Abstract
Breast cancer (BC) represents a multifaceted malignancy, with escalating incidence and mortality rates annually. Chemotherapy stands as an indispensable approach for treating breast cancer, yet drug resistance poses a formidable challenge. Through transcriptome data analysis, we have identified two sets of genes exhibiting differential expression in this context. Furthermore, we have confirmed the overlap between these genes and those associated with exosomes, which were subsequently validated in cell lines. The investigation screened the identified genes to determine prognostic markers for BC and utilized them to formulate a prognostic model. The disparities in prognosis and immunity between the high- and low-risk groups were validated using the test dataset. We have discerned different BC subtypes based on the expression levels of prognostic genes in BC samples. Variations in prognosis, immunity, and drug sensitivity among distinct subtypes were examined. Leveraging data from single-cell sequencing and prognostic gene expression, the AUCell algorithm was employed to score individual cell clusters and analyze the pathways implicated in high-scoring groups. Prognostic genes (CCT4, CXCL13, MTDH, PSMD2, and RAB27A) were subsewoquently validated using RT-qPCR. Consequently, we have established a model for predicting prognosis in breast cancer that hinges on drug resistance and ERGs. Furthermore, we have evaluated the prognostic value of this model. The genes identified as prognostic markers can now serve as a reference for precise treatment of this condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BC), a pervasive malignancy worldwide, is witnessing an increase in both incidence and mortality rates [1]. This trend underscores the necessity for effective BC management strategies, including prompt diagnosis, precise treatment, and notably, chemotherapy. Although chemotherapy remains a cornerstone of BC treatment, leading to tumor shrinkage and potential cures, it is increasingly impeded by the emergence of drug resistance in various BC subtypes [2]. This resistance not only diminishes the effectiveness of chemotherapy but also complicates the anti-tumor treatment process. Therefore, it is crucial to investigate the underlying mechanisms of resistance.
Drug resistance can manifest in various subtypes of BC through diverse mechanisms. For example, certain genes are commonly or differentially expressed across BC types and might impact drug resistance [3, 4]. Research has concluded that the overexpression of the human epidermal growth factor receptor (HER-2) significantly affects BC prognosis. The use of trastuzumab has markedly improved patient outcomes in cases of HER-2 overexpression, while resistance to HER-2 targeted agents is increasing [5]. Furthermore, mutations in the estrogen receptor gene have been specifically linked to harm in metastatic BC [6]. These genetic elements contribute to the growing complexity of drug resistance at the molecular level, potentially leading to unmanageable BC events and increased fatality. It has been demonstrated that genes associated with drug resistance have a significant impact in this context [6]. For instance, PTX-induced exosomal circBACH1 stimulates the migration and stemness of BC cells, influencing drug resistance mechanisms by absorbing miR-217 to increase the expression of G3BP2 [Correlation analysis of scRNA data The analysis of single-cell RNA data was conducted using the R package “Seurat.” The initial step involved performing data quality control, during which genes associated with mitochondria and erythrocytes were excluded. Subsequently, the following screening parameters were applied: the “nFeature_RNA” criterion selected values above 200 and below 100,000; the “percent.mt” criterion selected values below 20, and the “nCount_RNA” criterion selected values below 100,000. In the normalization process, we identified the top 2,000 genes with high variability across all cells using the “LogNormalize” and “vst” methods. Here, “nFeature_RNA” refers to the number of detected genes in each single-cell sample. “percent.mt” represents the percentage of mitochondrial gene expression in the total gene expression. “nCount_RNA” indicates the total number of RNA molecules detected in each single-cell sample. “LogNormalize” denotes the normalization of gene expression for each cell to make the data distribution more akin to a normal distribution. “vst” refers to the variance-stabilizing transformation applied to minimize technical noise in the data. We subsequently conducted principal component analysis (PCA) analysis, scaled the data, and retained the first 15 principal components for clustering purposes. The cells were then subjected to cluster analysis with a resolution of 0.5. The cell clusters were labeled using the R package "singleR." We then used the t-SNE