (400–1,000 nm) to estimate nitrogen (N), phosphorus (P), potassium (K), and calcium (Ca) concentrations in strawberry leaves, flowers, unripe fruit, and ripe fruit and to predict plant yield. Partial least squares regression (PLSR) models were developed to estimate nutrient concentrations. The determination coefficient of prediction (R2P) and ratio of performance to deviation (RPD) were used to evaluate prediction accuracy, which often proved to be greater for leaves, flowers, and unripe fruit than for ripe fruit. The prediction accuracies for N concentration were R2P = 0.64, 0.60, 0.81, and 0.30, and RPD = 1.64, 1.59, 2.64, and 1.31, for leaves, flowers, unripe fruit, and ripe fruit, respectively. Prediction accuracies for Ca concentrations were R2P = 0.70, 0.62, 0.61, and 0.03, and RPD = 1.77, 1.63, 1.60, and 1.15, for the same respective plant parts. Yield and fruit mass only had significant linear relationships with the Difference Vegetation Index (R2 = 0.256 and 0.266, respectively) among the eleven vegetation indices tested. Hyperspectral imaging showed potential for estimating nutrient status in strawberry crops. This technology will assist growers to make rapid nutrient-management decisions, allowing for optimal yield and quality.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Global demand for food is increasing due to the continued rise in the world population (Godfray et al. 2010). Fertilizers play an important role in maximizing food production but fruit, nut, and seed yields are often sustained by excessive fertilizer use (Islam et al. 2022; Trejo-Téllez and Gómez-Merino 2014). Non-optimal fertilizer applications cause negative impacts on plant growth and development (Kulkarni and Goswami 2019). Over-fertilization, for example, can reduce yield and have negative impacts on the environment, while insufficient fertilization can also reduce yield and decrease product quality (González et al. 2015; Hapuarachchi et al. 2022; Pereira et al. 2015; Trejo-Téllez and Gómez-Merino 2014). Farmers often make decisions on nutrient amendments without full knowledge of the nutrient status of their crop (Bindraban et al. 2015; Islam et al 2022; Sheriff 2005) because current methods to examine plant nutrient status are laborious, costly, and time consuming (Yanli et al. 2015). Knowledge of plant nutrient status in real time would allow timely decisions on how much fertilizer needs to be added to a crop.
Hyperspectral imaging has been applied widely in agriculture, food, medicine, and other fields to estimate the internal qualities of scanned objects (Bai et al. 2018; Davur et al. 2023; ElMasry et al. 2012; Farrar et al. 2023; Gowen et al. 2007; Han et al. 2021; Huang et al. 2014; Malmir et al. 2020; Moscetti et al. 2015). Hyperspectral imaging combines spectroscopy with imaging techniques to acquire both spectral and spatial information simultaneously (ElMasry et al. 2012; Huang et al. 2014). Hyperspectral imaging is potentially non-destructive, low-cost, and reliable and has been applied to fruit, nuts, grains, and vegetables to estimate internal qualities such as total soluble solid concentration and moisture content, as well as firmness, ripeness, and shelf life (Bai et al. 2018; Davur et al. 2023; Gómez et al. 2006; Han et al. 2021; Han et al. 2023; Peng and Lu 2008; Pérez-Marín et al. 2009; Rajkumar et al. 2012; Ravikanth et al. 2017). Hyperspectral imaging allows the estimation of mineral nutrient concentrations such as nitrogen (N), phosphorus (P), potassium (K), and calcium (Ca) in the soil and leaves of many crops (Ferwerda et al. 2005; Mahajan et al. 2017; Pacumbaba and Beyl 2011; Pandey et al. 2017; Rodriguez et al. 2006; Tahmasbian et al. 2018; Yu et al. 2014). Hyperspectral imaging can also be used to estimate the concentrations of mineral nutrients including N, P, K, and Ca in avocado fruit (Kämper et al. 2020). Hyperspectral images obtained from the canopy or leaves have also been used to predict crop yield (Aparicio et al. 2000; Babar et al. 2006; Cao et al. 2015; Prasad et al. 2007; ** models to estimate nitrogen, phosphorus, potassium, and calcium concentrations. Scale bars = 1 cm
Hyperspectral imaging
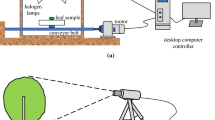
We used a laboratory-based hyperspectral imaging system (Benchtop System, Resonon, Bozeman, MT) with a 12-bit line scanner camera (Pika XC2, Bozeman, MT) to capture images (Farrar et al. 2021). The system comprised a hyperspectral camera with a 23-mm focal-length lens, a linear transition stage operated by a stepper motor, four current-controlled wide-spectrum quartz-halogen lights, and a main-control computer (Fig. A1).
