Abstract
Background
Somatic and germline variants are not distinguishable by circulating tumor DNA (ctDNA) testing without analyzing non-tumor samples. Although confirmatory germline testing is clinically relevant, the criteria for selecting presumed germline variants have not been established in ctDNA testing. In the present study, we aimed to evaluate the prevalence of pathogenic germline variants in clinical ctDNA testing through their variant allele fractions (VAFs).
Methods
A total of consecutive 106 patients with advanced solid tumors who underwent ctDNA testing (Guardant360®) between January 2018 and March 2020 were eligible for this study. To verify the origin of pathogenic variants reported in ctDNA testing, germline sequencing was performed using peripheral blood DNA samples archived in the Clinical Bioresource Center in Kyoto University Hospital (Kyoto, Japan) under clinical research settings.
Results
Among 223 pathogenic variants reported in ctDNA testing, the median VAF was 0.9% (0.02–81.8%), and 88 variants with ≥ 1% VAFs were analyzed in germline sequencing. Among 25 variants with ≥ 30% VAFs, seven were found in peripheral blood DNA (BRCA2: n = 6, JAK2: n = 1). In contrast, among the 63 variants with VAFs ranging from 1 to < 30%, only one variant was found in peripheral blood DNA (TP53: n = 1). Eventually, this variant with 15.6% VAF was defined to be an acquired variant, because its allelic distribution did not completely link to those of neighboring germline polymorphisms.
Conclusion
Our current study demonstrated that VAFs values are helpful for selecting presumed germline variants in clinical ctDNA testing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Next-generation sequencing (NGS)-based circulating tumor DNA (ctDNA) testing is an alternative method for comprehensive genomic profiling. This method is widely used in the clinical practice of cancer treatment because of its minimal invasiveness and no requirement for tissues. In ctDNA testing, small fractions of ctDNA can be detected among the total cell-free DNAs by incorporating a combination of molecular barcoding technology and bioinformatics methods [1]. Previous studies reported that the variant allele fractions (VAFs) of ctDNA range widely from 0.1 to ≥ 90%, although the median VAFs are less than 1% [2, 3]. VAFs reported in ctDNA testing are affected by several clinical variables, including the cancer type, stage, and overall tumor load [4–7].
Comprehensive genomic profiling sometimes results in secondary germline findings (SFs). The American College of Medical Genetics and Genomics (ACMG) recommended reporting SFs for several genes responsible for hereditary diseases [8–10]. However, in tumor-only testing and ctDNA testing, confirmatory germline testing is required, which uses non-tumor samples such as peripheral blood DNA. The European Society of Medical Oncology Precision Medicine Working Group recommended a threshold of ≥ 30% (or ≥ 20% for small insertions–deletions) for VAFs as an indication of presumed germline pathogenic variants in tumor-only tissue sequencing [11]. These recommendations are now widely used for managing SFs in tumor-only tissue sequencing in clinical practice. However, if the same threshold of ≥ 30% VAFs is feasible in clinical ctDNA testing is not well understood. Previous studies suggested that germline variants were presumable based on higher VAFs in ctDNA testing [12, 13], but the studies verifying this idea by germline testing are still limited.
In this present study, we performed germline sequencing to investigate the prevalence of true germline variants using peripheral blood DNA samples archived in the Clinical Bioresource Center in Kyoto University Hospital under clinical research settings.
Patients and methods
Patients
Consecutive 106 patients with unresectable advanced cancers who underwent Guardant360® liquid biopsy (Guardant Health, Redwood City, CA) in Kyoto University Hospital between January 2018 and March 2020 were eligible. Studies were approved by the Ethics Committee of the Kyoto University Graduate School of Medicine (Kyoto, Japan; G692 and G1223). All participants provided written informed consent to donate their blood samples to Clinical Bioresource Center in Kyoto University Hospital (Kyoto, Japan) for research use.
