Abstract
Pancreatic ductal adenocarcinoma (PDAC) is one of the most aggressive and lethal cancers, and develo** an efficient and reliable approach for its early-stage diagnosis is urgently needed. Precancerous lesions of PDAC, such as pancreatic intraepithelial neoplasia (PanIN) and intraductal papillary mucinous neoplasms (IPMN), arise through multiple steps of driver gene alterations in KRAS, TP53, CDKN2A, SMAD4, or GNAS. Hallmark mutations play a role in tumor initiation and progression, and their detection in bodily fluids is crucial for diagnosis. Recently, liquid biopsy has gained attention as an approach to complement pathological diagnosis, and in addition to mutation signatures in cell-free DNA, cell-free RNA, and extracellular vesicles have been investigated as potential diagnostic and prognostic markers. Integrating such molecular information to revise the diagnostic criteria for pancreatic cancer can enable a better understanding of the pathogenesis underlying inter-patient heterogeneity, such as sensitivity to chemotherapy and disease outcomes. This review discusses the current diagnostic approaches and clinical applications of genetic analysis in pancreatic cancer and diagnostic attempts by liquid biopsy and molecular analyses using pancreatic juice, duodenal fluid, and blood samples. Emerging knowledge in the rapidly advancing liquid biopsy field is promising for molecular profiling and diagnosing pancreatic diseases with significant diversity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the most dismal malignancy, with a 5-year survival rate of approximately 10% [1], as only < 25% of PDAC are localized and potentially curable during the first diagnostic phase [2, 3]. Owing to the lack of early-stage diagnosis, most patients with PDAC present with locally advanced or metastatic cancers [4]. Surgical resection that results in a better prognosis than other therapies [5] is an option; however, over 80% of PDAC cases are advanced owing to constant invasion and distant metastasis [6, 7]. FOLFIRINOX (fluorouracil, leucovorin, irinotecan, and oxaliplatin) and gemcitabine plus nanoparticle albumin-bound paclitaxel (GnP) are chemotherapeutic agents used to treat unresectable PDAC. However, their effectiveness is still unsatisfactory and limited [5].
For the early diagnosis of PDAC, imaging studies performed for screening or symptomatic examination should accurately detect pancreatic tumors, main pancreatic duct dilatation, and cystic changes [8]. Abdominal ultrasound is simple but does not have a high tumor detection rate; contrast agents have been reported to improve detection in recent years [9]. Computed tomography (CT) is the most widely used modality in diagnosing the presence and extent of PDAC [10]. However, to avoid radiation-associated malignancies in BRCA mutation carriers [11], follow-up is usually preferred using magnetic resonance imaging (MRI). To date, there is no clear evidence of a link between medical radiation exposure and accelerated pancreatic carcinogenesis; however, the long-term effects of frequent dynamic scanning need to be carefully evaluated.
Endoscopic ultrasonography (EUS) was developed in the early 1980s, and EUS-guided tissue acquisition (TA) was first reported in 1992 [12]. Since then, EUS has become increasingly widespread. EUS achieves the best detection performance [13]. However, it differs from other endoscopic procedures in its operation, making it challenging to train physicians and limiting its applicability as its diagnostic performance is influenced by the examiner's skill [14]. Attempts to strengthen cooperation among medical institutions are also being made to utilize these examination modalities accurately. [15, 16]. Significant efforts have been made to identify the etiology of carcinogenesis and its risk factors. However, strategies that integrate this knowledge into clinically feasible scoring are yet to be established [17]. Recently, scattered reports on using specific biomarkers for the minimally invasive identification of high-risk PDAC groups are available. However, to date, no effective biomarker for early detection has been established, despite an understanding of the mechanism of pancreatic carcinogenesis.
A liquid biopsy system using body fluids to detect reliable biomarkers is crucial for the early diagnosis of PDAC to improve diagnostic applications. Carbohydrate antigen 19–9 (CA19-9) and carcinoembryonic antigen (CEA) are commonly used tumor markers for PDAC. CA19-9 has the highest sensitivity at 70–80% and a < 50% specificity for PDAC diagnosis. Moreover, the sensitivity of CA19-9 for stage I PDAC is 55.6% [18]. Notably, several studies have reported the usefulness of CA19-9 as an anchor marker for PDAC [19, 20]; however, its utility is still restricted, particularly in Lewis antigen-negative patients characterized by inadequate secretion of CA19-9 and fucosylation deficiency [21].
In contrast, many clinical trials have been conducted to identify new biomarkers using liquid biopsy tools, such as cfDNA, cell-free RNA (cfRNA), proteins, extracellular vesicles (EVs) including exosomes, and circulating tumor cells (CTCs) [22, 23]. For example, the cell surface proteoglycan glypican-1 (GPC1) is present in PDAC cell-derived exosomes, and circulating exosome GPC1 in serum has been reported to serve as a biomarker for detecting early stages of PDAC [24]. Moreover, several miRNAs or exosomal miRNAs are highly expressed in body fluids such as blood, duodenal fluid (DF), and pancreatic juice (PJ) [25,26,27]. We previously encountered a case of early-stage PDAC in which KRAS mutations were detected in PJ within the resected PDAC tissue but not in the plasma, suggesting that PJ is superior to blood for PDAC diagnosis as it may reflect DNA mutations or abnormal RNA expression from the original tumor more precisely owing to the short distance between the primary lesion and the position from where PJ is collected [28]. DF can also be a valuable sample collected using a minimally invasive method compared with PJ collection using a catheter. Nevertheless, there is a need to discover and validate new markers in PJ and DF for their utility for liquid biopsy to enable the early diagnosis of PDAC and increase the opportunities for curative surgery.
This review summarizes the gene alterations associated with initiating PDAC precursor lesions, such as intraepithelial neoplasia (PanIN) and intraductal papillary mucinous neoplasms (IPMN). It discusses the molecular subtypes of invasive PDAC. We also discuss literature and trials on the early diagnosis of PDAC. Finally, we summarize the key themes and recent progress in liquid biopsy using body fluids, mainly PJ and DF, to analyze oncogenic mutations, RNA expression, and other factors, focusing on high-grade IPMN and PDAC.
Precancerous lesions and gene alterations
Pancreatic intraepithelial neoplasia (PanIN) and gene alteration
PanIN is the most common precursor lesion for PDAC. Oncogenic Kras can induce PanIN in mice [29], and humans with PDAC harbor KRAS mutations. Tumor development is initiated by KRAS mutation at the earliest stages of pancreatic tumorigenesis [30]. Mutations are mainly detected in codon 12, while during the initial stage of pancreatic carcinogenesis, mutations are occasionally identified in codons 13 and 61 [31]. Approximately 70–80% of PDAC patients harbor G12V, G12D, or G12R KRAS mutations [32, 33]. KRAS G12D mutation is the most frequent and is a predictive factor for worse PDAC prognosis [34]. A metastatic mouse PDAC model with KrasG12D revealed that Kras mutant allele-specific imbalances, such as copy number gain or loss of heterozygosity, were observed even in human PanIN lesions and invasive PDAC (Fig. 1) [35].
Diagnostic material/information during pancreatic carcinogenesis. Oncogenic KRAS can initiate precursor lesions (dysplasia; see Fig. 2 for details), and allelic imbalance increases oncogenic dosage gain and tumor progression. In addition to circulating tumor cells (CTCs), mutations and methylations in circulating cell-free DNA (cfDNA), cell-free RNA (cfRNA), and extracellular vesicles (EVs) can serve as liquid biopsy targets. A multi-layer detection system covering these factors is necessary for early diagnosis, complementary to conventional pathological assessments (e.g., SPACE and FNA). In addition to blood, where tumor-derived factors are significantly diluted, the collection of DF and PJ is expected to provide an opportunity to capture such molecules. Such genomic information can be integrated with traditional diagnostic modalities, improving the accuracy of early detection of human pancreatic cancer with high accuracy
Following tumor initiation by KRAS mutation, PanIN lesions progress to invasive PDAC through subsequent inactivation of the tumor suppressor genes TP53, cyclin-dependent kinase inhibitor 2A (CDKN2A), and SMAD family member 4 (SMAD4) [36]. Mutations in several genes have been associated with PDAC; however, these four genes are the primary driver genes for human PDAC [37].