algorithm to generate a two-dimensional visualization of the cell clusters and their respective types. Additionally, we employed the R package "AUCell" to calculate the area under the curve (AUC) value for each cluster. Our study identified the population of cells with high scores and the pathway scores that had significant enrichment in this particular cell population based on prognostic genes. The gene set we used is "h.all.v2022.1.Hs.symbols.gmt." Total RNA was extracted from A10 breast epithelial cells and MDA-MB-231 breast cancer cells, both of which were obtained from the Cell Bank of the Chinese Academy of Sciences, using the AG RNA Extraction Kit II (AG21022). Reverse transcription was performed using the Evo M-MLV RT Kit, and gDNA Clean for qPCR (AG11711). RT-PCR was conducted with the SYBR Green Premix Pro Taq HS qPCR Kit (AG11701) in a real-time fluorescent quantitative PCR analyzer from Agilent. Primer information for the validated genes is provided in Table S1 in the Supplementary material. Statistical analyses were performed using R software (version 4.2.0) and SPSS software (version 26.0). For continuous and categorical variables, we utilized the Wilcoxon rank-sum and chi-square tests, respectively. All analyses were deemed statistically significant at a p-value of < 0.05. We denote p-values less than 0.05 and 0.01 with “*” and “**,” respectively.RT-qPCR method for the verification of prognostic genes
Statistical analysis
Result
Identification of differential drug resistance and exosome-related genes in BC
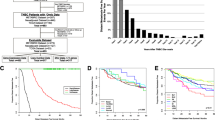
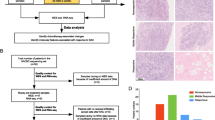
The technical process outlined in this article is depicted in Fig. 1. A differential analysis of transcriptome data was conducted to identify genes associated with drug resistance by comparing the drug-sensitive and drug-resistant groups from the GSE163882 dataset. The results of the differential analysis, including the heatmap and volcano plot, are presented in Fig. 2A, B. Details of DEGs are provided in the “GSE163882_diff.xls” file in the Supplementary Material. Figure 2C illustrates the Venn diagram produced from the intersection of DEGs and exosome-associated genes, resulting in 47 shared genes. Gene enrichment analysis using GO on these shared genes revealed pathways linked to BC and BC-associated medication responses, as shown in Fig. 2D. The biological significance of these pathways will be discussed in the following section. Box plots were utilized to evaluate the expression of communication-related genes in breast cancer patients and their controls from the TCGA cohort. The results showcased significant variations in the expression levels of most of these genes between the two groups.
Results of differential analysis between drug-sensitive and resistant groups in GEO cohort. A and B are the heat map and volcano map obtained from the difference analysis, respectively. C is the Venn diagram of the intersection of DEGs and exosome-related genes. D is the GO enrichment result of EGDR. E is the expression boxplot of EGDR in the transcriptome data of TCGA cohort
Establishment and verification of prognostic model
In this study, we developed a prognostic model for BC using Lasso-Cox regression analysis. Five gene markers were identified through the Lasso algorithm, based on the optimal λ value (refer to Fig. 3A, B). Specifically, LASSO calculates a coefficient for each gene. The first graph is the coefficient distribution graph. Each line in the graph represents a gene, and the end of these genes will point to an ordinate (representing the coefficient of the gene) from which the gene can be further screened after taking λ as the threshold value. Figure 3B shows that the coefficients of five genes could be retained. Utilizing the calculated risk scores, the model categorized BC samples into high- and low-risk groups. We presented the results of survival analysis and ROC curve examination for these groups on the TCGA dataset in Fig. 3C, D. Importantly, a significant variance in survival time was observed between the two cohorts. The model for estimating patient prognosis indicated that the risk score yielded area under the curve (AUC) values of 0.632, 0.655, and 0.631 for predicting patient survival after 1, 3, and 5 years, respectively. PCA of the samples from the training group revealed a clear separation of low and high-risk groups on a two-dimensional plot (refer to Fig. 3F). The forest plot in Fig. 3F emphasizes the identification of five genes as prognostic markers through multivariate Cox regression analysis. In addition, we performed an independent prognostic analysis to assess whether the five genes constituting the prognostic model could be used as independent prognostic factors to predict breast cancer survival (Fig. 5). The results showed that only CXCL13 and MTDH could be used as independent prognostic factors.
