Abstract
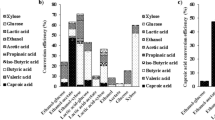
In this study, the cellular metabolic mechanisms regarding ammonium sulfate supplementation on erythromycin production were investigated by employing targeted metabolomics and metabolic flux analysis. The results suggested that the addition of ammonium sulfate stimulates erythromycin biosynthesis. Targeted metabolomics analysis uncovered that the addition of ammonium sulfate during the late stage of fermentation resulted in an augmented intracellular amino acid metabolism pool, guaranteeing an ample supply of precursors for organic acids and coenzyme A-related compounds. Therefore, adequate precursors facilitated cellular maintenance and erythromycin biosynthesis. Subsequently, an optimal supplementation rate of 0.02 g/L/h was determined. The results exhibited that erythromycin titer (1311.1 μg/mL) and specific production rate (0.008 mmol/gDCW/h) were 101.3% and 41.0% higher than those of the process without ammonium sulfate supplementation, respectively. Moreover, the erythromycin A component proportion increased from 83.2% to 99.5%. Metabolic flux analysis revealed increased metabolic fluxes with the supplementation of three ammonium sulfate rates.






Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Oliynyk M, Samborskyy M, Lester J, Mironenko T, Scott N, Dickens S, Leadlay P (2007) Complete genome sequence of the erythromycin-producing bacterium Saccharopolyspora erythraea NRRL23338. Nat Biotechnol 25:447–453
Mironov V, Sergienko O, Nastasyak I, Danilenko V (2004) Biogenesis and regulation of biosynthesis of erythromycins in Saccharopolyspora erythraea. Appl Biochem Microbiol 40:531–541
Li X, Ke X, Qiao L, Sui Y, Chu J (2022) Comparative genomic and transcriptomic analysis guides to further enhance the biosynthesis of erythromycin by an overproducer. Biotechnol Bioeng 119(6):1624–1640
Karničar K, Drobnak I, Petek M, Magdevska V, Horvat J, Vidmar R, Baebler Š, Rotter A, Jamnik P, Fujs Š et al (2016) Integrated omics approaches provide strategies for rapid erythromycin yield increase in Saccharopolyspora erythraea. Microb Cell Fact 15:93
Chen C, Hong M, Chu J, Huang M, Ouyang L, Tian X, Zhuang Y (2017) Blocking the flow of propionate into TCA cycle through a mutB knockout leads to a significant increase of erythromycin production by an industrial strain of Saccharopolyspora erythraea. Bioprocess Biosyst Eng 40:201–209
Xu F, Ke X, Hong M, Huang M, Chen C, Tian X, Hang H, Chu J (2021) Exploring the metabolic fate of propanol in industrial erythromycin-producing strain via 13C labeling experiments and enhancement of erythromycin production by rational metabolic engineering of Saccharopolyspora erythraea. Biochem Biophys Res Commun 542:73–79
Chen Y, Huang M, Wang Z, Chu J, Zhuang Y, Zhang S (2013) Controlling the feed rate of glucose and propanol for the enhancement of erythromycin production and exploration of propanol metabolism fate by quantitative metabolic flux analysis. Bioprocess Biosyst Eng 36:1445–1453
Xu F, Lu J, Ke X, Shao M, Huang M, Chu J (2022) Reconstruction of the genome-scale metabolic model of Saccharopolyspora erythraea and its application in the overproduction of erythromycin. Metabolites 12(6):509
Zou X, Hang H, Chu J, Zhuang Y, Zhang S (2009) Enhancement of erythromycin A production with feeding available nitrogen sources in erythromycin biosynthesis phase. Bioresour Technol 100:3358–3365
Technikova-Dobrova Z, Damiano F, Tredici S et al (2004) Design of mineral medium for growth of Actinomadura sp. ATCC 39727, producer of the glycopeptide A40926: effects of calcium ions and nitrogen sources. Appl Microbiol Biotechnol 65(6):671–677