IPMN and gene alteration
IPMN has been associated with GNAS mutation at codon 201 in 40–70%, while KRAS mutations in 40–65% of cases [36]. GNAS gene is located on chromosome 20q13.32 and encodes the G-protein alpha-subunit [G(s)α] of heterotrimeric G-protein-coupled receptors. Activation of protein kinase A (PKA) through stimulation of the messenger cAMP by the ligand induces the phosphorylation of cAMP-responsive element-binding protein (CREB) and extracellular signal-regulated kinase (ERK). In several tissues, GNAS–cAMP signaling maintains cellular differentiation and quiescence [38,39,60]. PDAC can be classified into two common molecular subtypes: basal like and classical. The basal-like subtype is characterized by a more aggressive phenotype and poor patient survival, whereas the classical subtype is highly sensitive to chemotherapeutic agents. Therefore, knowing which subtype is involved can help guide chemotherapy decisions [61,62,63]. Phenotype classification by RNA signatures may not be clinically relevant because of the high cost and difficulty in analyzing heterogeneous tumors. However, immunohistochemical analysis has several advantages in overcoming this issue; GATA6, CK5, and Vimentin may serve as relevant markers for defining the differential expression profile of heterogeneous tumors [64]. Like GATA6, GATA4 maintains the classical phenotype in cooperation with GATA6 [65].
Flowers et al. suggested that ductal cell-derived tumor signatures are associated with the basal-like subtype. However, acinar cell-derived tumor signatures are correlated with the classical subtype of human PDAC, using a gene set from GEMM in the context of oncogenic Kras and Tp53. They also suggested that specific genetic events, such as mutations in TP53 and KDM6A, may be associated with particular subtypes [66]. There is some confusion, however, regarding IPMN-related tumors. As IPMN likely originates from ductal cells, this would suggest that it is correlated with the more aggressive basal-like subtype of PDAC. Yet Collisson et al. have identified IPMN-related tumors as belonging to the classical subtype, which Flowers et al. associate with acinar cells related to PanIN [60]. This raises a discrepancy between these two theories: cell of origin and molecular subtypes (Fig. 2). Moreover, in IPMN, driver gene mutations, such as GNAS, STK11, RNF43, or KLF4, may also determine the molecular subtype and its transition [32, 67]. Therefore, the regulation of PDAC subtypes and their potential interaction with cancer cell origin must be further investigated in early-stage tumors to develop novel clinical applications for screening and surveillance.
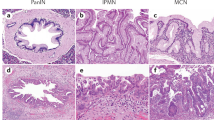
Cell of origin and molecular subtypes in pancreatic cancer. The pancreas contains several specialized cell types that can alter cellular identity in response to inflammation and subsequent regeneration/repair. Pancreatic acinar cells can undergo acinar-to-ductal metaplasia (ADM), a reprogramming event that induces transdifferentiation into a duct-like phenotype. Pancreatic intraepithelial neoplasia (PanIN)-like lesions can be generated in mice through the expression of oncogenic Kras in acinar cells; intraductal papillary mucinous neoplasms (IPMN) can arise from acinar cells in association with mutant Kras and Gnas or from ductal cells in the context of association with chromatin remodeling or transforming growth factor β signaling mutations. The subsequent inactivation of tumor suppressors accelerates the progression of precursor lesions, leading to invasive ductal adenocarcinoma. Molecular subty** approaches for pancreatic cancer based on transcriptomic data can help stratify patients in clinical practice, leading to optimized treatment algorithms. The recently identified classical and basal-like molecular subtypes of pancreatic cancer affect patient survival and can be profiled by the differential immunohistochemical expression of GATA6, CK5, and Vimentin. Phenotypic transitions can be observed during chemotherapy and even during the natural history of tumor progression via genetic and epigenetic alterations. It remains to be determined if mutant KRAS dosage and chromothripsis regulate PDAC development. Recent mouse and human studies have supported the significant role of the cell of origin and the associated transcriptomes in influencing PDAC phenotype
Body fluids transcriptomes can provide clinically relevant information to explore the origin of cancer and the roots of PDAC subtypes. Larson et al. reported that plasma cell-free RNAs (cfRNAs) in plasma help detect cancer tissue origin and subtypes [68]. The circulating cell-free genome atlas (NCT02889978) contains transcriptome data of blood samples from cancer and non-cancer patients. To date, no registry has collected mutation profiles combined with transcriptome analysis using DF or PJ, and we are currently investigating the states of driver mutations and transcriptomes in DF and PJ in collaboration with affiliated hospitals to establish a catalog for identifying PDAC subtypes and detecting the early stages of tumors more effectively than blood testing.
A diagnostic approach for early-stage PDAC
Endoscopic modalities for pathological assessment
The pathological diagnosis of PDAC is mandatory for patients with unresectable PDAC, and its significance has increased with advances in matched therapies following molecular profiling [69]. There has been an increase in the use of preoperative chemotherapy and a growing consensus that pathological diagnosis is crucial in cases where surgery is an option [70, 71].
EUS-TA, with a sensitivity of 84–91%, is the most widely performed tissue sampling method for pathological diagnosis [72, 73]. EUS-TA has progressed with a focus on puncture needles, such as forward facing, Franseen, and fork tips [74,75,76,77], and specimen collection and processing methods, such as rapid on-site evaluation and target sample check illuminator [78, 79]. Tissues obtained by EUS-FNA are occasionally small, and a diagnosis may not be made in samples containing a large amount of blood. Some reports have suggested that increased punctures improve diagnostic performance [80].
An unusual complication associated with EUS-TA is needle tract seeding. Kitano et al. reported that transgastric but not transduodenal EUS-TA causes seeding in approximately 0.9% of cases [81]. Therefore, careful monitoring is required following the resection of PDACs diagnosed using EUS-TA. In addition, the procedure should be avoided in cases where it is difficult to suspend antithrombotic drugs, where it is challenging to delineate the mass by EUS, or where the presence of a blood vessel or pancreatic duct in the puncture route is unavoidable.
Pancreatic juice cytology (PJC), with a sensitivity of 47–76%, is an alternative to EUS-TA [82]. Notably, several methods, including brushing PJC, washing PJC, endoscopic nasopancreatic drainage (ENPD) with serial pancreatic juice aspiration cytologic examination (SPACE) [83], and secretin-loaded PJC (S-PJC), have been attempted to improve the diagnostic performance of PJC [84]. However, all methods are associated with the risk of post-ERCP pancreatitis as a severe complication and are still considered alternatives to EUS-TA. Iiboshi et al. reported the usefulness of multiple consecutive PJC examinations with ENPD [83]. Sensitivity, specificity, and overall accuracy were 100%, 83.3%, and 95%, respectively, with an average of 5.3 mL PJ sampling. They also reported that SPACE is particularly useful for diagnosing intraepithelial carcinomas that cannot be delineated using EUS. Since then, there have been increasing reports on diagnosing intraepithelial pancreatic cancer using SPACE, mainly in Japan [85,86,87,88].
Until recently, S-PJC had been performed widely using porcine secretin [84, 89]; however, animal-derived products have been discontinued due to safety issues. Synthetic human secretin is now available as a secretin preparation. In the United States, synthetic secretin is licensed for use in (i) the exocrine pancreas function test, (ii) the Zollinger-Ellison Syndrome Test, and (iii) stimulation of pancreatic secretions to facilitate the identification of the ampulla of Vater and accessory papilla during ERCP. However, secretin has not been approved for clinical use in some countries, including Japan. The unstable supply of secretin is another issue. Mass production, distribution, and cost reductions are expected as its clinical utility becomes clearer. Other methods include (1) bile cytology in cases of pancreatic head cancer invading the distal bile duct [89], (2) ascites cytology in cases of peritoneal dissemination with exudative ascites effusion, and (3) tumor biopsy under transabdominal ultrasound or CT in institutions where EUS-TA is not feasible [90].