Results of prognostic model construction. A is the least absolute shrinkage and selection operator (LASSO) regression of prognostic genes. B is the cross-validation used to adjust the choice of parameters in the LASSO regression. C and D are the results of survival analysis and ROC analysis of the prognostic model, respectively. E is a scatter plot after dimensionality reduction using PCA for samples from the high- and low-risk groups. F is the forest plot obtained by multivariate Cox regression
Subsequently, a test dataset was utilized to validate the model’s performance. Figures 4A, B provide details on the Kaplan–Meier survival and ROC curves for the high-risk and low-risk groups in the test set, indicating a significant difference in survival between the two groups. The AUC values for predicting survival at 1, 3, and 5 years were 0.766, 0.623, and 0.633, respectively. Figure 4C, D presents the outcomes of both PCA and multivariate Cox regression analyses. The results further confirmed that CXCL13 and MTDH can be used as independent prognostic factors of breast cancer to predict the survival of breast cancer patients.
External validation of prognostic models. A is the KM survival curve of the prognostic model on the validation set. B is the result of ROC analysis on the validation set. C is the forest plot obtained from multivariate COX regression analysis to screen independent prognostic factors. D is the result of PCA analysis on the validation set
Prognostic validation was conducted for the five prognostic genes identified in the training set (refer to Fig. 5). Notable disparities in survival were observed between the high and low expression groups of each of the five genes when breast cancer samples were stratified accordingly. Additionally, DMFS for the high and low expression groups, segregated by these prognostic genes, was validated. The KM survival curves displayed in Fig. 6 reveal significant DMFS survival differences for the two expression groups.
Construction and validation of nomogram model
The present study outlines the development of a nomogram model (depicted in Fig. 7A) utilizing baseline information from patients (including age and clinical stage) and a risk score. This is done to assess the potential risk associated with BC. The calibration curve for this model, as shown in Fig. 7B, illustrates the extent of deviation from the ideal model. Additionally, the ROC analysis results for the nomogram model are presented in Fig. 7C, revealing AUC values of 0.867, 0.769, and 0.725 for predicting the 1-, 3-, and 5-year survival rates of patients, respectively. Notably, these values surpass those obtained using the prognostic model. Figure 7D displays curves exhibiting the C-index over time for various models, with the nomogram model achieving the highest C-index. According to the DCA depicted in Fig. 7E, the predictive accuracy of the nomogram model surpasses that of age and clinical stage.
Construction and validation of the nomogram model. A is a nomogram model constructed based on clinical stage, baseline information of patients, and risk score. B is the calibration curve of the nomogram model. C is the result of ROC curve analysis of the nomogram model. D is the result of C-index analysis. E is the DCA result of the nomogram model
Results of correlation analysis of immune infiltration
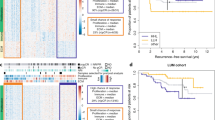
The ssGSEA algorithm was utilized to assess variances in the abundance of immune cell infiltration and immune function scores of multiple immune cells in high- and low-risk groups, determined by gene expression levels in BC specimens (refer to Fig. 8A). Significantly different infiltration abundance and immune function scores were observed in immunocytes of both groups. Figure 8C–L displays scatter plots delineating specific genes that exhibit substantial links with immune cells. Further outcomes can be found in the supplementary materials’ Immune section.
Evaluation of immune infiltration in BC samples based on ssGSEA analysis. A is the boxplot of the difference in the abundance of immune cell infiltration in the high- and low-risk groups. B is the boxplot of the difference in immune function between the high- and low-risk groups. C–L is the correlation analysis of prognostic genes and immune cells; correlation analysis of risk scores and immune cells
The dissimilarities in the immune landscape between the high-risk and low-risk groups were evaluated from multiple viewpoints in this study. Notably, the results show statistically significant differences between the two groups in ESTIMATE score, immune score, stromal score, and tumor purity utilizing the ESTIMATE analysis assessment outcomes (refer to Fig. 9A–D). Additionally, we conducted an investigation into variances in the expression of HLA-related genes and immunoassay sites among both groups (see Fig. 9E, F) and examined variations in immunotherapy (see Fig. 10). Figure 11 presents IC50 values of specific drugs that significantly differ between the two groups (further information can be found in the supplementary materials, in the Drug1 folder).
