Abstract
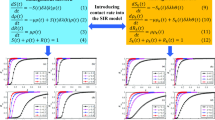
Communities are commonly not isolated but interact asymmetrically with each other, allowing the propagation of infectious diseases within the same community and between different communities. To reveal the impact of asymmetrical interactions and contact heterogeneity on disease transmission, we formulate a two-community SIR epidemic model, in which each community has its contact structure while communication between communities occurs through temporary commuters. We derive an explicit formula for the basic reproduction number \({\mathcal {R}}_0\), give an implicit equation for the final epidemic size z, and analyze the relationship between them. Unlike the typical positive correlation between \({\mathcal {R}}_0\) and z in the classic SIR model, we find a negatively correlated relationship between counterparts of our model deviating from homogeneous populations. Moreover, we investigate the impact of asymmetric coupling mechanisms on \({\mathcal {R}}_0\). The results suggest that, in scenarios with restricted movement of susceptible individuals within a community, \({\mathcal {R}}_0\) does not follow a simple monotonous relationship, indicating that an unbending decrease in the movement of susceptible individuals may increase \({\mathcal {R}}_0\). We further demonstrate that network contacts within communities have a greater effect on \({\mathcal {R}}_0\) than casual contacts between communities. Finally, we develop an epidemic model without restriction on the movement of susceptible individuals, and the numerical simulations suggest that the increase in human flow between communities leads to a larger \({\mathcal {R}}_0\).















Similar content being viewed by others
Data availibility
This manuscript has no associated data.
References
Amini H, Minca A (2022) Epidemic spreading and equilibrium social distancing in heterogeneous networks. Dyn Games Appl 1–30
Anderson RM, May RM (1992) Infectious diseases of humans: dynamics and control. Oxford University Press, Oxford
Ball F, Neal P (2002) A general model for stochastic SIR epidemics with two levels of mixing. Math Biosci 180(1–2):73–102
Barthélemy M, Barrat A, Pastor-Satorras R et al (2005) Dynamical patterns of epidemic outbreaks in complex heterogeneous networks. J Theor Biol 235(2):275–288
Bidari S, Chen X, Peters D et al (2016) Solvability of implicit final size equations for SIR epidemic models. Math Biosci 282:181–190
Cadoni M (2020) How to reduce epidemic peaks kee** under control the time-span of the epidemic. Chaos Solitons Fractals 138:109940
Cui J, Wu Y, Guo S (2022) Effect of non-homogeneous mixing and asymptomatic individuals on final epidemic size and basic reproduction number in a meta-population model. Bull Math Biol 84(3):1–22
Diekmann O (1978) Thresholds and travelling waves for the geographical spread of infection. J Math Biol 6(2):109–130
Diekmann O, De Jong MCM, Metz JAJ (1998) A deterministic epidemic model taking account of repeated contacts between the same individuals. J Appl Probab 35(2):448–462
El Sayed NM, Gomatos PJ, Beck-Sagué CM et al (2000) Epidemic transmission of human immunodeficiency virus in renal dialysis centers in Egypt. J Infect Dis 181(1):91–97
Ferrari’s solution. https://en.wikipedia.org/wiki/Quartic_function#Ferraris_solution
Fitzgibbon WE, Morgan JJ, Webb GF et al (2019) Spatial models of vector-host epidemics with directed movement of vectors over long distances. Math Biosci 312:77–87
Fitzgibbon WE, Morgan JJ, Webb GF et al (2020) Analysis of a reaction-diffusion epidemic model with asymptomatic transmission. J Biol Syst 28(03):561–587
Graham M, House T (2014) Dynamics of stochastic epidemics on heterogeneous networks. J Math Biol 68(7):1583–1605
Großmann G, Backenköhler M, Wolf V (2021) Heterogeneity matters: contact structure and individual variation shape epidemic dynamics. PLoS ONE 16(7):e0250050
Hales S, Weinstein P, Souares Y et al (1999) El Niño and the dynamics of vectorborne disease transmission. Environ Health Perspect 107(2):99–102
Hethcote HW (2000) The mathematics of infectious diseases. SIAM Rev 42(4):599–653
** Z, Sun G, Zhu H (2014) Epidemic models for complex networks with demographics. Math Biosci Eng 11(6):1295
Kamp C (2010) Untangling the interplay between epidemic spread and transmission network dynamics. PLoS Comput Biol 6(11):e1000984
Kiss IZ, Green DM, Kao RR (2006) The effect of contact heterogeneity and multiple routes of transmission on final epidemic size. Math Biosci 203(1):124–136
Koch D, Illner R, Ma J (2013) Edge removal in random contact networks and the basic reproduction number. J Math Biol 67(2):217–238
Krasnosel’skii MA (1964) Positive solutions of operator equations. Nordhoff, Groningen
Lajmanovich A, Yorke JA (1976) A deterministic model for gonorrhea in a nonhomogeneous population. Math Biosci 28(3–4):221–236
