Abstract
A number of crop diseases are emerging at an alarming rate worldwide. Bacterial wilt of dry beans, caused by Curtobacterium flaccumfaciens pv. flaccumfaciens (Cff), is one of them. In Iran, this disease was first reported in 2012, which, since then, has rapidly spread across the major dry bean growing areas of the country causing severe yield losses. Previously, only two colony variants (yellow and orange) of the pathogen had been described from Iran in association with bacterial wilt of dry beans. In this study, we describe a new red-pigmented variant of Cff, isolated from dry bean seeds stored in seed banks of Khomein Bean Research Station, the major seed supplier in the region. Because Cff is a quarantine pathogen in Iran and elsewhere, with a potential threat for dry bean productions, more knowledge about the biology of this pathogen and epidemiology of the disease it causes are a prerequisite for the development of effective disease management strategies. Within this framework, we performed phenotypic and genetic characterization of the red-pigmented variant of the pathogen, in comparison with previously isolated yellow and orange variants, including pathogenicity, host range, bacteriocin production and genetic diversity. Our results showed a similar host range of different Cff variants although they differed in their aggressiveness. Yellow and orange variants of the pathogen were more aggressive on cowpea and common bean, respectively while the red variant showed the same level of aggressiveness on both hosts. Orange- and red-pigmented strains were separated from yellow-pigmented strains in the phylogeny of gyrB sequences. All orange- or red-pigmented strains were clustered in a separate branch from yellow-pigmented strains, except strain CffK31, in phylogeny based on rpoB sequences. In BOX-PCR analysis, Cff strains used in this study were clustered in two distinct genetic groups, with yellow variants of the pathogen separated from the orange and red variants. Overall, our results provide evidence of a remarkable diversity of Cff in Iran, which needs further in-depth investigation.











Similar content being viewed by others
References
Agarkova, I. V., Lambrecht, P. A., Vidaver, A. K., & Harveson, R. M. (2012). Genetic diversity among Curtobacterium flaccumfaciens pv. flaccumfaciens populations in the American High Plains. Canadian Journal of Microbiology, 58, 788–801. doi:10.1139/W2012-052.
Alipour, Y. (2013). A guide for diagnosis and detection of quarantine pests. Ministry of Jihad-e-Agriculture, Plant Protection Organization. http://www.ppo.ir/Uploads/English/Articles/Bacterium%20phytoplasma/Bacterial-tan-spot-Curtobacterium-flaccumfaciens-pv.pdf.
Chen, Y. F., Yin, Y. N., Zhang, X. M., & Guo, J. H. (2007). Curtobacterium flaccumfaciens pv. beticola, a new pathovar of pathogens in sugar beet. Plant Disease, 91, 677–684. doi:10.1094/PDIS-91-6-0677.
Collins, M. D., & Jones, D. (1983). Reclassification of Corynebacterium flaccumfaciens, Corynebacterium betae, Corynebacterium oortii and Corynebacterium poinsettiae in the genus Curtobacterium, as Curtobacterium flaccumfaciens comb. nov. Journal of General Microbiology, 129, 3545–3548.
EPPO. (2011). Curtobacterium flaccumfaciens pv. flaccumfaciens. Bulletin OEPP/EPPO, 41, 320–328. doi:10.1111/j.1365-2338.2011.02496.x.
Everitt, B. S., Landau, S., Leese, M., & Stahl, D. (2011). Cluster analysis (5th ed.). Chichester: Wiley. 348 pp.
Gross, D. C., & Vidaver, A. K. (1979). Bacteriocins of phytopathogenic Corynebacterium species. Canadian Journal of Microbiology, 25, 367–374.
Guimaraes, P. M., Palmano, S., Smith, J. J., Grossi, M. F. S., & Saddler, G. S. (2001). Development of a PCR test for the detection of Curtobacterium flaccumfaciens pv. flaccumfaciens. Antonie van Leeuwenhoek Journal of Microbiology, 80, 1–10. doi:10.1023/A:1012077425747.
Harveson, R. M., & Vidaver, A. K. (2007). First report of the natural occurrence of soybean bacterial wilt isolates pathogenic to dry beans in Nebraska. Plant Health Progress. doi:10.1094/PHP-2007-0822-01-BR.