Clinical application of genetic testing
A highly sensitive method for detecting even minor changes in crucial genes is required, particularly in the early stages of cancer. Invariably, numerous techniques have been developed to quantify mutations in driver oncogenes and tumor suppressor genes. One method fundamental in mutation analyses is polymerase chain reaction (PCR). Real-time PCR for mutation-specific amplification has been widely validated with high sensitivity and reliability, allowing for the real-time detection and quantification of DNA in a comparatively short turnaround time.
Further technological developments aimed at improving detection sensitivity have led to the advent of technologies such as the BEAMing (Beads, Emulsion, Amplification, and Magnetics) method [91] and digital PCR (dPCR) [92]. dPCR is a highly sensitive method that performs endpoint PCR on more than 10,000 individual reaction wells (such as oil droplets or chambers), allowing for accurate detection and absolute quantification of DNA mutations without calibration reactions [93]. The dPCR assays achieved a shallow variant allele frequency (VAF) detection limit (approximately 20-fold; 0.05% vs. 1%). The multiplex droplet digital PCR assay was developed to target many genomic regions using a few primers and probes [94,95,96]. As tissue specimens are occasionally minimal, comprehensive analysis of genomic alterations in a single reaction is a powerful tool for clinical testing, potentially compensating for pathological diagnosis. The multiplex ddPCR assay can detect point mutations in cfDNA to minimize false positives while retaining sensitivity, making it feasible to analyze cfDNA samples [97].
Comprehensive genomic profiling (CGP) using next-generation sequencing (NGS) is the most powerful tool for identifying tumor-specific genomic alterations in clinical oncology, enabling precision medicine [98,99,100]. In addition to clinically available CGP assays such as FoundationOne CDx and OncoGuide, whole exome sequencing (WES) and whole genome sequencing (WGS) are used to evaluate to fine-tune cancer therapy for patients with unresectable diseases [101]. As a super-sensitive analysis protocol for cfDNA, unique molecular identifiers (UMIs) that are attached to unique DNA sequences in each original molecule of DNA are standard tools for overcoming PCR and sequencing errors to reliably quantitate low-frequent mutations at ≤ 0.1% VAFs [99, 100]. In addition to determining the best cancer therapy, the possibility of early PDAC screening through cfDNA diagnostics has been widely discussed, and many trials on cfDNA mutation profiling have been conducted [102]. Such a technique would be useful in stratifying high-risk individuals, such as those with a family history of PDA and pancreatic cysts, including IPMN. Furthermore, several other CGP panels for tissue and plasma cfDNA using genetic or epigenetic biomarkers, such as FoundationOne Liquid [103] and Guardant360, as well as the more recent CGP: GuardantOMNI and GuardantINFINITY [104], have also been developed. Currently, the last two panels are even more comprehensive for research use only and may eventually allow diagnosis at an earlier tumor stage [105].
Molecular diagnosis using body fluids
Advantage of the analysis using PJ or DF
Molecular diagnostic methods using body fluids have been developed in recent years, and we outlined the benefits and the challenges that need to be addressed in Table 1. Significant efforts have been made to detect mutations in plasma cfDNA in many cancer types early. Such detection may be a biomarker of tumor burden; however, mutation detection rates are lower in patients with PDAC than in those with other neoplasms, primarily because of low tumor cellularity [106]. Even in metastatic PDAC, a previous study demonstrated that the accuracy only reached around 40% [107]. Considering the low yield of circulating plasma cfDNA, specifically in patients with small PDAC, detecting tumor-derived DNA remains challenging [108]. Therefore, alternative approaches must be considered to overcome this limitation.
Analysis of blood cfDNA from patients with pancreatic neoplasms has been performed using high-sensitivity dPCR technology [109]. dPCR detects KRAS codon12 mutations even in pancreatic cancer cohorts with many early-stage patients [110]. We developed a dPCR method to overcome subsampling errors, an issue in testing for extremely low copy number mutations [111]. Using the pre-amplification method, tumor-derived mutant KRAS in the plasma of patients with resectable PDAC was accurately detected (AUC, 0.861–0.876), and the dPCR method improved post-resection recurrence prediction over that of the marker CA19-9 [26]. DNA in blood exosomes (exoDNA) is protected from degradation and fragmentation and can be extracted as high molecular weight DNA compared with cfDNA. It has been reported that exoDNA levels increase after neoadjuvant therapy; exoDNA also yields a higher detection rate of KRAS mutations with dPCR and is more amenable as a template for CGP [112].
In gastrointestinal organs, secreted digestive juices may enrich genetic abnormalities associated with cancerous lesions. Therefore, liquid biopsy using body fluids could resolve the limitations of mutation detection in plasma cfDNA. Initially, KRAS mutations were targeted using PJ collected from patients with PDAC/IPMN for PJC and could complement the less-sensitive cytology evaluation and help in follow-up [113,114,115]. Using high-resolution melt curve analysis and pyrosequencing methods [31], mutations in KRAS are detected in PJ from PDAC patients and in 50% of asymptomatic individuals at high risk for neoplasms [116]. KRAS mutations have also been observed in individuals without pancreatic abnormalities, likely because of invisible PanIN lesions. TP53 mutations have also been detected in PJ from patients with HGD [31].
Sequencing technologies enable comprehensive cancer genome profiling using PJ and efficient detection of tumor suppressor mutations, including TP53, KRAS, and GNAS mutations, serving as a tool to stratify tumor grades in patients with IPMN [117, 118]. Yu et al. developed a digital NGS method in which many aliquots of DNA from each patient’s fluid were individually subjected to sequencing, and a mutation score of one was given to each mutation-containing aliquot [119]. Canonical mutations during pancreatic carcinogenesis (e.g., KRAS, TP53, SMAD4, and CDKN2A), using this method, were efficiently detected in the PJ of PDAC/IPMN patients, proving useful in the evaluation of patients undergoing pancreatic surveillance [120].
Epigenetics, fragmentomics, and cfDNA topology are crucial for develo** DNA-based cancer screening [121]. Advanced library preparation was adapted to allow for the methylome profiling of samples with a low input of DNA (1–10 ng), potentially applicable to early cancer detection [122]. In addition to this technology, detailed informatics regarding pan-cancer methylome analysis offer a better classification of cancer origins [123]. Methylated DNA in the PJ can also provide clinically relevant information to diagnose PDAC, and three methylated DNA markers (C13orf18, FER1L4, and BMP3) can distinguish non-PDAC patients from PDAC patients, including those in the early stage or those with HG IPMN [124].
Recently, cfRNAs and EVs were investigated as potential tools for liquid biopsy (Table 2) [23, 125]. EVs comprise exosomes, apoptotic bodies, and microvesicles and are released by most cells, including cancer cells. EVs are microstructures with a lipid bilayer membrane containing RNAs, proteins, or lipids and transfer their contents from donor to recipient cells [126,127,128]. As EVs exist in circulating body fluids, such as blood, PJ, and DF, the expression levels of their contents, mRNA, protein, and non-coding RNAs (ncRNAs), have been used as biomarkers for PDAC diagnosis [25]. ncRNAs are non-protein-coding RNAs that regulate diverse biological processes in several diseases [129]. In particular, ncRNAs, non-protein-coding RNAs regulating various biological processes in several diseases, have recently been investigated regarding disease pathogenesis, including several cancers. They are divided into two main groups based on their transcript size. Short ncRNAs, which are less than 200 nucleotides in length, include microRNAs (miRNAs) and other small RNA classes, whereas long non-coding RNAs (lncRNAs) are greater than 200 nucleotides long [129,130,131].
