Abstract
Rice source- and sink-associated traits are important for grain yield and are sensitive to environmental conditions. The continuing increase of CO2 concentrations in the atmosphere will become a major challenge for rice growth and development in the future due to changes in our climate such as extremes in temperature. To guarantee food safety, novel genetic loci need to be identified for source- and sink-associated traits that are specifically expressed under elevated CO2 conditions. Eighty chromosome segment substitution lines carrying japonica (Nipponbare) chromosome segments in the indica (9311) background were used in this study. QTL analysis was conducted for source- and sink-related traits, including flag leaf length, flag leaf width, flag leaf fresh weight, flag leaf dry weight, primary branch number, secondary branch number, grain number per panicle, panicle weight per plant, chlorophyll a, chlorophyll b, and carotenoid contents, under ambient CO2 concentrations and free-air CO2 enrichment. A total of 49 QTLs for these traits were detected on chromosomes 1, 3, 5, 6, 7, 9, and 12 under the two conditions; the variance explained by these QTLs varied from 6.22 to 38.15%. Among these QTLs, 19 of them were detected under the natural field conditions and 30 were detected in the elevated CO2 conditions. In addition, 2 and 13 QTLs were specifically expressed in the natural and CO2-enriched conditions, respectively. Our findings have important implications on the utilization of germplasm resources for ensuring food security under elevated CO2 levels, especially for QTLs that were specifically detected under the elevated CO2 condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa L.) is a major cereal crop consumed worldwide and increase of its grain yield is extremely important to meet the challenge of increasing population (Gross and Zhao 2014). Grain yield is sensitive to climate change, such as atmospheric CO2 concentration, temperature, and precipitation (Shi et al. 2012; Zhang et al. 2014; Liu et al. 2014). One study showed that the fast decline of total chlorophyll content, photosynthetic rate, and transpiration rate of the flag leaf in the late growth period led to a decrease of the seed setting rate, which greatly influences grain yield potential (Fu et al. 2012). Chlorophyll content is positively correlated with spike length, spikelet number per panicle, seed setting rate, and 1000-grain weight from heading to maturity in rice, which indicates that chlorophyll content in the flag leaf is closely related to yield after heading, and within a certain range, improvement of the chlorophyll content at the medium-late growth stage contributes to an increase of yield (Liu et al. 2018).
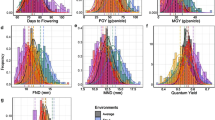
CO2 is a major greenhouse gas and the basic substrate for photosynthates (Xu et al. 2014; Jia 2015; Wang et al. 2018b). Plant growth and development are influenced by climate change, such as CO2 concentrations in the air (Chang et al. 2014). The increase of CO2 in the atmosphere not only promotes photosynthesis and the water utilization rate of plants, but also contributes to plant growth and crop yield (Song et al. 2008; Wang et al. 2018b). Furthermore, grain yield, grain number, panicle number, and spikelet fertility were significantly increased under elevated CO2 concentration (Fan et al. 2008a, b, The free-air CO2 enrichment (FACE) facilities were located in **. The CSSLs and the parents were planted in Yangzhou from May to October in 2018. Uniform seeds of all CSSLs and their parents were soaked in distilled water at 30 °C for 2 days and then germinated, covered by a damp washcloth at 30 °C for 12 h. The germinated seeds were sown in a paddy field at Hangzhou and transferred to Yangzhou after 30 days. The 30-day-old seedlings were transplanted into a 20-by-20-cm grid pattern in the flooded paddy in Yangzhou (119° 42′ 0″ E, 32° 35′ 5″ N). A randomized complete block design was used with three replications. In each