Abstract
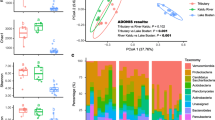
The surface water is an important habitat for freshwater microorganisms, but there is a lack of understanding of the pattern of microbial diversity and structure in stream continuums of small subtropical forest watersheds. Therefore, this study aimed to understand the variations in microbial diversity and community structure along stream orders (1–5) in the small subtropical forest catchments of the Wuyi Mountains. Using GIS software, 20 streams were chosen and classified into 5 orders. Illumina sequencing was used to analyze the dynamics of microbial communities, along with stream orders and hydro-chemical properties of stream water were also determined. Our results indicated that the bacterial and fungal richness (ACE index) was higher in low-order (1 and 2 orders) streams than in high-order (3, 4, and 5 orders) streams, with the highest value in the order 2 streams (P < 0.05). The water temperature and dissolved oxygen were positively correlated with fungal richness (P < 0.05). The bacterial rare taxa had a significant correlation with the abundance taxa (P < 0.05). The relative abundances of Bacteroidetes, Actinobacteria, and Chytridiomycota microbial phyla were significantly different among different order streams (P < 0.05). Using the neutral community model, we found that the fungal community structure was significantly shaped by hydro-chemical properties, while the bacterial community structure was largely regulated by stochastic processes. Our findings suggest that variations in microbial community structure in subtropical headwaters are largely shaped by the water temperature and dissolved oxygen.







Similar content being viewed by others
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Palmer M, Ruhi A (2019) Linkages between flow regime, biota, and ecosystem processes: implications for river restoration. Science 365:1264. https://doi.org/10.1126/science.aaw2087
Cavaco MA, St Louis VL, Engel K, St Pierre KA, Schiff SL, Stibal M, Neufeld JD (2019) Freshwater microbial community diversity in a rapidly changing High Arctic watershed. FEMS Microbiol Ecol 95:fiz161. https://doi.org/10.1093/femsec/fiz161
Zhou SL, Sun Y, Huang TL, Cheng Y, Yang X, Zhou ZZ, Li Y, Li ZX, Cui JS, **ao L (2020) Reservoir water stratification and mixing affects microbial community structure and functional community composition in a stratified drinking reservoir. J Environ Manage 267:110456. https://doi.org/10.1016/j.jenvman.2020.110456
Hassell N, Tinker KA, Moore T, Ottesen EA (2018) Temporal and spatial dynamics in microbial community composition within a temperate stream network. Environ Microbiol 20:3560–3572. https://doi.org/10.1111/1462-2920.14311
Akinwole PO, Kan JJ, Kaplan LA, Findlay RH (2021) Spatial variability in streambed microbial community structure across two watersheds. Microbiol Spectr 9:e0197221. https://doi.org/10.1128/Spectrum.01972-21
Gweon HS, Bowes MJ, Moorhouse HL, Oliver AE, Bailey MJ, Acreman MC, Read DS (2021) Contrasting community assembly processes structure lotic bacteria metacommunities along the river continuum. Environ Microbiol 23:484–498. https://doi.org/10.1111/1462-2920.15337
Nyirabuhoro P, Liu M, **ao P, Liu L, Yu Z, Wang L, Yang J (2020) Seasonal variability of conditionally rare taxa in the water column bacterioplankton community of subtropical reservoirs in China. Microb Ecol 80:14–26. https://doi.org/10.1007/s00248-019-01458-9
Jones SE, Lennon JT (2010) Dormancy contributes to the maintenance of microbial diversity. P Natl Acad Sci Usa 107:5881–5886. https://doi.org/10.1073/pnas.0912765107
Zeglin LH (2015) Stream microbial diversity in response to environmental changes: review and synthesis of existing research. Front Microbiol 6:454. https://doi.org/10.3389/fmicb.2015.00454
Chen W, Ren K, Isabwe A, Chen H, Liu M, Yang J (2019) Stochastic processes shape microeukaryotic community assembly in a subtropical river across wet and dry seasons. Microbiome 7:1. https://doi.org/10.1186/s40168-019-0749-8
Tang X, **e G, Shao K, Hu Y, Cai J, Bai C, Gong Y, Gao G (2020) Contrast diversity patterns and processes of microbial community assembly in a river-lake continuum across a catchment scale in northwestern China. Environ Microbiome 15:13. https://doi.org/10.1186/s40793-020-00356-9
Guo K, Wu N, Li W, Baattrup-Pedersen A, Riis T (2021) Microbial biofilm community dynamics in five lowland streams. Sci Total Environ 798:149169. https://doi.org/10.1016/j.scitotenv.2021.149169
Artigas J, Gaudes A, Munoz I, Romani AM, Sabater S (2011) Fungal and bacterial colonization of submerged leaf litter in a mediterranean stream. Int Rev Hydrobiol 96:221–234. https://doi.org/10.1002/iroh.201111355