In a blood examination, serum EV-encapsulated miR-21 was upregulated in patients with PDAC compared with healthy individuals and has been identified as a prognostic and chemoresistance marker [132]. Combined analysis of serum EVs miRNAs, miR-1246, miR-4644, miR-3976, and miR-4306, with PDAC-initiating cell markers, such as CD44 and EpCAM, could contribute to the detection of the early stages of PDAC [133]. Yang et al. investigated the usefulness of the expression levels of 16 plasma EV-related proteins as biomarkers of the malignant potential of IPMN. They found that MUC5AC expression was significantly higher in IPMN-associated carcinoma than in low-grade IPMN [134]. In contrast, some lncRNAs are detected as cfRNAs or EV-encapsulated RNAs in blood [25]. lncRNA MALAT-1 and HOTAIR expression is significantly increased in blood derived from patients with PDAC, suggesting their utility as biomarkers [141, 142]. Furthermore, lncRNA Sox2ot is present in plasma EVs and is highly expressed in patients with PDAC [137].
Nakamura et al. reported that expression levels of miR-21 and miR-155 within exosomes distinguished PDAC patients from chronic pancreatitis (CP) patients [27]; however, PDAC and CP patients could not be characterized based on miR-21 and miR-155 levels when whole RNAs from PJ were analyzed. The accuracy of exosomal miR-21 and miR-155 levels and PJC was 83%, 89%, and 74%, respectively. Another study revealed that EV-derived miR-21, miR-25, and miR-16 in the PJ were increased in patients with PDAC than in non-malignant controls. Combining these EV miRNAs with serum miR-210 and CA-19-9 showed an area under the curve of 0.91, a specificity of 84.2%, and a sensitivity of 81.5% for PDAC diagnosis [138]. They also investigated the size and concentration of EVs in PJ following secretin stimulation. EV concentrations did not differ between healthy controls and patients with PDAC; however, the PJ from PDAC had more large vesicles [143]. Moreover, in PJ-derived EVs, MUC1, MUC4, MUC5AC, MUC6, MUC16, CFTR, and MDR1 have been recognized as candidate markers for PDAC [139]. In addition, exosomal proteins carcinoembryonic antigen-related cell adhesion molecule (CEACAM) 1/5 and tenascin C have been identified as the most discriminating proteins between patients with PDAC and benign controls [140].
Besides miRNAs, mRNAs, and proteins, EV-encapsulated lncRNAs in PJ are yet to be investigated. We previously reported that EV lncRNA HULC in the serum was highly expressed in PDAC patients compared with IPMN patients or healthy individuals and served as a potential biomarker [144]. Therefore, similar to blood samples, several lncRNAs in the PJ may be crucial markers for detecting the early stages of PDAC. Moreover, no studies have evaluated EVs or cfRNAs to diagnose PDAC using DF. Considering the evidence showing that ncRNAs in the blood can be assessed through liquid biopsy [25], the analysis of ncRNAs and EVs is needed to discover previously unrecognized biomarkers in body fluids such as PJ and DF.
The relationship between the microbiome and PDAC has been increasingly reported in the past decade. In 2015, Fusobacterium species were found in PDAC tumors [145]. In 2019, Malassezia, a type of fungus, was shown to be increased in the pancreas of PDAC patients by 18S rRNA sequencing [146]. In 2022, certain strains of gut microbiota were shown to be more prevalent in patients with pancreatic cancer across racial groups (Japanese, Spanish, and German) using shotgun metagenomic analysis [147]. Examination of the PJ microbiome in resected pancreatic cancer tissue samples revealed no specific trends [148]. There remains much potential for microbiome research, not only as a direct cause of pancreatic carcinogenesis but also for the search for biomarkers for early diagnosis, the identification of new risk factors, and the prediction of treatment efficacy after diagnosis.
In this review, we have examined various studies conducted in different laboratories. However, it is difficult to determine the levels of evidence in each study. To address this issue, we need to establish specific protocols for liquid biopsies that standardize the sample collection process and ensure quality control throughout the molecular analysis process. By doing so, we can improve the consistency between investigators and better identify the origin tumor using liquid biopsy. In addition, integrating the information on where and how liquid biopsy factors are released into the body fluid and collected, sample size, single institute or multicenter study, and prospective study or study using existing samples will enhance compatibility across studies and deepen our understanding of the topics.
Ongoing projects using PJ or DF
We have been studying the PJC using synthetic secretin since 2012 and are conducting a multicenter study on MRI imaging and liquid biopsy in addition to PJC (CRB6200003). Synthetic secretins such as ChiRhoStim (ChiRho-Clin, Inc., Burtonsville, MD, USA) and Secrelux (Sanochemia, Vienna, Austria) have been used in the past; the former is currently in use. We previously reported that administering 0.6 μg of synthetic human secretin increases the amount of PJ collected using a catheter-inserted transpapillary [149, 150]. For the substantial masses, collected volume increased from 2.0 ± 2.1 mL (range, 0–14.0 mL) to 3.7 ± 2.5 mL (range, 1–15.0 mL) in PDAC patients [149] and from 3.7 ± 7.3 mL (range:0–79.0 mL) to 5.1 ± 6.2 mL (range:1–64.0 mL) in IPMN patients [150]. Furthermore, in diagnosing malignancy using PJC, we found that the sensitivity improved from 50.9 to 74.0% and from 50.0 to 70.8% in the case of a substantial mass and IPMN, respectively [149, 150].
We also evaluated mutation detection in DNA from DF (DFDNA) as an alternative to plasma DNA (UMIN000028284). In 150 patients suspected of PDAC and IPMN, DF containing PJ was collected from the duodenal lumen through an endoscopic channel. DFDNA yielded significantly higher levels of plasma cfDNA and exhibited characteristic fragmentation patterns. Using molecular barcode sequencing, hotspot mutations detected in DF showed higher concordance with resected tissue than with plasma cfDNA. In contrast to plasma cfDNA, in which liquid tissue matches were limited to advanced-stage cancer, the DFDNA assay enabled higher concordance across a wide range of PDA stages. We showed that DF-derived DNA could be a potent biospecimen for detecting mutations with higher yield and sensitivity than plasma cfDNA [120].
In contrast, genuine PJ-derived DNA had a much higher mutation detection rate. During endoscopic retrograde cholangiopancreatography (ERCP) testing, pancreatic fluid was aspirated via a catheter from the main pancreatic duct using a contrast agent. Mutation analysis using molecular barcoding of DNA from 1 mL of PJ showed a much higher concordance rate for PJDNA than for DFDNA. Furthermore, in addition to mutations derived from the index lesion, several other mutations that might have originated from multicentric lesions in the residual pancreas were detected in the PJDNA (manuscript in preparation) [151]. Therefore, mutational analysis using PJDNA may be helpful for auxiliary diagnosis when PJ is collected for cytological analysis.
Future directions
To investigate the usefulness of PJ-derived genes or EVs, we have initiated a prospective multicenter study to analyze EV RNAs, including lncRNAs and cfDNAs, in the PJ and plasma as liquid biopsy tools for the early diagnosis of PDAC (CRB6200003). Based on our previous report demonstrating the efficacy of synthetic secretin injections on the diagnostic accuracy of PJC [149], PJ samples are now collected following secretin administration. The dataset of DNA mutations and EV RNA expression in plasma and PJ will be integrated to develop a multi-layer diagnostic agonism for the early stages of PDAC diagnosis. These trials may offer new insights for develo** diagnostic strategies and identifying therapeutic targets for deadly cancers.