replication, 10 plants per line were planted in one row with a spacing of 20 cm between plants and 20 cm between rows. Five plants in the middle of each row were randomly selected for measurement of chlorophyll a (Chl a), chlorophyll b (Chl b), carotenoids (Car), flag leaf length (LL), flag leaf width (LW), flag leaf fresh weight (FLFW), and flag leaf dry weight (FLDW) at 20 days after heading. Chl a, Chl b, and Car were measured according to Yang et al. (2016). At maturity, five plants in the middle were harvested, air-dried, and used for measurement of primary branch number (PBN), secondary branch number (SBN), grain number per panicle (GN), and panicle weight per plant (PWPP: grain weight with branch stem weight). The mean values of the five plants within each row were used as the trait value of each line, and the trait values of three replications of each line were averaged and used for data analysis. Using the NPB/9311 CSSLs, the effects of putative QTLs were tested according to variance analysis and t test with a significance level of P < 0.001. When a significant phenotypic difference between a CSSL and the recurrent parent was observed, it was considered the presence of a QTL. The marker interval of a corresponding QTL on a chromosomal location was determined according to the different phenotypic performance between the two contiguous CSSLs with overlap** segments. The phenotypic performance of CSSLs and parents is summarized in Fig. 1. All measured traits exhibited obvious differences between the parents (NPB and 9311) under both natural field and elevated CO2 conditions. The contents of Chl a, Chl b, and Car in NPB were higher than those of 9311. By contrast, the other traits measured were lower in NPB than in 9311. This indicates that NPB may have more alleles that enhance Chl a, Chl b, and Car but fewer alleles that enhance the remaining traits. With the exception of Chl a, Chl b, and Car, the mean phenotypic values of the other eight yield-related traits in the CSSL population were closer to those 9311, which is in accordance with the fact that the CSSLs were in the 9311 genetic background. Performance of source- and sink-associated traits in the CSSL population and parents. Histograms with dots represent traits measured under FACE and without dots represent traits measured under natural field conditions. Black and white arrows represent parental phenotypic values measured under FACE and natural field conditions, respectively. Arrows with blunt end represent Nipponbare, and arrows with notched end represent 9311 The correlation coefficients among the agronomic traits under natural and CO2-enriched conditions are presented in Table 1. For the photosynthetic pigments, Chl a, Chl b, and Car content showed significant positive correlations between each other under both conditions. For the flag leaf size traits, LL, FLFW, and FLDW displayed significant positive correlations between each other under both conditions, whereas LW only showed significant positive correlations with other traits under the natural field condition. For the panicle traits, PBN, SBN, GN, and PWPP exhibited significant positive correlations between each other under both conditions, except for PWPP and PBN, and PWPP and SBN under CO2 enrichment. These results indicate the existence of an interdependent relationship within the photosynthetic pigments, flag leaf size, and panicle traits. Significant negative correlations were detected between most of the photosynthetic pigments and flag leaf size as well as panicle traits under the natural field condition, whereas nonsignificant negative correlations were detected between most of the photosynthetic pigments and flag leaf size and panicle traits under CO2 enrichment. These results suggest that there is probably a negative association between photosynthetic pigment traits and flag leaf size and panicle traits under natural field conditions, whereas their association is diminished under high CO2 conditions. Under the natural