ESTIMATE analysis between high- and low-risk groups, expression analysis of HLA-related genes and immunoassay sites. A–D is the result of ESTIMATE score, immune score, Stromal score, and tumor purity in the high- and low-risk groups based on ESTIMATE analysis, respectively. E and F is a boxplot based on the expression of HLA-related genes and immunoassay sites in the high- and low-risk groups, respectively
BC ty** based on prognostic genes
In this study, we utilized prognostic gene expression profiles to classify BC using the NMF clustering algorithm. As depicted in Fig. 12A, the NMF algorithm exhibits distinct phenotypic characteristics, RSS, and dispersion distribution across various cluster numbers. According to the most cophenetic curve in this figure, it can be judged that the leading point of the largest decrease is 2. Therefore, the NMF algorithm achieves the best performance when the number of clusters is 2. The figure highlights that the performance of the NMF algorithm peaks when the cluster number is set to 2, as illustrated in the consensus plot in Fig. 12B. Additionally, Fig. 12C and D depicts the survival curves for the two subtypes and the level of immune cell infiltration. Notably, significant differences in both survival and immune cell infiltration were observed between the two subtypes. Finally, we investigated the disparities in drug sensitivity between the two BC subtypes, as presented in Fig. 13 in the Drug2 folder in the supplementary material.
Identification of BC subtypes based on prognostic genes. A is phenotype distribution, rss distribution and dispersion distribution when rank = 2–10; B is the consensus map of NMF clustering. C is the prognostic survival curve of the two molecular subtypes. D is the difference in the abundance of immune cell infiltration among different subtypes
Results of immune landscape and AUCell analysis at the single-cell level
To generate multiple cell clusters, we conducted quality control, normalization, dimensionality reduction, and clustering using the R package Seurat. The cell clusters were then annotated using the R package singleR. Figure 14A depicts the scores of different cell clusters associated with various cell types. Figure 14B presents the final cell type annotations based on these scores. In Fig. 14C–F, the bubble and violin plots illustrate the expression of prognostic genes in different cell clusters. Notably, we observed high expression of MTDH and RAB27A in various immune cell types. Furthermore, we utilized the AUCell algorithm (Fig. 15A, B) to evaluate the scores for different cell types, with macrophages exhibiting the highest score. Figure 15C displays the different cell types within the macrophage population with elevated AUCell scores. The Discussion section provides a comprehensive explanation of the relationship between these pathways and breast cancer, along with potential drug targets.
Results of cell clustering and annotation analysis of single-cell sequencing data. A is a heatmap that annotates the cell population based on the single-cell algorithm. B is the result of the visualization of the cell population based on tnse dimensionality reduction. C is a bubble plot of the expression of prognostic genes on different cell populations. D–F is a violin plot of the expression of prognostic genes on different cell populations
Results of the analysis based on AUCell. A is a scatter plot obtained by scoring different cell types by AUCell based on tsne. B is a violin plot based on the distribution of AUCell scores for different cell populations. C is the bar graph drawn for the top cell types and their corresponding AUC scores in the macrophage population
Additionally, based on the gene expression matrix and cell type information from scRNA-seq data, we used AUCell to calculate the expression levels of genes in specified gene sets within each cell. Scatter plots of the top 10 pathways before scoring are provided in Supplementary Material Fig. S1. We will analyze the relationship between these pathways and the progression of breast cancer in detail in the Discussion section.
The results of RT-qPCR were verified for the prognostic genes
Differences in the expression of prognostic genes between normal mammary epithelial cells and breast cancer cells were subsequently verified. Each sample was tested in triplicate, and a melting curve analysis was conducted to assess the specificity of amplification. Figure 16A–E presents a box plot illustrating the disparities in gene expression between the two cell groups. The verification process confirmed distinctions in the expression of the majority of genes between the two cell groups.
Discussion
The heterogeneity observed in BC presents a significant challenge to the efficacy of isochemoth erapy across different stages and subtypes. Our study addresses this challenge by investigating personalized treatment strategies, with a particular focus on the role of exosomes in inhibiting tumor growth and regulating the immune response. We performed an overlap analysis of DEGs and ERGs between resistant and sensitive groups, followed by GO enrichment analysis of the common genes. This approach highlights the potential for tailored therapies based on genetic profiling, as shown in our flowchart. We also elucidated the role of exosomes in BC treatment by examining their interactions with key genes and pathways. These findings from our comprehensive analysis underscore the importance of the exosome pathway in the development of targeted BC therapies. A study by Du et al. demonstrated that everolimus has the potential to decrease the expression of protein kinase B (AKT) in BC cells, indicating its promise as a therapeutic agent [15]. According to research by Zheng et al., adriamycin-induced apoptotic MCF-7 cells show potential for BC treatment [16]. A study by Irene Lopez-Mateo suggested that thyroid hormone receptor β may act as a tumor suppressor in BC [17]. Ji-Young Park and colleagues induced the expression of manganese superoxide dismutase in BC cells through the protein kinase B and extracellular signal-regulated kinase signaling pathways [18]. Philip Bischoff and colleagues found that the progression of BC depends on the disruption of adherent tissue integrity [19]. In this study, Siyoung Choi and colleagues discovered that mineralization of the matrix inhibited integrin-mediated mechanical signal transduction by using a collagen matrix with adjustable mineralization, thereby inducing a less proliferative stem cell-like phenotype in BC cells [20].