Perspectives
The utility of liquid biopsy is increasingly being explored, and several markers for the early diagnosis of human PDAC in humans are being identified. Novel cancer-associated RNAs as biomarkers and their roles in regulating tumor phenotypes are being investigated. In particular, ncRNAs seem promising for defining disease pathogenesis and serving as diagnostic and prognostic markers and therapeutic targets. Mutation analysis of liquid samples can supplement pathological diagnosis or may even be an alternative; however, there is a need to improve the diagnostic accuracy and identification of early PDAC using body fluids such as blood and PJ, and DF obtained via endoscopy. Therefore, a clinical application system must be promptly developed based on current and future trials. In addition, focused studies on the genes implicated in pancreatic diseases must be conducted to improve our understanding of disease pathogenesis and to aid their utility as diagnostic markers, so they can eventually lead to novel clinical applications.
References
Siegel RL, Miller KD, Fuchs HE, et al. Cancer statistics, 2021. CA Cancer J Clin. 2021;71:7–33.
Rahib L, Smith BD, Aizenberg R, et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014;74:2913–21.
Lee JC, Ahn S, Cho IK, et al. Management of recurrent pancreatic cancer after surgical resection: a protocol for systematic review, evidence map** and meta-analysis. BMJ Open. 2018;8:e017249.
Vincent A, Herman J, Schulick R, et al. Pancreatic cancer. Lancet. 2011;378:607–20.
Kamisawa T, Wood LD, Itoi T, et al. Pancreatic cancer. Lancet. 2016;388:73–85.
Lai E, Puzzoni M, Ziranu P, et al. New therapeutic targets in pancreatic cancer. Cancer Treat Rev. 2019;81:101926.
Hidalgo M. Pancreatic cancer. N Engl J Med. 2010;362:1605–17.
Tanaka S, Nakao M, Ioka T, et al. Slight dilatation of the main pancreatic duct and presence of pancreatic cysts as predictive signs of pancreatic cancer: a prospective study. Radiology. 2010;254:965–72.
Tanaka S, Fukuda J, Nakao M, et al. Effectiveness of contrast-enhanced ultrasonography for the characterization of small and early stage pancreatic adenocarcinoma. Ultrasound Med Biol. 2020;46:2245–53.
Chu LC, Goggins MG, Fishman EK. Diagnosis and detection of pancreatic cancer. Cancer J. 2017;23:333–42.
Pijpe A, Andrieu N, Easton DF, et al. Exposure to diagnostic radiation and risk of breast cancer among carriers of BRCA1/2 mutations: retrospective cohort study (GENE-RAD-RISK). BMJ. 2012;345:e5660.
Vilmann P, Jacobsen GK, Henriksen FW, et al. Endoscopic ultrasonography with guided fine needle aspiration biopsy in pancreatic disease. Gastrointest Endosc. 1992;38:172–3.
Dewitt J, Devereaux BM, Lehman GA, et al. Comparison of endoscopic ultrasound and computed tomography for the preoperative evaluation of pancreatic cancer: a systematic review. Clin Gastroenterol Hepatol. 2006;4:717–25.
Hou X, Liang J, Konge L, et al. Training and certification of EUS operators in China. Endosc Ultrasound. 2022;11:133–40.
Hanada K, Okazaki A, Hirano N, et al. Diagnostic strategies for early pancreatic cancer. J Gastroenterol. 2015;50:147–54.
Matsumoto K, Kushiyama Y, Izumi A, et al. Usefulness of practitioner-led pancreatic cancer screening. Diagnostics (Basel). 2022. https://doi.org/10.3390/diagnostics12092257.
Chari ST, Maitra A, Matrisian LM, et al. Early detection initiative: a randomized controlled trial of algorithm-based screening in patients with new onset hyperglycemia and diabetes for early detection of pancreatic ductal adenocarcinoma. Contemp Clin Trials. 2022;113:106659.
Ni XG, Bai XF, Mao YL, et al. The clinical value of serum CEA, CA19-9, and CA242 in the diagnosis and prognosis of pancreatic cancer. Eur J Surg Oncol. 2005;31:164–9.
Fahrmann JF, Schmidt CM, Mao X, et al. Lead-time trajectory of CA19-9 as an anchor marker for pancreatic cancer early detection. Gastroenterology. 2021;160(1373–1383): e6.
Liu J, Gao J, Du Y, et al. Combination of plasma microRNAs with serum CA19-9 for early detection of pancreatic cancer. Int J Cancer. 2012;131:683–91.
Ono M, Ono Y, Nakamura T, et al. Predictors of long-term survival in pancreatic ductal adenocarcinoma after pancreatectomy: TP53 and SMAD4 mutation scoring in combination with CA19-9. Ann Surg Oncol. 2022;29:5007–19.
Crowley E, Di Nicolantonio F, Loupakis F, et al. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol. 2013;10:472–84.
Yoshioka Y, Katsuda T, Ochiya T. Extracellular vesicles and encapusulated miRNAs as emerging cancer biomarkers for novel liquid biopsy. Jpn J Clin Oncol. 2018;48:869–76.
Melo SA, Luecke LB, Kahlert C, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523:177–82.
Takahashi K, Taniue K, Ono Y, et al. Long non-coding RNAs in epithelial-mesenchymal transition of pancreatic cancer. Front Mol Biosci. 2021;8:717890.
Okada T, Mizukami Y, Ono Y, et al. Digital PCR-based plasma cell-free DNA mutation analysis for early-stage pancreatic tumor diagnosis and surveillance. J Gastroenterol. 2020;55:1183–93.
Nakamura S, Sadakari Y, Ohtsuka T, et al. Pancreatic juice exosomal MicroRNAs as biomarkers for detection of pancreatic ductal adenocarcinoma. Ann Surg Oncol. 2019;26:2104–11.
Okada T, Iwano H, Ono Y, et al. Utility of “liquid biopsy” using pancreatic juice for early detection of pancreatic cancer. Endosc Int Open. 2018;6:E1454–61.
Morris JPt, Cano DA, Sekine S, et al. Beta-catenin blocks Kras-dependent reprogramming of acini into pancreatic cancer precursor lesions in mice. J Clin Invest. 2010;120:508–20.
Krasinskas AM, Moser AJ, Saka B, et al. KRAS mutant allele-specific imbalance is associated with worse prognosis in pancreatic cancer and progression to undifferentiated carcinoma of the pancreas. Mod Pathol. 2013;26:1346–54.
Kanda M, Matthaei H, Wu J, et al. Presence of somatic mutations in most early-stage pancreatic intraepithelial neoplasia. Gastroenterology. 2012;142:730-733 e9.
Bailey P, Chang DK, Nones K, et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature. 2016;531:47–52.
Cao L, Huang C, Cui Zhou D, et al. Proteogenomic characterization of pancreatic ductal adenocarcinoma. Cell. 2021;184:5031-5052 e26.
Bournet B, Muscari F, Buscail C, et al. KRAS G12D mutation subtype is a prognostic factor for advanced pancreatic adenocarcinoma. Clin Transl Gastroenterol. 2016;7:e157.
Mueller S, Engleitner T, Maresch R, et al. Evolutionary routes and KRAS dosage define pancreatic cancer phenotypes. Nature. 2018;554:62–8.
Ryan DP, Hong TS, Bardeesy N. Pancreatic adenocarcinoma. N Engl J Med. 2014;371:1039–49.
Patra KC, Bardeesy N, Mizukami Y. Diversity of precursor lesions for pancreatic cancer: the genetics and biology of intraductal papillary mucinous neoplasm. Clin Transl Gastroenterol. 2017;8:e86.
Drelon C, Berthon A, Sahut-Barnola I, et al. PKA inhibits WNT signalling in adrenal cortex zonation and prevents malignant tumour development. Nat Commun. 2016;7:12751.
Kawabata H, Ono Y, Tamamura N, et al. Mutant GNAS limits tumor aggressiveness in established pancreatic cancer via antagonizing the KRAS-pathway. J Gastroenterol. 2022;57:208–20.
**ng F, Luan Y, Cai J, et al. The anti-Warburg effect elicited by the cAMP-PGC1alpha pathway drives differentiation of glioblastoma cells into astrocytes. Cell Rep. 2017;18:468–81.