Gonzalez R, Islas L, Obregon AM, Escalante L, Sanchez S (1995) Gentamicin formation in Micromonospora purpurea: stimulatory effect of ammonium. J Antibiot (Tokyo) 48(6):479–483
Zhang Q, Hang H, Tian X, Zeng W, Yu Z, Wang X, Tang Y, Zhuang Y, Chu J (2019) Combined available nitrogen resources enhanced erythromycin production and preliminary exploration of metabolic flux analysis under nitrogen perturbations. Bioprocess Biosyst Eng 42(11):1747–1756
Zhu C, Lu F, He Y, Han Z, Du L (2007) Regulation of avilamycin biosynthesis in Streptomyces viridochromogenes: effects of glucose, ammonium ion, and inorganic phosphate. Appl Microbiol Biotechnol 73(5):1031–1038
Sévin D, Kuehne A, Zamboni N, Sauer U (2015) Biological insights through nontargeted metabolomics. Curr Opin Biotechnol 34:1–8
Ellis D, Goodacre R (2012) Metabolomics-assisted synthetic biology. Curr Opin Biotechnol 23:22–28
**a M, Huang D, Li S, Wen J, Jia X, Chen Y (2013) Enhanced FK506 production in Streptomyces tsukubaensis by rational feeding strategies based on comparative metabolic profiling analysis. Biotechnol Bioeng 110:2717–2730
Taymaz-Nikerel H, De Mey M, Baart G, Maertens J, Heijnen J, van Gulik W (2013) Changes in substrate availability in Escherichia coli lead to rapid metabolite, flux and growth rate responses. Metab Eng 16:115–129
de Jonge LP, Buijs NA, ten Pierick A, Deshmukh A, Zhao Z, Kiel JA, Heijnen JJ, van Gulik WM (2011) Scale-down of penicillin production in Penicillium chrysogenum. Biotechnol J 6:944–958
Jordà J, Rojas H, Carnicer M, Wahl A, Ferrer P, Albiol J (2014) Quantitative metabolomics and instationary 13C-metabolic flux analysis reveals impact of recombinant protein production on trehalose and energy metabolism in Pichia pastoris. Metabolites 4:281–299
Zamboni N, Fendt S-M, Ruhl M, Sauer U (2009) 13C-based metabolic flux analysis. Nat Protoc 4:878–892
Antoniewicz M (2013) 13C metabolic flux analysis: optimal design of isotopic labeling experiments. Curr Opin Biotechnol 24:1116–1121
Niedenführ S, Wiechert W, Nöh K (2015) How to measure metabolic fluxes: a taxonomic guide for 13C fluxomics. Curr Opin Biotechnol 34:82–90
Driouch H, Melzer G, Wittmann C (2012) Integration of in vivo and in silico metabolic fluxes for improvement of recombinant protein production. Metab Eng 14:47–58
Lu H, Liu X, Huang M, **a J, Chu J, Zhuang Y, Zhang S, Noorman H (2015) Integrated isotope-assisted metabolomics and (13)C metabolic flux analysis reveals metabolic flux redistribution for high glucoamylase production by Aspergillus niger. Microb Cell Fact 14:147
Xu W, Xu F, Song W, Dong L, Qian J, Huang M (2022) Untargeted metabolomics combined with metabolic flux analysis reveals the mechanism of sodium citrate for high S-adenosyl-methionine production by Pichia pastoris. Fermentation 8(12):681
Hong M, Liao J, Chu J (2018) High-throughput optimization of the chemically defined synthetic medium for the production of erythromycin A. Bioprocess Biosyst Eng 41:1529–1538
Hong M, Mou H, Liu X, Huang M, Chu J (2017) 13C-assisted metabolomics analysis reveals the positive correlation between specific erythromycin production rate and intracellular propionyl-CoA pool size in Saccharopolyspora erythraea. Bioprocess Biosyst Eng 40:1337–1348
Xu F, Zhang X, Liu L, Ke X, Wu J, Guo Y, Tian X, Chu J (2022) Engineering the methyltransferase through inactivation of the genK and genL leads to a significant increase of gentamicin C1a production in an industrial strain of Micromonospora echinospora 49-92S. Bioprocess Biosyst Eng 45(10):1693–1703
Douma R, de Jonge L, Jonker CT, Seifar R, Heijnen J, van Gulik W (2010) Intracellular metabolite determination in the presence of extracellular abundance: application to the penicillin biosynthesis pathway in Penicillium chrysogenum. Biotechnol Bioeng 107:105–115