Confirmatory germline sequencing
According to the established recommendations for SFs [8, 9, 14], we selected 18 genes associated with hereditary cancer syndromes targeted by Guardant360® and designed a custom amplicon sequencing panel covering these 18 genes (Ion AmpliSeq On-Demand Panel, Thermo Fisher Scientific, MA, USA) (Supplementary Table S1). This panel covered the coding sequences and the splice sites of target genes. In addition, we also used the Ion AmpliSeq Cancer Hotspot Panel v2 (Thermo Fisher Scientific), which covered common pathogenic mutations in additional 29 cancer-related genes targeted by Guardant360® (Supplementary Table S1). Germline sequencing was performed in clinical research settings regardless of clinical indication.
Peripheral blood DNA was extracted using the Gene Prep Star NA-480 system (Kurabo Industries, Osaka, Japan) and archived in Clinical Bioresource Center in Kyoto University Hospital. To prepare library samples, the DNA was processed using the Ion AmpliSeq Library Kit Plus (Thermo Fisher Scientific). It was analyzed using an Ion S5 sequencing system equipped with Torrent Suite 5.10.1 (Thermo Fisher Scientific). Variants identified by Torrent Suite variantCaller were annotated using SnpEff tools. To avoid false-positive mutation calls, variant reads less than ten or VAFs less than 1.0% were excluded. Variants detected with VAFs between 1 and 30% in peripheral blood DNA were suspected to be somatically acquired variants that possibly include clonal hematopoiesis (CH) and other genetic mosaicisms rather than inherited germline variants [15, 16], and further information regarding their copy number and allelic distribution were obtained as follows. Copy-number alterations were analyzed using CovCopCan software (v.1.3.3) [17] to evaluate possible effects on VAFs. In addition, the distribution of variant allele was compared with those of heterozygous common single-nucleotide polymorphisms (SNPs) on the same sequencing reads using the Integrative Genomics Viewer (Broad Institute, MA, USA). Acquired variants were defined if their allelic distribution did not completely link to those of neighboring SNPs.
Definition of variant pathogenicity
To define variant pathogenicity, we first referred to ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/). If variants were registered as “pathogenic” or “likely-pathogenic” in ClinVar, they were classified into pathogenic variants. The variants registered as “conflicting interpretations of pathogenicity” were also classified as pathogenic if they demonstrated at least one pathogenic or likely-pathogenic report in ClinVar. For frameshift or truncation variants with no registration in ClinVar, their pathogenicity was considered by the annotations of neighboring truncated variants. If variants exhibited no information in ClinVar, their pathogenicity was evaluated based on the ACMG and the Association for Molecular Pathology guidelines [18].
Results
Patient characteristics
The characteristics of patients who underwent ctDNA testing are summarized in Table 1. The patients’ median age was 64 years (28–82), and 57 patients (53.8%) were male. The most common type of cancer was pancreatic (n = 37), followed by colorectal (n = 12), and lung (n = 11). Twenty patients (18.9%) presented with a family history of cancer in both first-degree and second-degree relatives.
Distribution of pathogenic variant allele fractions in ctDNA testing
In total, 409 short variants were reported in 106 patients, and 224 of those variants were classified as pathogenic. The median VAF of pathogenic variants was 0.9% (0.02–81.8%). The number of pathogenic variants with ≥ 1%, ≥ 5%, and ≥ 30% VAFs was 111, 73, and 27, respectively (Fig. 1A). Among 27 variants with ≥ 30% VAFs (high VAFs), TP53 variants were the most common (n = 10), followed by BRCA2 (n = 6), and then KRAS (n = 4) (Fig. 1B). Similarly, among 84 pathogenic variants with moderate VAFs ranging from 1 to < 30%, TP53 variants were most common (n = 28), followed by KRAS (n = 17), APC (n = 7), SMAD4 (n = 5), and PIK3CA (n = 5) (Fig. 1C).