field condition, 19 main-effect QTLs were detected, including one each for Chl a, Chl b, and Car content, and two for each of the remaining traits investigated (Fig. 2; Table 2). The contributions of a single QTL to the phenotypic variance ranged from 7.96 to 27.95%. There were six regions that had effects on two or more traits, among which three regions showed pleiotropic effects on both source- and sink-associated traits, while the remaining three regions exhibited pleiotropic effects on either source- or sink-associated traits. Main-effect QTLs detected in the natural field and elevated CO2 conditions. Chl a chlorophyll a, Chl b chlorophyll b, Car carotenoids, LL flag leaf length, LW flag leaf width, FLFW flag leaf fresh weight, FLDW flag leaf dry weight, PBN primary branch number, SBN secondary branch number, GN grain number per panicle, PWPP panicle weight per plant For the photosynthetic pigments, three QTLs, qChla-1N, qChlb-3N, and qCar-1N, were detected for Chl a, Chl b, and Car content, respectively. Two of the enhancing alleles were from NPB, and one was from 9311. This result is in accordance with the content of Chl a, Chl b, and Car being higher in NPB than in 9311. For the flag leaf-associated traits, eight QTLs were detected; seven of the enhancing alleles were from 9311 and one was from NPB. For the panicle-associated traits, we detected eight QTLs; five of the enhancing alleles were from 9311 and three were from NPB. This result is in accordance with 9311 showing higher phenotypic values for flag leaf- and panicle-associated traits than NPB. Under the elevated CO2 condition, 30 main-effect QTLs were detected and summarized in Fig. 2 and Table 2. The contributions of a single QTL to the phenotypic variance ranged from 6.22 to 38.15%. There were nine regions that had effects on two or more traits, among which only three regions showed pleiotropic effects on both source- and sink-associated traits, while the remaining six regions only exhibited pleiotropic effects on either source- or sink-associated traits. For the photosynthetic pigments, five QTLs were detected: only one each for Chl a and Car, and three for Chl b. Four of the enhancing alleles were from NPB, and one was from 9311. This result is in accordance with the content of Chl a, Chl b, and Car being higher in NPB than in 9311. For the flag leaf-associated traits, 13 QTLs were detected; nine of the enhancing alleles were from 9311 and four were from NPB. For the panicle-associated traits, 12 QTLs were detected; nine of the enhancing alleles were from 9311 and three were from NPB. This result is in accordance with 9311 having higher phenotypic values for flag leaf- and panicle-associated traits than those of NPB. A total of 49 main-effect QTLs distributed on chromosomes 1, 3, 5, 6, 7, 9, and 12 were detected among the two conditions, with more of the QTLs detected in the elevated CO2 condition than in the natural field condition (Fig. 2; Table 2). In addition, 2 and 13 QTLs were specifically detected under the natural and CO2 enriched conditions, respectively. These results suggest that some QTLs were detected depending on the CO2 concentration and that a considerable number of QTLs were specifically expressed under the elevated CO2 condition. Rice grain yield is largely determined by the source–sink relationship. Flag leaf area and its photosynthetic pigment content are key factors that influence source ability, while panicle traits greatly affect the sink capacity. The genetic basis underlying these traits remains poorly understood, particularly in elevated CO2 environments. In this study, QTL analysis for source- and sink-associated traits was conducted under natural field and elevated CO2 conditions. A total of 49 QTLs for Chl a, Chl b, Car, LL, LW, FLFW, FLDW, PBN, SBN, GN, and PWPP were detected among the two conditions, among which 19 and 30 QTLs were detected under natural field and elevated CO2 conditions, respectively. They fell into 16 regions of 7 chromosomes, and