Abstract
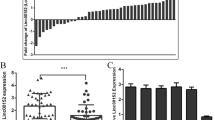
Hypoxic microenvironments are intricately linked to malignant characteristics of glioblastoma multiforme (GBM). Long non-coding ribonucleic acids (lncRNAs) have been reported to be involved in the progression of GBM and closely associated with hypoxia. Nevertheless, the differential expression profiles as well as functional roles of lncRNAs in GBM cells under hypoxic conditions remain largely obscure. We explored the expression profiles of lncRNAs in hypoxic U87 cells as well as T98G cells using sequencing analysis. The effect of differentially expressed lncRNAs (DElncRNAs) was assessed through bioinformatic analysis. Furthermore, the expression of lncRNAs significantly dysregulated in both U87 and T98G cells was further validated using quantitative reverse-transcription polymerase chain reaction (qRT-PCR). Relevant cell functional experiments were also conducted. We used predicted RNA-binding proteins (RBPs) to construct an interaction network via the interaction prediction module. U87 and T98G cells showed dysregulation of 1115 and 597 lncRNAs, respectively. Gene Ontology (GO) analysis indicated that altered lncRNA expression was associated with nucleotide-excision repair and cell metabolism in GBM cells. Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis revealed the association between dysregulated lncRNAs and the Hippo signaling pathway under hypoxia. The dysregulation of six selected lncRNAs (ENST00000371192, uc003tnq.3, ENST00000262952, ENST00000609350, ENST00000610036, and NR_046262) was validated by qRT-PCR. Investigation of lncRNA-microRNA (miRNA)-mRNA networks centered on HIF-1α demonstrated cross-talk between the six validated lncRNAs and 16 related miRNAs. Functional experiments showed the significant inhibition of GBM cell proliferation, invasion, and migration by the knockdown of uc003tnq.3 in vitro. Additionally, uc003tnq.3 was used to construct a comprehensive RBP-transcription factor (TF)-miRNA interaction network. The expression of LncRNAs was dysregulated in GBM cells under hypoxic conditions. The identified six lncRNAs might exert important effect on the development of GBM under hypoxic microenvironment.







Similar content being viewed by others
Availability of data and materials
The raw data supporting the conclusion of this article have been uploaded to Supplementary Appendix 4.
Abbreviations
- GBM:
-
Glioblastoma multiforme
- lncRNA:
-
Long non-coding ribonucleic acid
- HIF-1α:
-
Hypoxia-induced factor-1α
- qRT-PCR:
-
Quantitative reverse-transcription polymerase chain reaction
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ncRNA:
-
Non-coding ribonucleic acid
- ceRNA:
-
Competing endogenous RNA
- miRNA:
-
MicroRNA
- DElncRNA:
-
Differentially expressed lncRNA
- siRNA:
-
Small interfering RNA
- CCK-8 assay:
-
Cell counting kit-8 solution assay
- OD:
-
Optical density
- one-way ANOVA:
-
One-way analysis of variance
- FC:
-
Fold change
- BP:
-
Biological process
- CC:
-
Cellular component
- MF:
-
Molecular function
- UHRF1:
-
Ubiquitin-like with PHD and RING finger domains 1
- UPAT:
-
UHRF1-protein associated transcript
- HIE:
-
Hypoxic ischemic encephalopathy
- IGFBP3:
-
Insulin-like growth factor binding protein 3
- RBP:
-
RNA-binding protein
- TF:
-
Transcription factor
References
Adams BD, Parsons C, Walker L, Zhang WC, Slack FJ (2017) Targeting noncoding RNAs in disease. J Clin Invest 127:761–771
Alifieris C, Trafalis DT (2015) Glioblastoma multiforme: pathogenesis and treatment. Pharmacol Ther 152:63–82
Amer RG, Ezz El Arab LR, Abd El Ghany D, Saad AS, Bahie-Eldin N, Swellam M (2022) Prognostic utility of lncRNAs (LINC00565 and LINC00641) as molecular markers in glioblastoma multiforme (GBM). J Neurooncol 158:435–444
Amjadi N, Talayeh M, Momeni M, Mansouri N (2023) The comparison of umbilical cord artery pH in newborns with and without thick meconium stained amniotic fluid. Cell Mol Biomed Rep 3:222–226