A The distribution of variant allele fractions of pathogenic variants found in ctDNA testing. Two hundred and twenty-four pathogenic variants detected in ctDNA testing are arranged by their variant allele fractions (VAFs) from highest to lowest. B The number of pathogenic variants with 30% or higher VAFs arranged by gene in ctDNA testing (n = 27). C The number of pathogenic variants with VAFs ranged from 1 to < 30% by gene in ctDNA testing (n = 84). Other includes one variant in BRAF, BRCA2, CDKN2A, CCNB1, ERBB2, FBXW7, FGFR2, NFE2L2, and PTEN
Results of germline sequencing
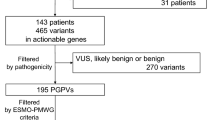
To confirm the origin of pathogenic variants, germline sequencing was performed using archived DNA from 44 patients whose ctDNA testing indicated pathogenic variants with ≥ 1% VAFs (Fig. 2). As a result, six BRCA2 variants, which VAFs ranged from 43.7 to 60.0% in ctDNA testing, were confirmed to be germline in patients with pancreatic (n = 2), breast (n = 2), ovarian (n = 1), and prostate cancers (n = 1) (Fig. 3 and Table 2). In addition, germline sequencing illustrated possible acquired variants in TP53 and JAK2 genes (Fig. 3 and Table 2). VAFs of the TP53 C238Y variant were 14.0 and 15.6% in ctDNA testing and germline sequencing, respectively (Table 2). In germline sequencing, no copy-number abnormality was detected in the TP53 locus (Supplementary Figure S1). Further analysis of allelic read distribution in germline sequencing revealed that this variant was found only in a fraction of sequencing reads originating from haploid genome with two neighboring heterozygous SNPs (rs12951053, rs12947788) (Fig. 4). These results indicate that this variant was likely to be an acquired variant rather than a germline variant. VAFs of the JAK2 V617F variant, commonly found in myeloproliferative neoplasm [19], were 37.7 and 46.1% in ctDNA testing and germline sequencing, respectively (Table 2). Since no heterozygous SNPs existed near this variant to compare the allelic distributions, we could not further clarify the origin of this variant.
The distributions of the TP53 C238Y variant and its common SNPs on the germline DNA sequencing reads. The TP53 C238Y variant was detected in a fraction of sequencing reads originating from haploid genome harboring two neighboring heterozygous SNPs (rs12951053 and rs12947788) in the germline DNA sequencing
Discussion
Since VAFs detected in ctDNA testing tend to be much lower (median VAF < 1%) than those in tissue-based testing [2, 3], variants with higher VAFs around 50% are expected to be of germline rather than somatic origin. The present study performed germline sequencing using peripheral blood DNA samples for pathogenic variants with ≥ 1% VAFs reported in ctDNA testing. Although peripheral blood DNA analysis found seven pathogenic variants out of 25 with ≥ 30% VAFs, only six were confirmed to be true pathogenic germline variants. In contrast, out of the 63 variants with VAFs ranging from 1 to < 30%, there were no true pathogenic germline variants. These results support the idea that a threshold of VAFs ≥ 30% is feasible to select presumed germline pathogenic variants, which require confirmatory germline testing in clinical ctDNA testing. A recent study reported that 89% of 36 confirmed germline variants exhibited ctDNA VAFs between 40 and 60%; the remaining 11% were out of this range [20]. Their report is consistent with our results; however, research also suggests that using a tight threshold of the ctDNA VAF range to screen germline variants can lead to missing true germline variants.
In the present study, we confirmed the presence of true pathogenic germline variants only in BRCA2, which could possibly be due to the following reasons: First, BRCA2 was the most common cancer susceptibility gene in patients with advanced cancers who undergo universal germline sequencing [21]. Second, ctDNA panel (Guardant360®) utilized in this study did not cover several of the common cancer susceptibility genes, such as CHEK2, MSH2, MSH6, and PMS2. Third, the sample size was small, and over 50% of the patients had one of the top three cancer types (pancreatic, colorectal, and lung).
Variants of CH can contaminate the results of the ctDNA testing [22, 23]. CH is commonly observed in aged people and patients undergoing anticancer therapies [24, 25]. CH variants are found in various hereditary cancer-associated genes such as TP53 and ATM [15, 23]. The VAFs for the TP53 C238Y variant detected in this study were relatively low in ctDNA and peripheral blood DNA (14.0 and 15.6%, respectively), suggesting that this variant is not likely to be of true germline origin. Supporting this idea, the family history and clinical course of the patient did not meet the criteria for the Li-Fraumeni syndrome. Further analysis of allelic read distribution compared to common SNPs located nearby the TP53 C238Y variant was helpful to confirm it is an acquired genetic variation. The JAK2 V617F variant was detected in another patient with 46.1% VAFs in germline sequencing. This variant is a well-known somatic variant hotspot in myeloproliferative neoplasms and CH [19, 26], and this patient demonstrated a clinical manifestation of thrombocytosis. Although we could not obtain further information to identify its origin by germline sequencing, according to the patient’s phenotype, familial history, and known properties of this variant, we presumed that this was likely to be an acquired variant rather than a germline one. The recent publication of the ACMG statements suggested that analyzing additional control samples from other normal tissues from the patients or their family members would be helpful to distinguish the origin of variants [15].