the phenotypic variation explained by a single QTL ranged from 6.22 to 38.15%. Notably, five of these regions were reported cloned genes that related to source- and sink-associated traits (Fig. 2). The five regions are RM6703–RM3738 that is co-located with GNP1, RM3513–RM1350 that covers GL3.1 and NOL, RM234–RM3555 that coincided in position with DEP2, RM3555–RM1306 that co-localized with Ghd7.1, and RM3700–STS9-6 that contains DEP1 (Huang et al. 2018). In this study, significant positive correlations were only detected between a few source- and sink-associated traits, while negative correlations were detected between multiple source- and sink-associated traits, such as Chl a and PWPP, Chl b and GN, and leaf length and PBN (Table 1). However, both the positive and negative correlations between the two types of traits were relatively weak. Furthermore, we detected six and nine QTL clusters under the natural field and enhanced CO2 conditions, respectively, but only three regions showed pleiotropic effects on both source- and sink-associated traits under each condition. The remaining QTL regions exhibited allelic variation for either source- or sink-associated traits. Thus, the source- and sink-associated traits may have relatively weak relationship genetically. With regard to these results, pyramiding of genes that have superior effects on source- or sink-associated traits is an effective and practive way to develop high-yielding cultivars. Rice source- and sink-associated traits are controlled by multiple genes and are easily influenced by environmental change, such as the temperature and CO2 concentration. It is important to identify genes that confer increased yield potential under elevated CO2 concentrations. In this study, a total of 34 QTLs were detected under both conditions. Among which, 13 QTLs were specifically detected under the elevated CO2 condition, and only two QTLs were specifically detected under the natural field condition. The QTLs expressed only under the elevated CO2 condition will be useful in breeding to improve rice yield potential. The size of the flag leaf is an important indicator for breeding cultivars with high yield potential. QTLs with pleiotropism on both leaf size and yield traits were also reported. For example, Wang et al. (2011) detected a QTL, qFL1, for flag leaf size on the short arm of chromosome 1 and fine-mapped it to a 31-kb region and found that qFL1 has pleiotropic effects on secondary branch number, number of spikelets per panicle, and panicle weight. Shen et al. (2012) fine-mapped a major QTL, qFLL6.2, that simultaneously controlled flag leaf length and yield traits. Zhang et al. (2015) fine-mapped a major QTL, qFLW7.2, responsible for flag leaf size and grain yield. In this study, three regions, RM3513–RM1350, RM3555–RM1306, and RM1246–RM7376, showed pleiotropic effects on both source- and sink-associated traits. However, whether the effects of these three regions result from a single QTL or a QTL cluster needs to be validated by further study.Materials and methods
Free-air CO2 enrichment
Trait evaluation
Data and QTL Analysis
Results
Phenotypic performance of CSSLs and parents
Correlation analysis of traits
QTL analysis under the natural field condition
QTL analysis under elevated CO2
Similarities and differences of QTL identification under the two conditions
Discussion
Data availability
The datasets supporting the conclusions of this article are provided within the article.
References
Ainsworth EA (2008) Rice production in a changing climate: a meta-analysis of responses to elevated carbon dioxide and elevated ozone concentration. Glob Change Biol 14(7):1642–1650
Bai X, Wu B, **ng Y (2012) Yield-related QTLs and their applications in rice genetic improvement. J Integr Plant Biol 54(5):300–311
Chang CY, Lin WD, Tu SL (2014) Genome-wide analysis of heat-sensitive alternative splicing in Physcomitrella patens. Plant Physiol 165(2):826–840