Watanabe K, Nakamura T, Onodera S, et al. A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors. Tumour Biol. 2020;42:1010428320962588.
Lee JH, Kim Y, Choi JW, et al. KRAS, GNAS, and RNF43 mutations in intraductal papillary mucinous neoplasm of the pancreas: a meta-analysis. Springerplus. 2016;5:1172.
Omori Y, Ono Y, Kobayashi T, et al. How does intestinal-type intraductal papillary mucinous neoplasm emerge? CDX2 plays a critical role in the process of intestinal differentiation and progression. Virchows Arch. 2020;477:21–31.
Patra KC, Kato Y, Mizukami Y, et al. Mutant GNAS drives pancreatic tumourigenesis by inducing PKA-mediated SIK suppression and reprogramming lipid metabolism. Nat Cell Biol. 2018;20:811–22.
Iglesias-Bartolome R, Torres D, Marone R, et al. Inactivation of a Galpha(s)-PKA tumour suppressor pathway in skin stem cells initiates basal-cell carcinogenesis. Nat Cell Biol. 2015;17:793–803.
He X, Zhang L, Chen Y, et al. The G protein alpha subunit Galphas is a tumor suppressor in Sonic hedgehog-driven medulloblastoma. Nat Med. 2014;20:1035–42.
Jenne DE, Reimann H, Nezu J, et al. Peutz-Jeghers syndrome is caused by mutations in a novel serine threonine kinase. Nat Genet. 1998;18:38–43.
Omori Y, Ono Y, Morikawa T, et al. Serine/Threonine Kinase 11 Plays a Canonical Role in Malignant Progression of KRAS-mutant and GNAS-wild-type Intraductal Papillary Mucinous Neoplasms of the Pancreas. Ann Surg. 2021.
Sakamoto H, Kuboki Y, Hatori T, et al. Clinicopathological significance of somatic RNF43 mutation and aberrant expression of ring finger protein 43 in intraductal papillary mucinous neoplasms of the pancreas. Mod Pathol. 2015;28:261–7.
Jiang X, Hao HX, Growney JD, et al. Inactivating mutations of RNF43 confer Wnt dependency in pancreatic ductal adenocarcinoma. Proc Natl Acad Sci U S A. 2013;110:12649–54.
Gala MK, Mizukami Y, Le LP, et al. Germline mutations in oncogene-induced senescence pathways are associated with multiple sessile serrated adenomas. Gastroenterology. 2014;146:520–9.
Noe M, Niknafs N, Fischer CG, et al. Genomic characterization of malignant progression in neoplastic pancreatic cysts. Nat Commun. 2020;11:4085.
Fujikura K, Hosoda W, Felsenstein M, et al. Multiregion whole-exome sequencing of intraductal papillary mucinous neoplasms reveals frequent somatic KLF4 mutations predominantly in low-grade regions. Gut. 2021;70:928–39.
Fischer CG, Beleva Guthrie V, Braxton AM, et al. Intraductal papillary mucinous neoplasms arise from multiple independent clones, each with distinct mutations. Gastroenterology. 2019;157:1123-1137 e22.
Greaves M, Maley CC. Clonal evolution in cancer. Nature. 2012;481:306–13.
Kopp JL, von Figura G, Mayes E, et al. Identification of Sox9-dependent acinar-to-ductal reprogramming as the principal mechanism for initiation of pancreatic ductal adenocarcinoma. Cancer Cell. 2012;22:737–50.
Aichler M, Seiler C, Tost M, et al. Origin of pancreatic ductal adenocarcinoma from atypical flat lesions: a comparative study in transgenic mice and human tissues. J Pathol. 2012;226:723–34.
Grimont A, Leach SD, Chandwani R. Uncertain beginnings: acinar and ductal cell plasticity in the development of pancreatic cancer. Cell Mol Gastroenterol Hepatol. 2022;13:369–82.
Huang L, Desai R, Conrad DN, et al. Commitment and oncogene-induced plasticity of human stem cell-derived pancreatic acinar and ductal organoids. Cell Stem Cell. 2021;28:1090-1104 e6.
Collisson EA, Bailey P, Chang DK, et al. Molecular subtypes of pancreatic cancer. Nat Rev Gastroenterol Hepatol. 2019;16:207–20.
Connor AA, Gallinger S. Pancreatic cancer evolution and heterogeneity: integrating omics and clinical data. Nat Rev Cancer. 2022;22:131–42.
Baba T, Finetti P, Pancreatic Cancer G, et al. A lesson in transcriptional plasticity: classical identity is silenced, but not lost, in pancreatic ductal adenocarcinoma cell lines. Gastroenterology. 2022;163:1450-1453 e3.
Hayashi A, Hong J, Iacobuzio-Donahue CA. The pancreatic cancer genome revisited. Nat Rev Gastroenterol Hepatol. 2021;18:469–81.
Kokumai T, Omori Y, Ishida M, et al. GATA6 and CK5 stratify the survival of patients with pancreatic cancer undergoing neoadjuvant chemotherapy. Mod Pathol. 2023;36:100102.
de Andres MP, Jackson RJ, Felipe I, et al. GATA4 and GATA6 loss-of-expression is associated with extinction of the classical programme and poor outcome in pancreatic ductal adenocarcinoma. Gut. 2023;72:535–48.
Flowers BM, Xu H, Mulligan AS, et al. Cell of origin influences pancreatic cancer subtype. Cancer Discov. 2021;11:660–77.
Moffitt RA, Marayati R, Flate EL, et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat Genet. 2015;47:1168–78.
Larson MH, Pan W, Kim HJ, et al. A comprehensive characterization of the cell-free transcriptome reveals tissue- and subtype-specific biomarkers for cancer detection. Nat Commun. 2021;12:2357.
Pishvaian MJ, Blais EM, Brody JR, et al. Overall survival in patients with pancreatic cancer receiving matched therapies following molecular profiling: a retrospective analysis of the Know Your Tumor registry trial. Lancet Oncol. 2020;21:508–18.
Motoi F, Kosuge T, Ueno H, et al. Randomized phase II/III trial of neoadjuvant chemotherapy with gemcitabine and S-1 versus upfront surgery for resectable pancreatic cancer (Prep-02/JSAP05). Japanese J Clin Oncol. 2019;49:190–4.
Ye M, Zhang Q, Chen Y, et al. Neoadjuvant chemotherapy for primary resectable pancreatic cancer: a systematic review and meta-analysis. HPB (Oxford). 2020;22:821–32.
Yang Y, Li L, Qu C, et al. Endoscopic ultrasound-guided fine needle core biopsy for the diagnosis of pancreatic malignant lesions: a systematic review and meta-analysis. Sci Rep. 2016;6:22978.
Banafea O, Mghanga FP, Zhao J, et al. Endoscopic ultrasonography with fine-needle aspiration for histological diagnosis of solid pancreatic masses: a meta-analysis of diagnostic accuracy studies. BMC Gastroenterol. 2016;16:108.
Gkolfakis P, Crino SF, Tziatzios G, et al. Comparative diagnostic performance of end-cutting fine-needle biopsy needles for EUS tissue sampling of solid pancreatic masses: a network meta-analysis. Gastrointest Endosc. 2022;95:1067-1077 e15.
Itonaga M, Yasukawa S, Fukutake N, et al. Comparison of 22-gauge standard and Franseen needles in EUS-guided tissue acquisition for diagnosing solid pancreatic lesions: a multicenter randomized controlled trial. Gastrointest Endosc. 2022;96:57-66 e2.
Adler DG, Witt B, Chadwick B, et al. Pathologic evaluation of a new endoscopic ultrasound needle designed to obtain core tissue samples: a pilot study. Endosc Ultrasound. 2016;5:178–83.
Bang JY, Hebert-Magee S, Hasan MK, et al. Endoscopic ultrasonography-guided biopsy using a Franseen needle design: Initial assessment. Dig Endosc. 2017;29:338–46.