This study exhibits the other limitation. Because little information was available about copy-number alterations from ctDNA testing, their effects on VAFs remain unknown in our germline analysis. Detailed information about copy-number alterations can improve the presumption accuracy of germline variants in ctDNA testing, as shown in previous reports [12, 27].
In conclusion, our current study demonstrated that VAFs information helps select putative germline variants in clinical ctDNA testing.
References
Lanman RB, Mortimer SA, Zill OA et al (2015) Analytical and clinical validation of a digital Sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PLoS ONE 10(10):e0140712. https://doi.org/10.1371/journal.pone.0140712
Odegaard JI, Vincent JJ, Mortimer S et al (2018) Validation of a plasma-based comprehensive cancer genoty** assay utilizing orthogonal tissue- and plasma-based methodologies. Clin Cancer Res 24(15):3539–3549. https://doi.org/10.1158/1078-0432.ccr-17-3831
Mack PC, Banks KC, Espenschied CR et al (2020) Spectrum of driver mutations and clinical impact of circulating tumor DNA analysis in non-small cell lung cancer: analysis of over 8000 cases. Cancer 126(14):3219–3228. https://doi.org/10.1002/cncr.32876
Phallen J, Sausen M, Adleff V et al (2017) Direct detection of early-stage cancers using circulating tumor DNA. Sci Transl. https://doi.org/10.1126/scitranslmed.aan2415
Chaudhuri AA, Chabon JJ, Lovejoy AF et al (2017) Early detection of molecular residual disease in localized lung cancer by circulating tumor DNA profiling. Cancer Discov 7(12):1394–1403. https://doi.org/10.1158/2159-8290.cd-17-0716
Newman AM, Bratman SV, To J et al (2014) An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med 20(5):548–554. https://doi.org/10.1038/nm.3519
Osumi H, Shinozaki E, Takeda Y et al (2019) Clinical relevance of circulating tumor DNA assessed through deep sequencing in patients with metastatic colorectal cancer. Cancer Med 8(1):408–417. https://doi.org/10.1002/cam4.1913
Green RC, Berg JS, Grody WW et al (2013) ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med 15(7):565–574. https://doi.org/10.1038/gim.2013.73
Kalia SS, Adelman K, Bale SJ et al (2017) Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): a policy statement of the American College of Medical Genetics and Genomics. Genet Med 19(2):249–255. https://doi.org/10.1038/gim.2016.190
Miller DT, Lee K, Chung WK et al (2021) ACMG SF v3.0 list for reporting of secondary findings in clinical exome and genome sequencing: a policy statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med 23(8):1381–1390. https://doi.org/10.1038/s41436-021-01172-3
Mandelker D, Donoghue MTA, Talukdar S et al (2019) Germline-focused analysis of tumour-only sequencing: recommendations from the ESMO precision medicine working group. Ann Oncol. https://doi.org/10.1093/annonc/mdz136
Slavin TP, Banks KC, Chudova D et al (2018) Identification of incidental germline mutations in patients with advanced solid tumors who underwent cell-free circulating tumor DNA sequencing. J Clin Oncol. https://doi.org/10.1200/jco.18.00328
Nakamura Y, Taniguchi H, Ikeda M et al (2020) Clinical utility of circulating tumor DNA sequencing in advanced gastrointestinal cancer: SCRUM-Japan GI-Screen and GOZILA studies. Nat Med 26(12):1859–1864. https://doi.org/10.1038/s41591-020-1063-5
Pujol P, Vande Perre P, Faivre L et al (2018) Guidelines for reporting secondary findings of genome sequencing in cancer genes: the SFMPP recommendations. Eur J Hum Genet. https://doi.org/10.1038/s41431-018-0224-1
Chao EC, Astbury C, Deignan JL et al (2021) Incidental detection of acquired variants in germline genetic and genomic testing: a points to consider statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. https://doi.org/10.1038/s41436-021-01138-5