Wu L, Mashego M, van Dam J, Proell A, Vinke J, Ras C, van Winden W, van Gulik W, Heijnen J (2005) Quantitative analysis of the microbial metabolome by isotope dilution mass spectrometry using uniformly C-13-labeled cell extracts as internal standards. Anal Biochem 336:164–171
Pang Z, Zhou G, Ewald J, Chang L, Hacariz O, Basu N, **a J (2022) Using MetaboAnalyst 5.0 for LC–HRMS spectra processing, multi-omics integration and covariate adjustment of global metabolomics data. Nat Protoc 17:1735–1761
Hong M, Huang M, Chu J, Zhuang Y, Zhang S (2016) Impacts of proline on the central metabolism of an industrial erythromycin-producing strain Saccharopolyspora erythraea via (13)C labeling experiments. J Biotechnol 231:1–8
Mckenna M, Rae C (2015) A new role for α-ketoglutarate dehydrogenase complex: regulating metabolism through post-translational modification of other enzymes. J Neurochem 134(1):3–6
Ke X, Jiang X, Huang M, Tian X, Chu J (2022) Engineering of succinyl-CoA metabolism in view of succinylation regulation to improve the erythromycin production. Appl Microbiol Biotechnol 106(13–16):5153–5165
Li X, Chen C, Zhu F, Li Y (2005) Optimization of producing erythromycin and transforming its efficient components. Chin J Antibiot 02:65–69
Elmahdi I, Baganz F, Dixon K, Harrop T, Sugden D, Lye G (2003) pH control in microwell fermentations of S. erythraea CA340: influence on biomass growth kinetics and erythromycin biosynthesis. Biochem Eng J 16(3):299–310
Maeda RN, Silva MMPD, Anna LMMS Jr (2010) Nitrogen source optimization for cellulase production by Penicillium funiculosum, using a sequential experimental design methodology and the desirability function. Appl Biochem Biotechnol 161(1–8):411–422
Reeves A, Brikun I, Cernota W, Leach B, Gonzalez M, Weber JM (2007) Engineering of the methylmalonyl-CoA metabolite node of Saccharopolyspora erythraea for increased erythromycin production. Metab Eng 9(3):293–303
Jung WS, Yoo YJ, Park JW, Park SR, Han AR, Ban YH, Kim EJ, Kim E, Yoon YJ (2011) A combined approach of classical mutagenesis and rational metabolic engineering improves rapamycin biosynthesis and provides insights into methylmalonyl-CoA precursor supply pathway in Streptomyces hygroscopicus ATCC 29253. Appl Microbiol Biotechnol 91(5):1389–1397
Lee WH, Chin YW, Han NS, Kim MD, Seo JH (2011) Enhanced production of GDP-L-fucose by overexpression of NADPH regenerator in recombinant Escherichia coli. Appl Microbiol Biotechnol 91(4):967–976
Acknowledgements
This work was financially supported by the National Key Research and Development Program of China (Grant No. 2019YFA0904300), National Natural Science Foundation of China (Grant No. 32071461), and the National Key Research and Development Program of China (Grant No. 2018YFA0900300).
Author information
Authors and Affiliations
Contributions
Conceptualization: [Mingzhi Huang, Ju Chu]; methodology: [Yujie Yuan, Feng Xu]; formal analysis and investigation: [Yujie Yuan, **ang Ke, Ju Lu, Feng Xu]; writing—original draft preparation: [Yujie Yuan, Feng Xu]; writing—review and editing: [Feng Xu, Mingzhi Huang, Ju Chu]; funding acquisition: [Mingzhi Huang, Ju Chu]; supervision: [Mingzhi Huang, Ju Chu].
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yuan, Y., Xu, F., Ke, X. et al. Ammonium sulfate supplementation enhances erythromycin biosynthesis by augmenting intracellular metabolism and precursor supply in Saccharopolyspora erythraea. Bioprocess Biosyst Eng 46, 1303–1318 (2023). https://doi.org/10.1007/s00449-023-02898-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-023-02898-x