Bilal I, **e S, Elburki MS, Aziziaram Z, Ahmed SM, Jalal Balaky ST (2021) Cytotoxic effect of diferuloylmethane, a derivative of turmeric on different human glioblastoma cell lines. Cell Mol Biomed Rep 1:14–22
Bischoff FC, Werner A, John D, Boeckel JN, Melissari MT, Grote P, Glaser SF, Demolli S, Uchida S, Michalik KM, Meder B, Katus HA, Haas J, Chen W, Pullamsetti SS, Seeger W, Zeiher AM, Dimmeler S, Zehendner CM (2017) Identification and functional characterization of hypoxia-induced endoplasmic reticulum stress regulating lncRNA (HypERlnc) in pericytes. Circ Res 121:368–375
Cai X, Qiu W, Qian M, Feng S, Peng C, Zhang J, Wang Y, Wang Y (2020) A Candidate prognostic biomarker complement factor i promotes malignant progression in glioma. Front Cell Dev Biol 8:615970
Cai X, Chen Z, Huang C, Shen J, Zeng W, Feng S, Liu Y, Li S, Chen M (2022) Development of a novel glycolysis-related genes signature for isocitrate dehydrogenase 1-associated glioblastoma multiforme. Front Immunol 13:950917
Chang YN, Zhang K, Hu ZM, Qi HX, Shi ZM, Han XH, Han YW, Hong W (2016) Hypoxia-regulated lncRNAs in cancer. Gene 575:1–8
Chen Z, Su S, Yang M, Wang F, Chen M (2022) Profiling and bioinformatics analyses of differential circular RNA expression in glioblastoma multiforme cells under hypoxia. J Mol Neurosci 72:2451–2463
Colwell N, Larion M, Giles AJ, Seldomridge AN, Sizdahkhani S, Gilbert MR, Park DM (2017) Hypoxia in the glioblastoma microenvironment: sha** the phenotype of cancer stem-like cells. Neuro Oncol 19:887–896
Domenech M, Hernandez A, Plaja A, Martinez-Balibrea E, Balana C (2021) Hypoxia: the cornerstone of glioblastoma. Int J Mol Sci 22:12608
Fattahi S, Kosari-Monfared M, Golpour M, Emami Z, Ghasemiyan M, Nouri M, Akhavan-Niaki H (2020) LncRNAs as potential diagnostic and prognostic biomarkers in gastric cancer: a novel approach to personalized medicine. J Cell Physiol 235:3189–3206
Hua Q, Mi B, Xu F, Wen J, Zhao L, Liu J, Huang G (2020) Hypoxia-induced lncRNA-AC020978 promotes proliferation and glycolytic metabolism of non-small cell lung cancer by regulating PKM2/HIF-1alpha axis. Theranostics 10:4762–4778
Jonas K, Calin GA, Pichler M (2020) RNA-binding proteins as important regulators of long non-coding RNAs in cancer. Int J Mol Sci 21:2969
Kanehisa M, Sato Y, Furumichi M, Morishima K, Tanabe M (2019) New approach for understanding genome variations in KEGG. Nucleic Acids Res 47:D590–D595
Kechin A, Boyarskikh U, Kel A, Filipenko M (2017) cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J Comput Biol 24:1138–1143
Koehler J, Sandey M, Prasad N, Levy SA, Wang X, Wang X (2020) Differential expression of miRNAs in hypoxia (“HypoxamiRs”) in three canine high-grade glioma cell lines. Front Vet Sci 7:104
Le Rhun E, Preusser M, Roth P, Reardon DA, van den Bent M, Wen P, Reifenberger G, Weller M (2019) Molecular targeted therapy of glioblastoma. Cancer Treat Rev 80:101896
Liao Y, Luo Z, Lin Y, Chen H, Chen T, Xu L, Orgurek S, Berry K, Dzieciatkowska M, Reisz JA, D’Alessandro A, Zhou W, Lu QR (2022) PRMT3 drives glioblastoma progression by enhancing HIF1A and glycolytic metabolism. Cell Death Dis 13:943
Linhares P, Carvalho B, Vaz R, Costa BM (2020) Glioblastoma: is there any blood biomarker with true clinical relevance? Int J Mol Sci 21:5809
Liu L, Shen P, Zheng B, Yu W, Ji J, **ao Y (2020) Comparative genomic analysis of 19 clinical isolates of tigecycline-resistant Acinetobacter baumannii. Front Microbiol 11:1321
Peng Z, Liu C, Wu M (2018) New insights into long noncoding RNAs and their roles in glioma. Mol Cancer 17:61
Postepska-Igielska A, Giwojna A, Gasri-Plotnitsky L, Schmitt N, Dold A, Ginsberg D, Grummt I (2015) LncRNA Khps1 regulates expression of the proto-oncogene SPHK1 via triplex-mediated changes in chromatin structure. Mol Cell 60:626–636
Qin Y, Zhang X, Chen Y, Zhang W, Du S, Ren C (2023) Prognostic analysis of a hypoxia-associated lncrna signature in glioblastoma and its pan-cancer landscape. J Neurol Surg A Cent Eur Neurosurg. https://doi.org/10.1055/a-2070-3715