Fenoy E, Pradhan A, Pascoal C, Rubio-Rios J, Batista D, Moyano-Lopez FJ, Cassio F, Casas JJ (2022) Elevated temperature may reduce functional but not taxonomic diversity of fungal assemblages on decomposing leaf litter in streams. Global Change Biol 28:115–127. https://doi.org/10.1111/gcb.15931
Krause S, Le Roux X, Niklaus PA, Van Bodegom PM, Lennon JT, Bertilsson S, Grossart H-P, Philippot L, Bodelier PLE (2014) Trait-based approaches for understanding microbial biodiversity and ecosystem functioning. Front Microbiol 5:251. https://doi.org/10.3389/fmicb.2014.00251
Staley C, Gould TJ, Wang P, Phillips J, Cotner JB, Sadowsky MJ (2015) Species sorting and seasonal dynamics primarily shape bacterial communities in the Upper Mississippi River. Sci Total Environ 505:435–445. https://doi.org/10.1016/j.scitotenv.2014.10.012
Brandani J, Peter H, Fodelianakis S, Kohler TJ, Bourquin M, Michoud G, Busi SB, Ezzat L, Lane S, Battin TJ (2023) Homogeneous environmental selection structures the bacterial communities of benthic biofilms in proglacial floodplain streams. Appl Environ Microbiol 89:e02010–e02022. https://doi.org/10.1128/aem.02010-22
Nan** People’s Government of Fujian Province (2022) Nan** Wuyi Mountain National Park Protection and Development Belt Water Resources White Paper. https://www.np.gov.cn/cms/siteresource/article.shtml?id=430559290014820000&siteId=340328840390080000
Strahler AN (1957) Quantitative analysis of watershed geomorphology. Eos Trans Am Geophys Union 38:913–920
Finn DS, Bonada N, Murria C, Hughes JM (2011) Small but mighty: headwaters are vital to stream network biodiversity at two levels of organization. J N Am Benthol Soc 30:963–980. https://doi.org/10.1899/11-012.1
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. P Natl Acad Sci Usa 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Wagg C, Schlaeppi K, Banerjee S, Kuramae EE, van der Heijden MGA (2019) Fungal-bacterial diversity and microbiome complexity predict ecosystem functioning. Nat Commun 10:4841. https://doi.org/10.1038/s41467-019-12798-y
Fujita SI, Senda Y, Nakaguchi S, Hashimoto T (2001) Multiplex PCR using internal transcribed spacer 1 and 2 regions for rapid detection and identification of yeast strains. J Clin Microbiol 39:3617–3622. https://doi.org/10.1128/jcm.39.10.3617-3622.2001
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:884–890. https://doi.org/10.1093/bioinformatics/bty560
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. https://doi.org/10.1093/bioinformatics/btr507
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072. https://doi.org/10.1128/aem.03006-05
Kaiser C, Kilburn MR, Clode PL, Fuchslueger L, Koranda M, Cliff JB, Solaiman ZM, Murphy DV (2015) Exploring the transfer of recent plant photosynthates to soil microbes: mycorrhizal pathway vs direct root exudation. New Phytol 205:1537–1551. https://doi.org/10.1111/nph.13138
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Gloeckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Nilsson RH, Larsson K-H, Taylor AFS, Bengtsson-Palme J, Jeppesen TS, Schigel D, Kennedy P, Picard K, Gloeckner FO, Tedersoo L, Saar I, Koljalg U, Abarenkov K (2019) The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res 47:D259–D264. https://doi.org/10.1093/nar/gky1022
Ding J, Tarokh V, Yang YH (2018) Model selection techniques an overview. IEEE Signal Process Mag 35:16–34. https://doi.org/10.1109/msp.2018.2867638
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP (2006) Quantifying the roles of immigration and chance in sha** prokaryote community structure. Environ Microbiol 8:732–740. https://doi.org/10.1111/j.1462-2920.2005.00956.x
Ning DL, Deng Y, Tiedje JM, Zhou JZ (2019) A general framework for quantitatively assessing ecological stochasticity. P Natl Acad Sci Usa 116:16892–16898. https://doi.org/10.1073/pnas.1904623116
Dai TJ, Zhang Y, Tang YS, Bai YH, Tao YL, Huang B, Wen DH (2016) Identifying the key taxonomic categories that characterize microbial community diversity using full-scale classification: a case study of microbial communities in the sediments of Hangzhou Bay. FEMS Microbiol Ecol 92:fiw150. https://doi.org/10.1093/femsec/fiw150
Teachey ME, McDonald JM, Ottesen EA (2019) Rapid and stable microbial community assembly in the headwaters of a third-order stream. Appl Environ Microbiol 85:e00188–e00119. https://doi.org/10.1128/aem.00188-19
de Oliveira LFV, Margis R (2015) The source of the river as a nursery for microbial diversity. PLoS One 10:e0120608. https://doi.org/10.1371/journal.pone.0120608