Harveson, R. M., & Vidaver, A. K. (2008). A new color variant of the dry bean bacterial wilt pathogen (Curtobacterium flaccumfaciens pv. flaccumfaciens) found in western Nebraska. Plant Health Progress. doi:10.1094/PHP-2008-0815-01-BR.
Harveson, R.M., Schwartz, H.F., Urrea C.A., & Yonts C.D. (2015). Bacterial wilt of dry-edible beans in the central high plains of the U.S.: past, present, and future. Plant Disease, 99(12):1665–1677.
Hedges, F. (1922). A bacterial wilt of the bean caused by Bacterium flaccumfaciens nov. sp. Science, 55, 433–434.
Hsieh, T. F., Huang, H. C., Mündel, H. H., & Erickson, R. S. (2003). A rapid indoor technique for screening common bean (Phaseolus vulgaris L.) for resistance to bacterial wilt [Curtobacterium flaccumfaciens pv. flaccumfaciens (Hedges) Collins and Jones]. Revista mexicana de fitopatología, 21, 370–374.
Huang, H. C., Erickson, R. S., Yanke, L. J., Chelle, C. D., & Mündel, H. H. (2006). First report of the purple variant of Curtobacterium flaccumfaciens pv. flaccumfaciens, causal agent of bacterial wilt of bean, in Canada. Plant Disease, 90, 1262. doi:10.1094/PD-90-1262A.
Huang, H. C., Erickson, R. S., Balasubramanian, P. M., Hsieh, T. F., & Conner, R. L. (2009). Resurgence of bacterial wilt of common bean in North America. Canadian Journal of Plant Pathology, 31, 290–300. doi:10.1080/07060660909507600.
Jack, R. W., Tagg, J. R., & Ray, B. (1995). Bacteriocins of gram-positive bacteria. Microbiological Reviews, 59, 171–200.
Jacobs, J. L., Carroll, T. L., & Sundin, G. W. (2005). The role of pigmentation, ultraviolet radiation tolerance, and leaf colonization strategies in the epiphytic survival of phyllosphere bacteria. Microbial Ecology, 49, 104–113. doi:10.1007/s00248-003-1061-4.
Jukes, T. H., & Cantor, C. (1969). Mammalian protein metabolism (pp. 21–132). New York: Academic.
Keyworth, W. G., Howell, J., & Dowson, W. J. (1956). Corynebacterium betae (sp. nov.) The causal organism of silvering disease of red beet. Plant Pathology, 5, 88–90.
Larkin, M. A., Blackshields, G., Brown, N. P., Chenna, R., McGettigan, P. A., McWilliam, H., Valentin, F., Wallace, I. M., Wilm, A., Lopez, R., Thompson, J. D., Gibson, T. J., & Higgins, D. G. (2007). ClustalW and ClustalX version 2. Bioinformatics, 23, 2947–2948. doi:10.1093/bioinformatics/btm404.
Madden, L.V., Hughes, G., & van den Bosch, F. (2007). The study of plant disease epidemics. APS Press, St. Paul. University of Minnesota. 421pp.
Mandel, M., Guba, E. F., & Litsky, W. (1961). The causal agent of bacterial blight of American holly. Bacteriological Proceedings, 61, 61.
Maringoni, A. C., & Kurozawa, C. (2002). Curtobacterium flaccumfaciens pv. flaccumfaciens typification by bacteriocin. Pesquisa Agropecuária Brasileira, 37, 1339–1346. doi:10.1590/S0100-204X2002000900019.
Osdaghi, E., & Lak, M. R. (2015). Occurrence of a new orange variant of Curtobacterium flaccumfaciens pv. flaccumfaciens, causing common bean wilt in Iran. Journal of Phytopathology, 163, 867–871. doi:10.1111/jph.12322.
Osdaghi, E., Alizadeh, A., Shams-bakhsh, M., & Lak, M. R. (2009). Evaluation of common bean lines for their reaction to the common bacterial blight pathogen. Phytopathologia Mediterranea, 48, 461–468.