We have also explored the prognostic potential of genes such as CCT4, CXCL13, MTDH, PSMD2, and RAB27A. These genes exhibit significant associations with BC prognosis in both TCGA and GEO cohorts. Our flowchart illustrates how these genetic markers can be employed in a clinical context to predict treatment outcomes and identify high-risk patients. This comprehension is crucial for advancing novel therapeutic targets and personalized treatment regimens. They have been validated as significantly associated with BC prognosis in the TCGA and GEO cohorts. Further exploration into the roles of these genes in BC initiation and progression may lead to the discovery of new treatment targets. According to Wang et al., the inhibition of CCT4 by anti-carlin-β treatment disrupts protein balance and specifically inhibits tumor cell growth [21]. The significance of CXCL13 in an effective T cell response to anti-PD-L1 therapy was demonstrated through the analysis of scRNA-seq and scRNA-ATAC data in triple-negative BC [22]. Shen et al. observed frequent overexpression of MTDH in BC patients with a poor prognosis. MTDH promotes metastasis and treatment resistance by interacting with staphylococcal nuclease domain-containing 1 (SND1). This complex plays a critical role in suppressing anti-tumor T cell responses in BC [23]. RAB27A has been identified as a mediator of human milk BC stem cells, promoting the growth of mammospheres. Additionally, RAB27A has been found to be synergistically inhibited [24].
The risk model utilized in our study is based on LASSO-Cox analysis, effectively categorizing breast cancer (BC) patients into high-risk and low-risk groups. This model has been seamlessly integrated into our flowchart and functions as a crucial tool for evaluating patient outcomes and customizing treatment strategies. We further investigated the variations in survival and immune landscapes between these risk groups, employing the ssGSEA algorithm to underscore noteworthy variations in immune cell infiltration and function. As depicted in Fig. 8C, five prognostic genes exhibit a statistically significant correlation with distinct immune cells. Consistent with the findings of Li et al.’s study, clinical evidence indicates an inhibitory effect on BC metastasis. Moreover, certain sources suggest that the TAM/CXCL1/NF-κB/FOXP3 signaling pathway could potentially serve as a therapeutic target to modulate Tregs and enhance immunotherapy for BC [25]. Janakiram and colleagues have comprehensively reviewed the interplay between T cell inhibition and immunotherapy in BC [26].
The immune-related aspects of breast cancer, particularly the role of HLA-associated genes, constitute the focal point of our discussion. Studies have illuminated the intricate relationship between these genes and the progression of breast cancer, along with their influence on therapy response. Our flowchart, incorporating these findings, underscores the significance of immunological assessments in formulating comprehensive treatment strategies. A substantial number of HLA-related genes and immunoassay loci exhibit noteworthy differences between high- and low-risk groups. The majority of these genes have been linked to breast cancer and its treatment. In their review, Zheng et al. expounded on the role of HLA-G in the immune microenvironment of breast cancer, emphasizing its utility in identifying new biomarkers for breast cancer [27]. Woll et al. reported that HLA-A2 dimers can accurately measure and track antigen-specific T cell immune responses in peptide vaccine clinical trials [28]. It was observed that a reduction in EZH435 expression in the human breast cancer cell line MDA-MB-231 significantly increased HLA-DRA mRNA expression, even without IFN-γ stimulation [29]. In a study involving 89 patients with metastatic breast cancer and 50 age- and sex-matched healthy volunteers, Song et al. investigated the percentage of peripheral blood T lymphocyte subsets and plasma cytokine levels. The study revealed that an elevated level of CD8(+) CD28(−) suppressor T lymphocytes may independently predict progression-free survival during follow-up after chemotherapy [30]. Abdullah et al. identified that pharmacological combinations of NRP1 with FGFR-targeted kinase inhibitors could be an effective treatment for patients with drug-resistant metastatic breast cancer [31].
We have identified BC subtypes by employing prognostic genes and subsequently explored variations in prognosis, immune response, and drug sensitivity among these subtypes. Notably, therapeutic agents such as oxaliplatin [32], paclitaxel [33], and gefitinib [34] have shown effectiveness in treating breast cancer.