Hebert-Magee S, Bae S, Varadarajulu S, et al. The presence of a cytopathologist increases the diagnostic accuracy of endoscopic ultrasound-guided fine needle aspiration cytology for pancreatic adenocarcinoma: a meta-analysis. Cytopathology. 2013;24:159–71.
Matsumoto K, Hara K, Yasuda I, et al. Usefulness of a target sample check illuminator in the detection of target specimens in endoscopic ultrasound-guided fine-needle biopsy samples: Multicenter prospective study. Dig Endosc. 2021;33:970–6.
Mohamadnejad M, Mullady D, Early DS, et al. Increasing number of passes beyond 4 does not increase sensitivity of detection of pancreatic malignancy by endoscopic ultrasound-guided fine-needle aspiration. Clin Gastroenterol Hepatol. 2017;15:1071-1078 e2.
Kitano M, Yoshida M, Ashida R, et al. Needle tract seeding after endoscopic ultrasound-guided tissue acquisition of pancreatic tumors: a nationwide survey in Japan. Dig Endosc. 2022;34:1442.
Uchida N, Kamada H, Tsutsui K, et al. Utility of pancreatic duct brushing for diagnosis of pancreatic carcinoma. J Gastroenterol. 2007;42:657–62.
Iiboshi T, Hanada K, Fukuda T, et al. Value of cytodiagnosis using endoscopic nasopancreatic drainage for early diagnosis of pancreatic cancer: establishing a new method for the early detection of pancreatic carcinoma in situ. Pancreas. 2012;41:523–9.
Nakaizumi A, Tatsuta M, Uehara H, et al. Cytologic examination of pure pancreatic juice in the diagnosis of pancreatic carcinoma. The endoscopic retrograde intraductal catheter aspiration cytologic technique. Cancer. 1992;70:2610–4.
Kitagawa K, Mitoro A, Tomooka F, et al. Diagnostic yield of liquid-based cytology in serial pancreatic juice aspiration cytological examination. DEN Open. 2023;3:e177.
Mie T, Sasaki T, Takeda T, et al. Diagnostic yield of serial pancreatic juice aspiration cytologic examination with brush cytology for pancreatic ductal stenosis. Pancreas. 2022;51:995–9.
Mikata R, Ishihara T, Tada M, et al. Clinical usefulness of repeated pancreatic juice cytology via endoscopic naso-pancreatic drainage tube in patients with pancreatic cancer. J Gastroenterol. 2013;48:866–73.
Iwata T, Kitamura K, Yamamiya A, et al. Evaluation of diagnostic cytology via endoscopic naso-pancreatic drainage for pancreatic tumor. World J Gastrointest Endosc. 2014;6:366–72.
Naitoh I, Nakazawa T, Kato A, et al. Predictive factors for positive diagnosis of malignant biliary strictures by transpapillary brush cytology and forceps biopsy. J Dig Dis. 2016;17:44–51.
Horwhat JD, Paulson EK, McGrath K, et al. A randomized comparison of EUS-guided FNA versus CT or US-guided FNA for the evaluation of pancreatic mass lesions. Gastrointest Endosc. 2006;63:966–75.
Li M, Diehl F, Dressman D, et al. BEAMing up for detection and quantification of rare sequence variants. Nat Methods. 2006;3:95–7.
Salipante SJ, Jerome KR. Digital PCR-an emerging technology with broad applications in microbiology. Clin Chem. 2020;66:117–23.
Hindson BJ, Ness KD, Masquelier DA, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem. 2011;83:8604–10.
Alcaide M, Cheung M, Bushell K, et al. A novel multiplex droplet digital PCR assay to identify and quantify KRAS mutations in clinical specimens. J Mol Diagn. 2019;21:214–27.
Whale AS, Huggett JF, Tzonev S. Fundamentals of multiplexing with digital PCR. Biomol Detect Quantif. 2016;10:15–23.
Chiho M, Ono Y, Hayashi A, et al. Multiplex digital PCR assay to detect multiple KRAS and GNAS mutations associated with pancreatic carcinogenesis from minimal specimen amounts. J Mol Diagn. 2023;25(6):367–77.
Rowlands V, Rutkowski AJ, Meuser E, et al. Optimisation of robust singleplex and multiplex droplet digital PCR assays for high confidence mutation detection in circulating tumour DNA. Sci Rep. 2019;9:12620.
Hoadley KA, Yau C, Hinoue T, et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell. 2018;173:291-304 e6.
Kou R, Lam H, Duan H, et al. Benefits and challenges with applying unique molecular identifiers in next generation sequencing to detect low frequency mutations. PLoS ONE. 2016;11:e0146638.
Narayan A, Carriero NJ, Gettinger SN, et al. Ultrasensitive measurement of hotspot mutations in tumor DNA in blood using error-suppressed multiplexed deep sequencing. Cancer Res. 2012;72:3492–8.
Ebi H, Bando H. Precision oncology and the universal health coverage system in Japan. JCO Precis Oncol. 2019. https://doi.org/10.1200/PO.19.00291.
Chung C, Galvin R, Achenbach E, et al. Characterization of blood-based molecular profiling in pancreatic adenocarcinoma. Oncology (Williston Park). 2021;35:794–803.
Hellmann MD, Ciuleanu TE, Pluzanski A, et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N Engl J Med. 2018;378:2093–104.
Gandara DR, Paul SM, Kowanetz M, et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat Med. 2018;24:1441–8.
Postel M, Roosen A, Laurent-Puig P, et al. Droplet-based digital PCR and next generation sequencing for monitoring circulating tumor DNA: a cancer diagnostic perspective. Expert Rev Mol Diagn. 2018;18:7–17.
Bettegowda C, Sausen M, Leary RJ, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6:22424.
Takai E, Totoki Y, Nakamura H, et al. Clinical utility of circulating tumor DNA for molecular assessment in pancreatic cancer. Sci Rep. 2015;5:18425.
Hadano N, Murakami Y, Uemura K, et al. Prognostic value of circulating tumour DNA in patients undergoing curative resection for pancreatic cancer. Br J Cancer. 2016;115:59–65.
Buscail L, Bournet B, Cordelier P. Role of oncogenic KRAS in the diagnosis, prognosis and treatment of pancreatic cancer. Nat Rev Gastroenterol Hepatol. 2020;17:153–68.
Brychta N, Krahn T, von Ahsen O. Detection of KRAS mutations in circulating tumor DNA by digital PCR in early stages of pancreatic cancer. Clin Chem. 2016;62:1482–91.
Ono Y, Sugitani A, Karasaki H, et al. An improved digital polymerase chain reaction protocol to capture low-copy KRAS mutations in plasma cell-free DNA by resolving “subsampling” issues. Mol Oncol. 2017;11:1448–58.
Allenson K, Castillo J, San Lucas FA, et al. High prevalence of mutant KRAS in circulating exosome-derived DNA from early-stage pancreatic cancer patients. Ann Oncol. 2017;28:741–7.
Tada M, Kawabe T, Arizumi M, et al. Pancreatic cancer in patients with pancreatic cystic lesions: a prospective study in 197 patients. Clin Gastroenterol Hepatol. 2006;4:1265–70.
Tada M, Tateishi K, Kawabe T, et al. Quantity of mutant K-ras gene in pancreatic secretions for diagnosis of pancreatic carcinoma with different assays: analysis of 100 patients. Clin Chim Acta. 2002;324:105–11.
Tateishi K, Tada M, Yamagata M, et al. High proportion of mutant K-ras gene in pancreatic juice of patients with pancreatic cystic lesions. Gut. 1999;45:737–40.
Eshleman JR, Norris AL, Sadakari Y, et al. KRAS and guanine nucleotide-binding protein mutations in pancreatic juice collected from the duodenum of patients at high risk for neoplasia undergoing endoscopic ultrasound. Clin Gastroenterol Hepatol. 2015;13:963-9 e4.