Cheng D, Yu H, Zou L, Teng Y, Zhu C (2018) Effects of elevated atmospheric CO2 concentration on the stability of soil organic carbon in different layers of a paddy soil. Chin Appl Ecol 29(8):2559–2565
Fan G, Cai Q, Wang C, Wan J, Zhu J (2005) QTL for yield and its components responded to elevated CO2 in rice (Oryza sativa L.). Acta Genet Sin 32(10):1066–1073
Fan G, Li J, Wang C, **e H, Xu C, Zhu J, Wan J, Cai Q (2006) QTL analysis for heading date of rice under free air CO2 enrichment conditions. Chin J Rice Sci 20(3):259–264
Fan G, Cai Q, Zhu J (2008a) Effect of elevated CO2 on yield and metabolism of carbohydrate during grain filling in rice. Chin Agric Sci Bull 24(10):272–275
Fan G, Cai Q, Wang C, Wan J, Zhu J (2008b) QTL analysis of panicle traits in rice (Oryza sativa L.) under free air CO2 enrichment. Sci Agric Sin 41(8):2227–2234
Fan G, Cai Q, **e H, Wang C, Wan J, Zhu J (2008c) Effect of elevated CO2 on the QTLs of grain shape and grain weight in rice. China Rice 6:28–32
Fu J, Cheng L, Huang Z, Wang Z, Yang J (2012) The relationship of leaf photosynthetic characteristics and root physiological traits with grain yield in super rice. Crop J 38(7):1264–1276
Gao ZY, Zhao SC, He WM, Guo LB, Peng YL, Wang JJ, Guo XS, Zhang XM, Rao YC, Zhang C, Dong GJ, Zheng FY, Lu CX, Hu J, Zhou Q, Liu HJ, Wu HY, Xu J, Ni PX, Zeng DL, Liu DH, Tian P, Gong LH, Ye C, Zhang GH, Wang J, Tian FK, Xue DW, Liao Y, Zhu L, Chen MS, Li JY, Cheng SH, Zhang GY, Wang J, Qian Q (2013) Dissecting yield-associated loci in super hybrid rice by resequencing recombinant inbred lines and improving parental genome sequences. Proc Natl Acad Sci USA 110(35):14492–14497
Gross BL, Zhao Z (2014) Archaeological and genetic insights into the origins of domesticated rice. Proc Natl Acad Sci USA 111(17):6190–6197
Hori K, Matsubara K, Yano M (1972) Genetic control of flowering time in rice: integration of mendelian genetics and genomics. Theor Appl Genet 67(5):717–725
Huang X, Qian Q, Liu Z, Sun H, He S, Luo D, **a G, Chu C, Li J, Fu X (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41(4):494–497
Jia X (2015) Study on determination method of available phosphorus content in soil. Agric Dev Equip 5:74
Jiang S, Zhang X, Zhang F, Xu Z, Chen W, Li Y (2012) Identification and fine map** of qCTH4, a quantitative trait loci controlling the chlorophyll content from tillering to heading in rice (Oryza sativa L.). J Hered 103(5):720–726
Li Z, Pinson SRM, Stansel JW, Paterson AH (1998) Genetic dissection of the source–sink relationship affecting fecundity and yield in rice (Oryza sativa L.). Mol Breed 4(5):419–426
Li F, Liu W, Tang J, Chen J, Tong H, Hu B, Li C, Fang J, Chen M, Chu C (2010) Rice DENSE AND ERECT PANICLE 2 is essential for determining panicle outgrowth and elongation. Cell Res 20(7):838–849
Li B, Wang N, Hao X, Li P (2019) Effects of interaction between elevated atmospheric CO2 concentration and drought on photosynthesis of soybean. J Shanxi Agric Sci 47(2):222–225, 258
Liu CG, Zhou XQ, Chen DG, Li JJ, Li JC, Chen YD (2014) Natural variation of leaf thickness and its association to yield traits in indica rice. J Integr Agric 13(2):316–325
Liu J, Yao X, Fan S, Li M, Guo N, Wang X, Wang J, Cheng W (2018) Map** of QTLs for chlorophyll content and panicle traits and their relationship in rice (Oryza sativa L.). J Shengyang Agri Univ 49(6):641–648
Nakano H, Yoshinaga S, Takai T, Arai-Sanoh Y, Kondo K, Yamamoto T, Sakai H, Tokida T, Usui Y, Nakamura H, Hasegawa T, Kondo M (2017) Quantitative trait loci for large sink capacity enhance rice grain yield under free-air CO2 enrichment conditions. Sci Rep 7:1827
Qi P, Lin YS, Song XJ, Shen JB, Huang W, Shan JX, Zhu MZ, Jiang L, Gao JP, Lin HX (2012) The novel quantitative trait locus GL3.1 controls rice grain size and yield by regulating Cyclin-T1;3. Cell Res 22(12):1666–1680
Sato Y, Morita R, Katsuma S, Nishimura M, Tanaka A, Kusaba M (2009) Two short-chain dehydrogenase/reductases, NON-YELLOW COLORING 1 and NYC1-LIKE, are required for chlorophyll b and light-harvesting complex II degradation during senescence in rice. Plant J 57(1):120–131