Finally, we employed single-cell RNA sequencing (scRNA-seq) data to offer a more intricate insight into gene expression across various cell types. This enabled the identification of cell populations at risk. Seamlessly integrated into our flowchart, this approach underscores the significance of comprehending gene expression patterns within the context of breast cancer (BC) treatment and prognosis. The utilization of the AUCell algorithm facilitated the identification of high-scoring cell populations and pathways, many of which are pertinent to BC and its treatment, thus affirming the practicality of our model in clinical settings. X-box binding protein 1 (XBP1) plays a crucial role in the unfolded protein response (UPR). Vahid Arabkari et al. discovered the involvement of XBP1 in UPR, correlating it with endocrine resistance in BC [35]. In BRCA2 mutation carriers, Rachel Joyce et al. identified mTORC1 as a potential target for breast cancer prevention [36]. Results from Theresa E Hickey et al.’s experiments suggest that androgen receptor (AR) exerts a tumor-suppressive role in estrogen receptor (ER)-α-positive BC [37]. Kurt W Evans et al. found that oxidative phosphorylation represents a metabolic vulnerability in triple-negative BC, potentially exploitable in combination therapy [38]. Yvette Drabsch et al. provided a comprehensive review of the crucial role of TGF-β in the invasion and metastasis of BC [39].
In conclusion, our study offers a comprehensive overview of the intricate interplay among genetic, immune, and therapeutic elements in breast cancer. This integrated approach enhances our understanding of breast cancer and paves the way for more effective and personalized treatments.
Conclusion
This study has developed and validated a prognostic model for the accurate stratification of BC samples using bioinformatics algorithms and biological experiments. The model predicts prognosis, immunity, and drug sensitivity in drug-resistant BC. It can serve as an independent prognostic factor, enhancing the comprehension of BC treatment. Furthermore, the prognostic genes identified can be utilized as a reference for the precise treatment of BC.
Data availability
The data of the paper were downloaded from the TCGA database (https://portal.gdc.cancer.gov/) and the GEO database (https://ncbi.nlm.nih.gov/geo/).
References
Harbeck N, Penault-Llorca F, Cortes J, Gnant M, Houssami N, Poortmans P, Ruddy K, Tsang J, Cardoso F. Breast cancer. Nat Rev Dis Primers. 2019;5:66.
Norton L. High-dose chemotherapy for breast cancer: “how do you know?” J Clin Oncol. 2001;19:2769–70.
Gote V, Nookala AR, Bolla PK, Pal D. Drug resistance in metastatic breast cancer: tumor targeted nanomedicine to the rescue. Int J Mol Sci. 2021;22:4673.
Koual M, Tomkiewicz C, Cano-Sancho G, Antignac JP, Bats AS, Coumoul X. Environmental chemicals, breast cancer progression and drug resistance. Environ Health. 2020;19:117.
Sun Z, Shi Y, Shen Y, Cao L, Zhang W, Guan X. Analysis of different HER-2 mutations in breast cancer progression and drug resistance. J Cell Mol Med. 2015;19:2691–701.
Herzog SK, Fuqua SAW. ESR1 mutations and therapeutic resistance in metastatic breast cancer: progress and remaining challenges. Br J Cancer. 2022;126:174–86.
**a W, Chen W, Ni C, Meng X, Wu J, Yang Q, Tang H, Yuan H, Fang S. Chemotherapy-induced exosomal circBACH1 promotes breast cancer resistance and stemness via miR-217/G3BP2 signaling pathway. Breast Cancer Res. 2023;25:85.
Sun C, Huang X, Li J, Fu Z, Hua Y, Zeng T, He Y, Duan N, Yang F, Liang Y, et al. Exosome-transmitted tRF-16-K8J7K1B promotes tamoxifen resistance by reducing drug-induced cell apoptosis in breast cancer. Cancers (Basel). 2023;15:899.
Pan BT, Johnstone RM. Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: selective externalization of the receptor. Cell. 1983;33:967–78.
Xue D, Zheng Y, Wen J, Han J, Tuo H, Liu Y, Peng Y. Role of chemokines in hepatocellular carcinoma (Review). Oncol Rep. 2021;45:809–23.