Li CH, Tsai CC, Yang SY (2014) Analysis of epidemic spreading of an SIRS model in complex heterogeneous networks. Commun Nonlinear Sci Numer Simul 19(4):1042–1054
Lieberthal B, Soliman A, Wang S et al (2023) Epidemic spread on patch networks with community structure. Math Biosci 359:108996
Magal P, Seydi O, Webb GF (2016) Final size of an epidemic for a two-group SIR model. SIAM J Appl Math 76(5):2042–2059
Magal P, Seydi O, Webb GF (2018) Final size of a multi-group SIR epidemic model: irreducible and non-irreducible modes of transmission. Math Biosci 301:59–67
Meng X, Cai Z, Si S et al (2021) Analysis of epidemic vaccination strategies on heterogeneous networks: based on SEIRV model and evolutionary game. Appl Math Comput 403:126172
Miller JC, Slim AC, Volz EM (2012) Edge-based compartmental modelling for infectious disease spread. J R Soc Interface 9(70):890–906
Moreno Y, Pastor-Satorras R, Vespignani A (2002) Epidemic outbreaks in complex heterogeneous networks. Eur Phys J B Condens Matter Complex Syst 26(4):521–529
Mossong J, Hens N, Jit M et al (2008) Social contacts and mixing patterns relevant to the spread of infectious diseases. PLoS Med 5(3):e74
Muroya Y, Kuniya T (2014) Further stability analysis for a multi-group SIRS epidemic model with varying total population size. Appl Math Lett 38:73–78
Muroya Y, Enatsu Y, Kuniya T (2013) Global stability for a multi-group SIRS epidemic model with varying population sizes. Nonlinear Anal Real World Appl 14(3):1693–1704
Olinky R, Stone L (2004) Unexpected epidemic thresholds in heterogeneous networks: the role of disease transmission. Phys Rev E 70(3):030902
Pastor-Satorras R, Vespignani A (2001) Epidemic dynamics and endemic states in complex networks. Phys Rev E 63(6):066117
Rass L, Radcliffe J (2003) Spatial deterministic epidemics. American Mathematical Society
Thieme HR (1977) A model for the spatial spread of an epidemic. J Math Biol 4:337–351
Thieme HR (1979) On a class of Hammerstein integral equations. Manuscr Math 29:49–84
Thieme HR (2003) Mathematics in population biology. Princeton University Press, Princeton
Tilman D, Kareiva P (1998) Spatial ecology. Princeton University Press, Princeton
Turkyilmazoglu M (2021) Explicit formulae for the peak time of an epidemic from the SIR model. Phys D 422:132902
Van den Driessche P, Watmough J (2002) Reproduction numbers and sub-threshold endemic equilibria for compartmental models of disease transmission. Math Biosci 180(1–2):29–48
Velazquez O, Stetler HC, Avila C et al (1990) Epidemic transmission of enterically transmitted non-A, non-B hepatitis in Mexico, 1986–1987. JAMA 263(24):3281–3285
Wallis J, Lee DR (1999) Primate conservation: the prevention of disease transmission. Int J Primatol 20(6):803–826
Wang L, Dai G (2008) Global stability of virus spreading in complex heterogeneous networks. SIAM J Appl Math 68(5):1495–1502
Wang Y, ** Z, Yang Z et al (2012) Global analysis of an SIS model with an infective vector on complex networks. Nonlinear Anal Real World Appl 13(2):543–557
Wang Y, Cao J, Li X et al (2018) Edge-based epidemic dynamics with multiple routes of transmission on random networks. Nonlinear Dyn 91(1):403–420
Wang Y, Wei Z, Cao J (2020) Epidemic dynamics of influenza-like diseases spreading in complex networks. Nonlinear Dyn 101(3):1801–1820
Yorke JA (1967) Invariance for ordinary differential equations. Math Syst Theory 1(4):353–372
Zhang J, ** Z (2012) Epidemic spreading on complex networks with community structure. Appl Math Comput 219(6):2829–2838
Zhu G, Fu X, Chen G (2012) Spreading dynamics and global stability of a generalized epidemic model on complex heterogeneous networks. Appl Math Model 36(12):5808–5817
Acknowledgements
Yi Wang’s research is partially supported by the National Natural Science Foundation of China under Grants 12171443 and 11801532, and the Fundamental Research Funds for the Central Universities under Grant G1323523061. Guiquan Sun’s research is partially supported by the National Natural Science Foundation of China under Grants 12022113 and 12271314. Hao Wang’s research is partially supported by the Natural Sciences and Engineering Research Council of Canada under Discovery Grant RGPIN-2020-03911 and Accelerator Supplement Award RGPAS-2020-00090. The authors thank the handling editor and the two anonymous referees for helpful comments and suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
We can obtain the roots of the quaternion equation (3.10) using the Ferrari’s solution (see, https://en.wikipedia.org/wiki/Quartic_function#Ferraris_solution for the exact solution procedure):
where
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, Z., Wang, Y., Gao, S. et al. Final epidemic size of a two-community SIR model with asymmetric coupling. J. Math. Biol. 88, 51 (2024). https://doi.org/10.1007/s00285-024-02073-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00285-024-02073-0