Takano S, Fukasawa M, Maekawa S, et al. Deep sequencing of cancer-related genes revealed GNAS mutations to be associated with intraductal papillary mucinous neoplasms and its main pancreatic duct dilation. PLoS ONE. 2014;9:e98718.
Takano S, Fukasawa M, Kadokura M, et al. Mutational patterns in pancreatic juice of intraductal papillary mucinous neoplasms and concomitant pancreatic cancer. Pancreas. 2019;48:1032–40.
Yu J, Sadakari Y, Shindo K, et al. Digital next-generation sequencing identifies low-abundance mutations in pancreatic juice samples collected from the duodenum of patients with pancreatic cancer and intraductal papillary mucinous neoplasms. Gut. 2017;66:1677–87.
Suenaga M, Yu J, Shindo K, et al. Pancreatic juice mutation concentrations can help predict the grade of dysplasia in patients undergoing pancreatic surveillance. Clin Cancer Res. 2018;24:2963–74.
Lo YMD, Han DSC, Jiang P, et al. Epigenetics, fragmentomics, and topology of cell-free DNA in liquid biopsies. Science. 2021. https://doi.org/10.1126/science.aaw3616.
Shen SY, Burgener JM, Bratman SV, et al. Preparation of cfMeDIP-seq libraries for methylome profiling of plasma cell-free DNA. Nat Protoc. 2019;14:2749–80.
Shimizu D, Taniue K, Matsui Y, et al. Pan-cancer methylome analysis for cancer diagnosis and classification of cancer cell of origin. Cancer Gene Ther. 2022;29:428–36.
Majumder S, Raimondo M, Taylor WR, et al. Methylated DNA in pancreatic juice distinguishes patients with pancreatic cancer from controls. Clin Gastroenterol Hepatol. 2020;18:676-683 e3.
Liang K, Liu F, Fan J, et al. Nanoplasmonic quantification of tumor-derived extracellular vesicles in plasma microsamples for diagnosis and treatment monitoring. Nat Biomed Eng. 2017. https://doi.org/10.1038/s41551-016-0021.
Kalra H, Simpson RJ, Ji H, et al. Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol. 2012;10:e1001450.
Thery C, Ostrowski M, Segura E. Membrane vesicles as conveyors of immune responses. Nat Rev Immunol. 2009;9:581–93.
Kalluri R, McAndrews KM. The role of extracellular vesicles in cancer. Cell. 2023;186:1610–26.
Takahashi K, Yan I, Haga H, et al. Long noncoding RNA in liver diseases. Hepatology. 2014;60:744–53.
Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172:393–407.
Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol. 2018;19:143–57.
Goto T, Fujiya M, Konishi H, et al. An elevated expression of serum exosomal microRNA-191, - 21, -451a of pancreatic neoplasm is considered to be efficient diagnostic marker. BMC Cancer. 2018;18:116.
Madhavan B, Yue S, Galli U, et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int J Cancer. 2015;136:2616–27.
Yang KS, Ciprani D, O’Shea A, et al. Extracellular vesicle analysis allows for identification of invasive IPMN. Gastroenterology. 2021;160:1345-1358 e11.
Kumar SR, Kimchi ET, Manjunath Y, et al. Rna cargos in extracellular vesicles derived from blood serum in pancreas associated conditions. Sci Rep. 2020;10:2800.
Ma Y, Hu M, Zhou L, et al. Long non-coding rna hotair promotes cancer cell energy metabolism in pancreatic adenocarcinoma by upregulating hexokinase-2. Oncol Lett. 2019;18:2212–9.
Li Z, Jiang P, Li J, et al. Tumor-derived exosomal lnc-Sox2ot promotes EMT and stemness by acting as a ceRNA in pancreatic ductal adenocarcinoma. Oncogene. 2018;37:3822–38.
Nesteruk K, Levink IJM, de Vries E, et al. Extracellular vesicle-derived microRNAs in pancreatic juice as biomarkers for detection of pancreatic ductal adenocarcinoma. Pancreatology. 2022;22:626–35.
Osteikoetxea X, Benke M, Rodriguez M, et al. Detection and proteomic characterization of extracellular vesicles in human pancreatic juice. Biochem Biophys Res Commun. 2018;499:37–43.
Zheng J, Hernandez JM, Doussot A, et al. Extracellular matrix proteins and carcinoembryonic antigen-related cell adhesion molecules characterize pancreatic duct fluid exosomes in patients with pancreatic cancer. HPB (Oxford). 2018;20:597–604.
Jiao F, Hu H, Han T, et al. Long noncoding RNA MALAT-1 enhances stem cell-like phenotypes in pancreatic cancer cells. Int J Mol Sci. 2015;16:6677–93.
Kim K, Jutooru I, Chadalapaka G, et al. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene. 2013;32:1616–25.
Nesteruk K, Levink IJM, Dits NFJ, et al. Size and concentration of extracellular vesicles in pancreatic juice from patients with pancreatic ductal adenocarcinoma. Clin Transl Gastroenterol. 2022;13:e00465.
Takahashi K, Ota Y, Kogure T, et al. Circulating extracellular vesicle-encapsulated HULC is a potential biomarker for human pancreatic cancer. Cancer Sci. 2020;111:98–111.
Mitsuhashi K, Nosho K, Sukawa Y, et al. Association of Fusobacterium species in pancreatic cancer tissues with molecular features and prognosis. Oncotarget. 2015;6:7209–20.
Aykut B, Pushalkar S, Chen R, et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature. 2019;574:264–7.
Nagata N, Nishijima S, Kojima Y, et al. Metagenomic identification of microbial signatures predicting pancreatic cancer from a multinational study. Gastroenterology. 2022;163:222–38.
Kohi S, Macgregor-Das A, Dbouk M, et al. Alterations in the duodenal fluid microbiome of patients with pancreatic cancer. Clin Gastroenterol Hepatol. 2022;20:e196–227.
Takeda Y, Matsumoto K, Kurumi H, et al. Efficacy and safety of pancreatic juice cytology by using synthetic secretin in the diagnosis of pancreatic ductal adenocarcinoma. Dig Endosc. 2018;30:771–6.
Takeda Y, Matsumoto K, Onoyama T, et al. Efficacy and safety of pancreatic juice cytology with synthetic secretin in diagnosing malignant intraductal papillary mucinous neoplasms of the pancreas. Diagnostics (Basel). 2022. https://doi.org/10.3390/diagnostics12030744.
Ono Y, Takahashi K, Karasaki H, et al. Abstract PR013: High sensitive mutation detection in liquid biopsy with duodenal fluid captures genetic mutations associated with pancreatic carcinogenesis. Cancer Res. 2022;82:PR013–PR013.
Acknowledgements
The authors thank their colleagues in the Department of Medicine, Asahikawa Medical University; Department of Multidisciplinary Internal Medicine, Faculty of Medicine, Tottori University; and Institute of Biomedical Research, Sapporo Higashi Tokushukai Hospital, for supporting the review.
Funding
This study was supported by JSPS KAKENHI (Grant Numbers 20H03655, 21K07954, and 22KK0125), the Pancreas Research Foundation of Japan (to YO), the Japanese Society of Gastroenterology Grant (to KT), and Princess Takamatsu Cancer Research Fund (to YM).
Author information
Authors and Affiliations
Contributions
KT, YT, and YO were involved in conceptualization, literature search, interpretation of the data, and drafting of the article. HI and YM performed critical revision of the article and supervision of the study.
Corresponding author
Ethics declarations
Conflict of interest
K.T., Y.O., and Y.M. received grant support from the Hitachi High-Tech Corporation. The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Takahashi, K., Takeda, Y., Ono, Y. et al. Current status of molecular diagnostic approaches using liquid biopsy. J Gastroenterol 58, 834–847 (2023). https://doi.org/10.1007/s00535-023-02024-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-023-02024-4