Zhou C, Wei W, Ma J, Yang Y, Liang L, Zhang Y, Wang Z, Chen X, Huang L, Wang W, et al. Cancer-secreted exosomal miR-1468-5p promotes tumor immune escape via the immunosuppressive reprogramming of lymphatic vessels. Mol Ther. 2021;29:1512–28.
Huang Y, Kanada M, Ye J, Deng Y, He Q, Lei Z, Chen Y, Li Y, Qin P, Zhang J, et al. Exosome-mediated remodeling of the tumor microenvironment: from local to distant intercellular communication. Cancer Lett. 2022;543: 215796.
Wang L, Wang B, Wen H, Mao J, Ren Y, Yang H. Exosomes: a rising star in breast cancer (Review). Oncol Rep. 2020;44:407–23.
Dong X, Bai X, Ni J, Zhang H, Duan W, Graham P, Li Y. Exosomes and breast cancer drug resistance. Cell Death Dis. 2020;11:987.
Du L, Li X, Zhen L, Chen W, Mu L, Zhang Y, Song A. Everolimus inhibits breast cancer cell growth through PI3K/AKT/mTOR signaling pathway. Mol Med Rep. 2018;17:7163–9.
Zheng J, Liu Q, Yang J, Ren Q, Cao W, Yang J, Yu Z, Yu F, Wu Y, Shi H, et al. Co-culture of apoptotic breast cancer cells with immature dendritic cells: a novel approach for DC-based vaccination in breast cancer. Braz J Med Biol Res. 2012;45:510–5.
López-Mateo I, Alonso-Merino E, Suarez-Cabrera C, Park JW, Cheng SY, Alemany S, Paramio JM, Aranda A. Thyroid hormone receptor β inhibits self-renewal capacity of breast cancer stem cells. Thyroid. 2020;30:116–32.
Park JY, Saeidi S, Kim EH, Kim DH, Na HK, Keum JS, Surh YJ. Heregulin-β1 activates NF-E2-related factor 2 and induces manganese superoxide dismutase expression in human breast cancer cells via protein kinase B and extracellular signal-regulated protein kinase signaling pathways. J Cancer Prev. 2021;26:54–63.
Bischoff P, Kornhuber M, Dunst S, Zell J, Fauler B, Mielke T, Taubenberger AV, Guck J, Oelgeschläger M, Schönfelder G. Estrogens determine adherens junction organization and E-cadherin clustering in breast cancer cells via amphiregulin. iScience. 2020;23:101683.
Choi S, Whitman MA, Shimpi AA, Sempertegui ND, Chiou AE, Druso JE, Verma A, Lux SC, Cheng Z, Paszek M, et al. Bone-matrix mineralization dampens integrin-mediated mechanosignalling and metastatic progression in breast cancer. Nat Biomed Eng. 2023;7:1455–72.
Wang G, Zhang M, Meng P, Long C, Luo X, Yang X, Wang Y, Zhang Z, Mwangi J, Kamau PM, et al. Anticarin-β shows a promising anti-osteosarcoma effect by specifically inhibiting CCT4 to impair proteostasis. Acta Pharm Sin B. 2022;12:2268–79.
Zhang Y, Chen H, Mo H, Hu X, Gao R, Zhao Y, Liu B, Niu L, Sun X, Yu X, et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell. 2021;39:1578-1593.e1578.
Shen M, Smith HA, Wei Y, Jiang YZ, Zhao S, Wang N, Rowicki M, Tang Y, Hang X, Wu S, et al. Pharmacological disruption of the MTDH-SND1 complex enhances tumor antigen presentation and synergizes with anti-PD-1 therapy in metastatic breast cancer. Nat Cancer. 2022;3:60–74.
Feng F, Zhang J, Fan X, Yuan F, Jiang Y, Lv R, Ma Y. Downregulation of Rab27A contributes to metformin-induced suppression of breast cancer stem cells. Oncol Lett. 2017;14:2947–53.
Li J, Wang S, Wang N, Zheng Y, Yang B, Wang X, Zhang J, Pan B, Wang Z. Aiduqing formula inhibits breast cancer metastasis by suppressing TAM/CXCL1-induced Treg differentiation and infiltration. Cell Commun Signal. 2021;19:89.
Janakiram M, Abadi YM, Sparano JA, Zang X. T cell coinhibition and immunotherapy in human breast cancer. Discov Med. 2012;14:229–36.
Zheng G, Jia L, Yang AG. Roles of HLA-G/KIR2DL4 in breast cancer immune microenvironment. Front Immunol. 2022;13: 791975.