Osdaghi, E., Pakdaman Sardrood, B., Bavi, M., Akbari Oghaz, N., Kimiaei, S., & Hadian, S. (2015a). First report of Curtobacterium flaccumfaciens pv. flaccumfaciens causing cowpea bacterial wilt in Iran. Journal of Phytopathology, 163, 653–656. doi:10.1111/jph.12300.
Osdaghi, E., Taghavi, S. M., Fazliarab, A., Elahifard, E., & Lamichhane, J. R. (2015b). Characterization, geographic distribution and host range of Curtobacterium flaccumfaciens: An emerging bacterial pathogen in Iran. Crop Protection, 78, 185–192. doi:10.1016/j.cropro.2015.09.015.
Pirone, P. P., & Bender, T. R. (1941). A new bacterial disease of poinsettiae. New Jersey Agriculture and Experimental Nursery Disease Notes, 14, 13–16.
Richert, K., Brambilla, E., & Stackebrandt, E. (2005). Development of PCR primers specific for the amplification and direct sequencing of gyrB genes from microbacteria, order Actinomycetales. Journal of Microbiological Methods, 60, 115–123. doi:10.1016/j.mimet.2004.09.004.
Richert, K., Brambilla, E., & Stackebrandt, E. (2007). The phylogenetic significance of peptidoglycan types: molecular analysis of the genera Microbacterium and Aureobacterium based upon sequence comparison of gyrB, rpoB, recA and ppk and 16SrRNA genes. Systematic and Applied Microbiology, 30, 102–108. doi:10.1016/j.syapm.2006.04.001.
Rohlf, F. J. (2008). NTSYSpc: numerical taxonomy system, ver. 2.20. Setauket: Exeter Publishing, Ltd.
Saaltink, G. J., & Maas Geesteranus, P. H. (1969). A new disease of tulip caused by Corynebacterium oortii nov. sp. Netherlands Journal of Plant Pathology, 75, 123–128.
SAS. (1999). SAS software, Version 8 of the SAS System for windows. Cary: SAS Institute Inc.
Schaad, N. W., Jones, J. B., & Chun, W. (2001). Laboratory guide for identification of plant pathogenic bacteria (3rd ed.). St. Paul: APS. 379 pp.
Schuster, M. L., & Christiansen, D. W. (1957). An orange-colored strain of Corynebacterium flaccumfaciens causing bean wilt. Phytopathology, 47, 51–53.
Schuster, M. L., & Sayre, R. M. (1967). A coryneform bacterium induces purple-colored seed and leaf hypertrophy of Phaseolus vulgaris and other leguminosae. Phytopathology, 57, 1064–1066.
Schuster, M. L., Vidaver, A. K., & Mandel, M. (1968). A purple pigment-producing bean wilt bacterium Corynebacterium flaccumfaciens var. violaceum n. var. Canadian Journal of Microbiology, 14, 423–427.
Smith, N. C., Hennessy, J., & Stead, D. E. (2001). Repetitive sequence-derived PCR profiling using the BOX-A1R primer for rapid identification of the plant pathogen Clavibacter michiganensis subspecies sepedonicus. European Journal of Plant Pathology, 107, 739–748. doi:10.1023/A:1011955811847.
Soares, R. M., Fantinato, G. G. P., Darben, L. M., Marcelino-Guimarães, F. C., Seixas, C. D. S., & Carneiro, G. E. S. (2013). First report of Curtobacterium flaccumfaciens pv. flaccumfaciens on soybean in Brazil. Tropical Plant Pathology, 38, 452–454. doi:10.1590/S1982-56762013000500012.
Souza, V. L., Maringoni, A. C., & Krause-Sakate, R. (2006). Genetic variability in Curtobacterium flaccumfaciens isolates. Summa Phytopathologica, 32, 170–176. doi:10.1590/S0100-54052006000200012.
Tagg, J. R., Dajani, A. S., & Wannamaker, L. W. (1976). Bacteriocins of gram-positive bacteria. Bacteriology Reviews, 40, 722–756.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., & Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30, 2725–2729. doi:10.1093/molbev/mst197.
Tegli, S., Sereni, A., & Surico, G. (2002). PCR-based assay for the detection of Curtobacterium flaccumfaciens pv. flaccumfaciens in bean seeds. Letters in Applied Microbiology, 35, 331–337. doi:10.1046/j.1472-765X.2002.01187.x.