Campbell IM, Shaw CA, Stankiewicz P et al (2015) Somatic mosaicism: implications for disease and transmission genetics. Trends Genet 31(7):382–392. https://doi.org/10.1016/j.tig.2015.03.013
Derouault P, Chauzeix J, Rizzo D et al (2020) CovCopCan: an efficient tool to detect copy number variation from amplicon sequencing data in inherited diseases and cancer. PLoS Comput Biol 16(2):e1007503. https://doi.org/10.1371/journal.pcbi.1007503
Richards S, Aziz N, Bale S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17(5):405–424. https://doi.org/10.1038/gim.2015.30
Baxter EJ, Scott LM, Campbell PJ et al (2005) Acquired mutation of the tyrosine kinase JAK2 in human myeloproliferative disorders. Lancet 365(9464):1054–1061. https://doi.org/10.1016/s0140-6736(05)71142-9
Stout LA, Kassem N, Hunter C et al (2021) Identification of germline cancer predisposition variants during clinical ctDNA testing. Sci Rep 11(1):13624. https://doi.org/10.1038/s41598-021-93084-0
Mandelker D, Zhang L, Kemel Y et al (2017) Mutation detection in patients With advanced cancer by universal sequencing of cancer-related genes in tumor and normal DNA vs guideline-based germline testing. JAMA 318(9):825–835. https://doi.org/10.1001/jama.2017.11137
Mayrhofer M, De Laere B, Whitington T et al (2018) Cell-free DNA profiling of metastatic prostate cancer reveals microsatellite instability, structural rearrangements and clonal hematopoiesis. Genome Med 10(1):85. https://doi.org/10.1186/s13073-018-0595-5
Razavi P, Li BT, Brown DN et al (2019) High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat Med 25(12):1928–1937. https://doi.org/10.1038/s41591-019-0652-7
**e M, Lu C, Wang J et al (2014) Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med 20(12):1472–1478. https://doi.org/10.1038/nm.3733
Suehara Y, Sakata-Yanagimoto M, Hattori K et al (2019) Mutations found in cell-free DNAs of patients with malignant lymphoma at remission can derive from clonal hematopoiesis. Cancer Sci 110(10):3375–3381. https://doi.org/10.1111/cas.14176
McKerrell T, Park N, Chi J et al (2017) JAK2 V617F hematopoietic clones are present several years prior to MPN diagnosis and follow different expansion kinetics. Blood Adv 1(14):968–971. https://doi.org/10.1182/bloodadvances.2017007047
Hu Y, Alden RS, Odegaard JI et al (2017) Discrimination of germline EGFR T790M mutations in plasma cell-free DNA allows study of prevalence across 31,414 cancer patients. Clin Cancer Res 23(23):7351–7359. https://doi.org/10.1158/1078-0432.ccr-17-1745
Acknowledgements
The authors thank all the patients who participated in this study; Mari Funakoshi, Hitomi Sakamoto, Kanami Ashida, Junko Suga, Momoko Sato, Kumi Mukai, and all staff members of the Clinical Bioresource Center in Kyoto University Hospital for their excellent technical and secretarial assistance.
Funding
This study was supported by the Japan Agency for Medical Research and Development, AMED, under Grant Number 17kk0305006h0001.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Masashi Kanai received lecture fees, honoraria, and other fees from Chugai Pharmaceutical Co., Ltd.; research funding from Molecular Health; stock ownership from Therabiopharma Inc. Manabu Muto received research funding from Chugai Pharmaceutical Co., Ltd.; research funding from Sysmex Corporation. All remaining authors have declared no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Yamamoto, Y., Fukuyama, K., Kanai, M. et al. Prevalence of pathogenic germline variants in the circulating tumor DNA testing. Int J Clin Oncol 27, 1554–1561 (2022). https://doi.org/10.1007/s10147-022-02220-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-022-02220-x