Woll MM, Fisher CM, Ryan GB, Gurney JM, Storrer CE, Ioannides CG, Shriver CD, Moul JW, McLeod DG, Ponniah S, et al. Direct measurement of peptide-specific CD8+ T cells using HLA-A2: Ig dimer for monitoring the in vivo immune response to a HER2/neu vaccine in breast and prostate cancer patients. J Clin Immunol. 2004;24:449–61.
Truax AD, Thakkar M, Greer SF. Dysregulated recruitment of the histone methyltransferase EZH2 to the class II transactivator (CIITA) promoter IV in breast cancer cells. PLoS ONE. 2012;7: e36013.
Song G, Wang X, Jia J, Yuan Y, Wan F, Zhou X, Yang H, Ren J, Gu J, Lyerly HK. Elevated level of peripheral CD8(+)CD28(−) T lymphocytes are an independent predictor of progression-free survival in patients with metastatic breast cancer during the course of chemotherapy. Cancer Immunol Immunother. 2013;62:1123–30.
Abdullah A, Akhand SS, Paez JSP, Brown W, Pan L, Libring S, Badamy M, Dykuizen E, Solorio L, Andy Tao W, et al. Epigenetic targeting of neuropilin-1 prevents bypass signaling in drug-resistant breast cancer. Oncogene. 2021;40:322–33.
Andreidesz K, Koszegi B, Kovacs D, Bagone Vantus V, Gallyas F, Kovacs K. Effect of oxaliplatin, olaparib and LY294002 in combination on triple-negative breast cancer cells. Int J Mol Sci. 2021;22:2056.
Abu Samaan TM, Samec M, Liskova A, Kubatka P, Büsselberg D. Paclitaxel’s mechanistic and clinical effects on breast cancer. Biomolecules. 2019;9:789.
Segovia-Mendoza M, González-González ME, Barrera D, Díaz L, García-Becerra R. Efficacy and mechanism of action of the tyrosine kinase inhibitors gefitinib, lapatinib and neratinib in the treatment of HER2-positive breast cancer: preclinical and clinical evidence. Am J Cancer Res. 2015;5:2531–61.
Arabkari V, Barua D, Hossain MM, Webber M, Smith T, Gupta A, Gupta S. miRNA-378 is downregulated by XBP1 and inhibits growth and migration of luminal breast cancer cells. Int J Mol Sci. 2023;25:186.
Joyce R, Pascual R, Heitink L, Capaldo BD, Vaillant F, Christie M, Tsai M, Surgenor E, Anttila CJA, Rajasekhar P, et al. Identification of aberrant luminal progenitors and mTORC1 as a potential breast cancer prevention target in BRCA2 mutation carriers. Nat Cell Biol. 2024;26:138–52.
Hickey TE, Selth LA, Chia KM, Laven-Law G, Milioli HH, Roden D, **dal S, Hui M, Finlay-Schultz J, Ebrahimie E, et al. The androgen receptor is a tumor suppressor in estrogen receptor-positive breast cancer. Nat Med. 2021;27:310–20.
Evans KW, Yuca E, Scott SS, Zhao M, Paez Arango N, Cruz Pico CX, Saridogan T, Shariati M, Class CA, Bristow CA, et al. Oxidative phosphorylation is a metabolic vulnerability in chemotherapy-resistant triple-negative breast cancer. Cancer Res. 2021;81:5572–81.
Drabsch Y, ten Dijke P. TGF-β signaling in breast cancer cell invasion and bone metastasis. J Mammary Gland Biol Neoplasia. 2011;16:97–108.
Acknowledgements
Not applicable.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
YL contributed to conceptualization, methodology, visualization, data analysis, writing—first draft preparation, writing—review and editing. LD (common 1) contributed to data analysis, software, visualization, writing review and editing. JM contributed to methodology, writing review and editing. LC contributed to software, writing—review and editing. LF (newsletter) contributed to research design, data management, review and revision, writing—review and editing. ZW (main newsletter) contributed to research design, financial support, data management, writing—review and editing. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interests
The author declares no potential conflicts of interest for the research, authorship, and/or publication of this article.
Ethics approval
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Y., Dong, L., Ma, J. et al. The prognostic genes model of breast cancer drug resistance based on single-cell sequencing analysis and transcriptome analysis. Clin Exp Med 24, 113 (2024). https://doi.org/10.1007/s10238-024-01372-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10238-024-01372-6