Urrea, C. A., & Harveson, R. M. (2014). Identification of sources of bacterial wilt resistance in common beans (Phaseolus vulgaris L.). Plant Disease, 98, 973–976. doi:10.1094/PDIS-04-13-0391-RE.
van Schoonhoven, V. A., & Pastor-Corrales, M. A. (1987). CIAT. Standard evaluation of bean germplasm. Cali: Centro Internacional de Agricultura Tropical (CIAT). 54 p.
Versalovic, J., Schneider, M., de Bruijn, F. J., & Lupski, J. R. (1994). Genomic fingerprinting of bacteria using repetitive sequence-based polymerase chain reaction. Methods in Molecular and Cell Biology, 5, 25–40.
Weisburg, W. G., Barns, S. M., Pelletier, D. A., & Lane, D. J. (1991). 16S ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology, 173, 697–703.
Yim, K. O., Lee, H. I., Kim, J. H., Lee, S. D., Cho, J. H., & Cha, J. S. (2012). Characterization of phenotypic variants of Clavibacter michiganensis subsp. michiganensis isolated from Capsicum annuum. European Journal of Plant Pathology, 133, 559–575. doi:10.1007/s10658-011-9927-7.
Young, J. M., Watson, D. R. W., & Dye, D. W. (2004). Reconsideration of Arthrobacter ilicis (Mandel et al. 1961) Collins et al. 1982 as a plant-pathogenic species. Proposal to emend the authority and description of the species. Request for an Opinion. International Journal of Systematic and Evolutionary Microbiology, 54, 303–305. doi:10.1099/ijs.0.02929-0.
Acknowledgments
We thank the following institutions: Negin Bazre Pars co. in Khomein County, Markazi province for providing common bean seeds (cvs. Sadri, Derakhshan, Dorsa), Plant Improvement Institute (SPII) in Karaj, Alborz province for providing soybean seeds (cv. Katool), and Agricultural and Natural Resources Research Center of Safiabad, in Khuzestan province for supplying mung bean (cvs. L173 and Partow) and cowpea (cvs. L1057 and Mashhad) seeds. Financial support for this study was provided by Shiraz University.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Twenty plant-associated bacterial strains used in this study as bacteriocin indicator strains. The grey color scale shows the differences in sensitivity patterns among the indicator strains to the bacteriocins of yellow- (20Y), red- (50R), and orange- (80O) pigmented variants of Curtobacterium flaccumfaciens pv. flaccumfaciens. The sensitivity groups show the statistically different groups generated in Duncan grou** analysis (Fig. 2) (DOCX 14 kb)
Fig. S1
Production of bacteriocin by Curtobacterium flaccumfaciens pv. flaccumfaciens (called as producer strains) against different plant pathogenic bacteria (called as indicator strains). Clear zones appeared around the colonies of producer strains indicate inhibition after 3 days post incubation. a: strain 80O vs Pectobacterium carotovorum subsp. carotovorum, b: strain 20Y vs Pseudomonas fluorescens (CHAO), c: strain 80O vs Rhodococcus fascians, and d: strain 20Y vs Agrobacterium tumefaciens (JPG 757 kb)
Fig. S2
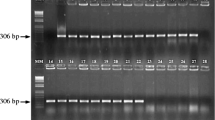
Agarose gel of PCR products obtained with the primer pairs CF4/CF5 (198 bp; a) and CffFOR2/CffREV4 (306 bp; b). Lanes: 1-20: bacterial strains as numbered in Table 1. 21: Xanthomonas axonopodis pv. phaseoli strain Araxa1, 22: 100 bp DNA Ladder. All the Curtobacterium flaccumfaciens strains were detected with the primer pairs, except strains CffG105 and Cmmeg20 which were isolated from tomato and eggplant, respectively (JPG 276 kb)
Rights and permissions
About this article
Cite this article
Osdaghi, E., Taghavi, S.M., Hamzehzarghani, H. et al. Occurrence and characterization of a new red-pigmented variant of Curtobacterium flaccumfaciens, the causal agent of bacterial wilt of edible dry beans in Iran. Eur J Plant Pathol 146, 129–145 (2016). https://doi.org/10.1007/s10658-016-0900-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-016-0900-3