We placed the samples on a black background on the transition stage of the camera. Images were captured using a visible/near-infrared hyperspectral imaging system in the spectral range of 400–1,000 nm. The spectral resolution was 1.3 nm, producing a total of 462 grayscale images that can be generated to form 462 bands of each hyperspectral image. The exposure time was adjusted to 27.3 ms and the stage moved at 1.23 mm s−1.
SpectrononPro software package (Version 2.94, Resonon, Bozeman, MT) was used to extract reflectance data (spectral information) of the acquired leaf, flower, unripe, and ripe fruit images. The mean corrected relative reflectance (R) was calculated from the raw spectral reflectance, R0, within SpectrononPro as shown in Eq. 1 (ElMasry et al. 2007):
where R0 was the raw spectral reflectance, D was the reflectance of a dark image (camera lens covered), and W was the reflectance of a white Teflon board that reflected approximately 99% of incident light. This corrected for the spectral curve of the leaf, flower, or fruit surface. The 100% reflectivity was scaled to 10,000 (integers) by default. The mean corrected relative reflectance was used for model development.
Nutrient analysis
Nitrogen concentrations of leaf, flower, unripe-fruit, and ripe-fruit samples were determined by combustion analysis using a LECO 928 analyzer (LECO, Saint Joseph, MI) (McGeehan and Naylor 1988; Muñoz-Huerta et al. 2013; Rayment and Higginson 1992). Calcium, phosphorus, and potassium concentrations were analyzed by using inductively coupled plasma–atomic emission spectroscopy on samples that were open-vessel digested with a 5:1 mixture of nitric and perchloric acids (Munter and Grande 1980).
Data analysis and model development
Image acquisition and data extraction were conducted using SpectrononPro software (Version 2.94, Resonon, Bozeman, MT). The mean raw reflectance was extracted by marking a region of interest (ROI) for each image. The ROI for leaf, flower, unripe-fruit, and ripe-fruit images contained the surface of one side of the leaf, flower, unripe fruit, or ripe fruit, respectively (Fig. A2). We mixed three fruit together for nutrient analysis in the case of composite fruit samples and so hyperspectral data from these three fruit were averaged prior to data analysis and model development. Spectral outliers in the samples, if any, were detected using a Hotelling’s T2 test (with 95% level of confidence) and removed from the data set (Farrar et al. 2021). The remaining data were divided randomly into two data sets, one used for calibration (80%) and the other (20%) used as a test data set to examine the precision of prediction using developed models (Table A1). We used a leave-one-out (full) cross-validation to evaluate the performance of the model (Dai et al. 2014; Tahmasbian et al. 2017; Zhang et al. 2013). This method uses the calibration data set but leaves one sample out of the calibration set each time and assesses the model using the remaining data. In the next iteration, another sample is left out randomly for the validation and this process continues until every sample is left out of the model once (Dai et al. 2014; Tahmasbian et al. 2017; Zhang et al. 2013).