Sharma L, Dalal M, Verma RK, Kumar SVV, Yadav SK, Pushkar S, Kushwaha SR, Bhowmik A, Chinnusamy V (2018) Auxin protects spikelet fertility and grain yield under drought and heat stresses in rice. Environ Exp Bot 150:9–24
Shen B, Yu WD, Zhu YJ, Fan YY, Zhuang JY (2012) Fine map** of a major quantitative trait locus, qFLL6.2, controlling flag leaf length and yield traits in rice (Oryza Sativa L.). Euphytica 184(1):57–64
Shi W, Yin X, Struik PC, Solis C, **e F, Schmidt RC, Huang M, Zou Y, Ye C, Jagadish SVK (2017) High day- and night-time temperatures affect grain growth dynamics in contrasting rice genotypes. J Exp Bot 68(18):5233–5245
Song C, Zhang X, Liu X, Gao C (2008) Effect of soil organic matter on soil fertility and crop productivity. Syst Compr Study Agric Sci 24(3):357–362
Wang P, Zhou G, Yu H, Yu S (2011) Fine map** a major QTL for flag leaf size and yield-related traits in rice. Theor Appl Genet 123(8):1319–1330
Wang W, Cai C, Lam SK, Liu G, Zhu J (2018a) Elevated CO2 cannot compensate for japonica grain yield losses under increasing air temperature because of the decrease in spikelet density. Eur J Agron 99:21–29
Wang Q, ** K, Cao J (2018b). Effects of atmospheric CO2 concentration enhancement on photosynthetic physiological indexes and yield of maize leaves. J Shanxi Agri Sci 46(12):2051–2053, 2061
Wu Y, Wang Y, Mi XF, Shan JX, Li XM, Xu JL, Lin HX (2016) The QTL GNP1 Encodes GA20ox1, which increases grain number and yield by increasing cytokinin activity in rice panicle meristems. PLoS Genet 12(10):e1006386
Xu J, Liu G, Zhao X, Zhao S, Ren J, Hu J (2014) Study on the major and trace elements in soil of Yunnan farmland. Chin Agric Sci Bull 20(23):150–154
Yan W, Liu H, Zhou X, Li Q, Zhang J, Lu L, Liu T, Liu H, Zhang C, Zhang Z, Shen G, Yao W, Chen H, Yu S, **e W, **ng Y (2013) Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res 23(7):969–971
Yang L, Wang Y, Dong G, Gu H, Huang J, Zhu J, Yang H, Liu G, Han Y (2007) The impact of free-air CO2 enrichment (FACE) and nitrogen supply on grain quality of rice. Field Crops Res 102:128–140
Yang Y, Xu J, Huang L, Leng Y, Dai L, Rao Y, Chen L, Wang Y, Tu Z, Hu J, Ren D, Zhang G, Zhu L, Guo L, Qian Q, Zeng D (2016) PGL, encoding chlorophyllide a oxygenase 1, impacts leaf senescence and indirectly affects grain yield and quality in rice. J Exp Bot 67(5):1297–1310
Zhang GH, Li SY, Wang L, Ye WJ, Zeng DL, Rao YC, Peng YL, Hu J, Yang YL, Xu J, Ren DY, Gao ZY, Zhu L, Dong GJ, Hu XM, Yan MX, Guo LB, Li CY, Qian Q (2014) LSCHL4 from japonica cultivar, which is allelic to NAL1, increases yield of indica super rice 93–11. Mol Plant 7:1350–1364
Zhang B, Ye W, Ren D, Tian P, Peng Y, Gao Y, Ruan B, Wang L, Zhang G, Guo L, Qian Q, Gao Z (2015) Genetic analysis of flag leaf size and candidate genes determination of a major QTL for flag leaf width in rice. Rice 8(1):39
Zhou N, Jiang L, Wang Y, Zhu J, Yang L, Wang Y (2017) Effects of elevated atmospheric CO2 and temperature on dynamics of leaf chlorophyll contents and SPAD Value of rice in open-air field conditions. China J Rice Sci 31(5):524–532
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 31661143006, 31971872, 91735303), “Science and technology innovation project” of the Chinese Academy of Agriculture Sciences; and Hangzhou Scientific and Technological Program (20170432B03).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. LP D and XL L performed most of the research. WW Z, CJ W, L S, J H, DY R, Q Z, G C, and GJ D performed the trait investigation. DW X, GH Z, ZY G, LB G, and L Z analyzed the data. LP D wrote the article. LP D, Q Q, TM M and DL Z designed the research. DLZ revised the article.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dai, LP., Lu, XL., Zou, WW. et al. Map** of QTLs for source and sink associated traits under elevated CO2 in rice (Oryza sativa L.). Plant Growth Regul 90, 359–367 (2020). https://doi.org/10.1007/s10725-019-00564-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-019-00564-5