Santangelo R, Rizzarelli E, Copani A (2020) Role for metallothionein-3 in the resistance of human U87 glioblastoma cells to temozolomide. ACS Omega 5:17900–17907
Touat M, Idbaih A, Sanson M, Ligon KL (2017) Glioblastoma targeted therapy: updated approaches from recent biological insights. Ann Oncol 28:1457–1472
Wang Y, Liu S (2021) LncRNA GHET1 promotes hypoxia-induced glycolysis, proliferation, and invasion in triple-negative breast cancer through the Hippo/YAP signaling pathway. Front Cell Dev Biol 9:643515
Wang H, Cao D, Wu F (2018) Long noncoding RNA UPAT promoted cell proliferation via increasing UHRF1 expression in non-small cell lung cancer. Oncol Lett 16:1491–1498
Wu W, Hu Q, Nie E, Yu T, Wu Y, Zhi T, Jiang K, Shen F, Wang Y, Zhang J, You Y (2017) Hypoxia induces H19 expression through direct and indirect Hif-1α activity, promoting oncogenic effects in glioblastoma. Sci Rep 7:45029
**ong LL, Xue LL, Du RL, Zhou HL, Tan YX, Ma Z, ** Y, Zhang ZB, Xu Y, Hu Q, Bobrovskaya L, Zhou XF, Liu J, Wang TH (2020) Vi4-miR-185-5p-Igfbp3 network protects the brain from neonatal hypoxic ischemic injury via promoting neuron survival and suppressing the cell apoptosis. Front Cell Dev Biol 8:529544
Xu W, Ying Y, Shan L, Feng J, Zhang S, Gao Y, Xu X, Yao Y, Zhu C, Mao W (2015) Enhanced expression of cohesin loading factor NIPBL confers poor prognosis and chemotherapy resistance in non-small cell lung cancer. J Transl Med 13:153
Yamazaki T, Souquere S, Chujo T, Kobelke S, Chong YS, Fox AH, Bond CS, Nakagawa S, Pierron G, Hirose T (2018) Functional domains of NEAT1 architectural lncRNA induce paraspeckle assembly through phase separation. Mol Cell 70:1038-1053.e1037
Yao ZT, Yang YM, Sun MM, He Y, Liao L, Chen KS, Li B (2022) New insights into the interplay between long non-coding RNAs and RNA-binding proteins in cancer. Cancer Commun (lond) 42:117–140
Zealy RW, Fomin M, Davila S, Makowsky D, Thigpen H, McDowell CH, Cummings JC, Lee ES, Kwon SH, Min KW, Yoon JH (2018) Long noncoding RNA complementarity and target transcripts abundance. Biochim Biophys Acta Gene Regul Mech 1861:224–234
Zhou X, Li X, Yu L, Wang R, Hua D, Shi C, Sun C, Luo W, Rao C, Jiang Z, Wang Q, Yu S (2019) The RNA-binding protein SRSF1 is a key cell cycle regulator via stabilizing NEAT1 in glioma. Int J Biochem Cell Biol 113:75–86
Zuin J, Casa V, Pozojevic J, Kolovos P, van den Hout M, van Ijcken WFJ, Parenti I, Braunholz D, Baron Y, Watrin E, Kaiser FJ, Wendt KS (2017) Regulation of the cohesin-loading factor NIPBL: role of the lncRNA NIPBL-AS1 and identification of a distal enhancer element. PLoS Genet 13:e1007137
Acknowledgments
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China (grant number: 81902521), Shanghai Sailing Program (grant number: 19YF1432800), and Research Project of **nhua Hospital (grant number: XH1936).
Author information
Authors and Affiliations
Contributions
MC and BWC designed and supervised the project. XMC, MSQ, KZ, and YZL conducted the experiments and performed data analysis. BWC and XMC wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10528_2023_10597_MOESM8_ESM.docx
Supplementary file8 Table 4. Top 10 BP, CC and MF terms corresponding to upregulated and downregulated lncRNAs in U87 (DOCX 24 KB)
10528_2023_10597_MOESM10_ESM.docx
Supplementary file10 Table 6. Top 10 BP, CC and MF terms corresponding to upregulated and downregulated lncRNAs in T98G(DOCX 23 KB)
10528_2023_10597_MOESM11_ESM.docx
Supplementary file11 Table 7. Top 10 pathways corresponding to upregulated and downregulated lncRNAs in T98G(DOCX 17 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cai, X., Qian, M., Zhang, K. et al. Profiling and Bioinformatics Analyses of Hypoxia-Induced Differential Expression of Long Non-coding RNA in Glioblastoma Multiforme Cells. Biochem Genet (2023). https://doi.org/10.1007/s10528-023-10597-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-023-10597-1