We developed partial least square regression (PLSR) models. PLSR is one of the most frequently used modeling methods applied in hyperspectral imaging studies (De Silva et al. 2023; Han et al. 2023; Kämper et al. 2020; Mayr et al. 2021). The PLSR is commonly recommended when the dataset is small (Wold et al. 2001a). The PLSR is a linear multivariate model and relates data matrices of X and Y, the predicted and observable variables, respectively, by identifying smaller sets of predictors to perform a series of regressions (Wold et al. 2001a). These predictors have linear combinations, and the model is able to analyze datasets with noise and incomplete variables (Wold et al. 2001b). We also applied spectral pre-processing methods to decrease noise and improve model performance (Qin et al. 2013). The applied pre-processing methods were Smoothing Savitzky-Golay (Smoothing S-Golay, 1st derivative), Normalize, Derivative Savitzky-Golay (Derivative S-Golay), and Standard Normal Variate (SNV) (Qin et al. 2013). Developed models were assessed using the following indices: determination coefficients of calibration (R2C), validation (R2V), and prediction (R2P); root mean squares error (RMSE) of calibration (RMSEC), validation (RMSEV), and prediction (RMSEP); and ratio of performance to deviation (RPD). Only the model with highest coefficient of determination (R2), ratio of performance to deviation (RPD), and lowest root mean squares error (RMSE) was selected to determine the accuracy of estimation for each targeted nutrient (Table 1). The R2 and RMSE were calculated using Eqs. 2 and 3 (Yanli et al. 2015):
where n indicated the number of samples, yi and ŷi represented the reference and predicted values of the ith sample, respectively, and ȳ represented the mean of each reference value.
We then calculated the RPD using the test set (Farrar et al. 2021; Morellos et al. 2016). The RPD indicates the appropriateness of prediction. RPD classifications include RPD of 1.5–2.0 discriminates between high and low values, RPD of 2.0–2.5 provides a coarse quantitative prediction, and RPD > 2.5 and RPD > 3.0 provide good and excellent predictions, respectively (Nicolaï et al. 2007). The higher the RPD, the more robust is the model (Farrar et al. 2021; Kamruzzaman et al. 2012). RPD was defined using Eq. 4:
where SDTEST was the standard deviation of the observed values and RMSETEST was the root mean square error of the prediction from the test set.
We identified specific wavelengths that were important for predicting N, P, K, and Ca concentrations using β-coefficient values that carry predictive information. The wavelengths with the highest β-coefficients contribute most to the predictive ability of the models (Iqbal et al. 2013; Malmir et al. 2019; Tahmasbian et al. 2021; Xu et al. 2018). We assessed the model accuracies in predicting nutrient concentrations of fruit, leaves, and flowers by comparing the best-fit model for each plant part, with this model being the one that had the highest RPD and R2 of the dataset. Unscrambler® X software version 11 (CAMO Software Inc., Trondheim, Norway) was used for all computation, spectral data transformations, PLSR computations, outlier detection, and model development (Farrar et al. 2021).
Predicting yield and fruit mass
We attempted to predict both fruit yield and fruit mass using hyperspectral images. We also attempted to predict fruit yield and fruit mass from the macronutrient concentrations in leaf samples (n = 150) (Fig. 1). Yield was calculated as the total mass of fruit harvested from each plant during the study period of July to October 2018. Hyperspectral data from leaves, nutrient concentrations of leaves, and fruit mass were averaged for each plant, while yield was calculated as total fruit mass per plant. Stepwise regression was performed to evaluate linear regressions between the concentrations of N, P, K, and Ca in leaves or vegetation indices as the independent variable and fruit mass or yield as the dependent variable. Vegetation indices used to predict yield were Difference Vegetation Index (DVI), Modified Chlorophyll Absorption Ratio Index (MCARI), Modified Triangle Vegetation Index (MTVI), Normalized Difference Vegetation Index (NDVI), Photochemical Reflectance Index (PRI), Enhanced Vegetation Index (EVI), Ratio Vegetation Index (RVI), Infrared Percentage Vegetation Index (IPVI), Structure Independent Pigments Index (SIPI), and Red Edge Vegetation Stress Index (RVSI) (Rathod et al. 2013; Wang et al. 2018; Yu et al. 2018). The equations for all indices are presented in Table A2. Linear regressions were regarded as significant at p < 0.05.