Miura A, Urabe J (2015) Spatial and seasonal changes in species diversity of epilithic fungi along environmental gradients of a river. Freshwat Biol 60:673–685. https://doi.org/10.1111/fwb.12514
Kominoski JS, Marczak LB, Richardson JS (2011) Riparian forest composition affects stream litter decomposition despite similar microbial and invertebrate communities. Ecology 92:151–159. https://doi.org/10.1890/10-0028.1
Chen J, Wang PF, Wang C, Wang X, Miao LZ, Liu S, Yuan QS, Sun SH (2020) Fungal community demonstrates stronger dispersal limitation and less network connectivity than bacterial community in sediments along a large river. Environ Microbiol 22:832–849. https://doi.org/10.1111/1462-2920.14795
David GM, Lopez-Garcia P, Moreira D, Alric B, Deschamps P, Bertolino P, Restoux G, Rochelle-Newall E, Thebault E, Simon M, Jardillier L (2021) Small freshwater ecosystems with dissimilar microbial communities exhibit similar temporal patterns. Mol Ecol 30:2162–2177. https://doi.org/10.1111/mec.15864
Ouellet V, Daniels MD, Peipoch M, Zgleszewski L, Watson N, Gibson E, Krause S, Kan JJ (2022) Beyond the light effect: How hydrologic and geomorphologic stream features control microbial distribution across pool sequences in a temperate headwater stream. Ecohydrology 15:e2380. https://doi.org/10.1002/eco.2380
Newton RJ, Jones SE, Eiler A, McMahon KD, Bertilsson S (2011) A Guide to the Natural History of Freshwater Lake Bacteria. Microbiol Mol Biol Rev 75:14–49. https://doi.org/10.1128/mmbr.00028-10
Seena S, Barlocher F, Sobral O, Gessner MO, Dudgeon D, McKie BG, Chauvet E, Boyero L, Ferreira V, Frainer A, Bruder A, Matthaei CD, Fenoglio S, Sridhar KR, Albarino RJ, Douglas MM, Encalada AC, Garcia E, Ghate SD et al (2019) Biodiversity of leaf litter fungi in streams along a latitudinal gradient. Sci Total Environ 661:306–315. https://doi.org/10.1016/j.scitotenv.2019.01.122
Zhang S, Li K, Hu J, Wang F, Chen D, Zhang Z, Li T, Li L, Tao J, Liu D, Che R (2022) Distinct assembly mechanisms of microbial sub-communities with different rarity along the Nu River. J Soils Sed 22:1530–1545. https://doi.org/10.1007/s11368-022-03149-4
Drew GC, Stevens EJ, King KC (2021) Microbial evolution and transitions along the parasite-mutualist continuum. Nat Rev Microbiol 19:623–638. https://doi.org/10.1038/s41579-021-00550-7
Hernandez DJ, David AS, Menges ES, Searcy CA, Afkhami ME (2021) Environmental stress destabilizes microbial networks. Isme J 15:1722–1734. https://doi.org/10.1038/s41396-020-00882-x
Song WW, Zhang LY, Li Y, Zhang WL, Wang LF, Niu LH, Zhang HJ, Ji Y, Liao ZY (2022) Hydrodynamic zones and the influence of microorganisms on nitrogen transformation in the diverging area of branched rivers. Environ Res 208:112778. https://doi.org/10.1016/j.envres.2022.112778
Roldan DM, Carrizo D, Sanchez-Garcia L, Menes RJ (2022) Diversity and effect of increasing temperature on the activity of methanotrophs in sediments of Fildes Peninsula Freshwater Lakes, King George Island, Antarctica. Front Microbiol 13:822552. https://doi.org/10.3389/fmicb.2022.822552
Duarte S, Barlocher F, Trabulo J, Cassio F, Pascoal C (2015) Stream-dwelling fungal decomposer communities along a gradient of eutrophication unraveled by 454 pyrosequencing. Fungal Divers 70:127–148. https://doi.org/10.1007/s13225-014-0300-y
Szabo-Taylor KE, Kiss KT, Logares R, Eiler A, Acs E, Toth B, Bertilsson S (2010) Composition and dynamics of microeukaryote communities in the River Danube. Fottea 10:99–113. https://doi.org/10.5507/fot.2010.005
Gad M, Hou LY, Li JW, Wu Y, Rashid A, Chen NW, Hu AY (2020) Distinct mechanisms underlying the assembly of microeukaryotic generalists and specialists in an anthropogenically impacted river. Sci Total Environ 748:141434. https://doi.org/10.1016/j.scitotenv.2020.141434
Acknowledgements
We thank Changshan ** and Wei He for the logistic support, and the anonymous reviewers for the valuable suggestions.
Funding
This work was supported by Fujian Forestry Science and Technology Project (grant nos: 2018-26 and 2020-29) and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
Investigation, conceptualization, visualization, formal analysis, and writing original draft by Boran Liu. Investigation, methodology, visualization, and formal analysis by Boran Liu and Yuchao Wang. Funding acquisition and project administration by Huiguang Zhang, Yan Zhou, and Chenhui Zhang. Writing review and editing by Nan Yang and Weifeng Wang. Conceptualization, supervision, and writing—review and editing—by Weifeng Wang. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Consent to Participate
Informed consent was obtained from all individual participants included in the study.
Consent to Publish
The authors affirm that all participants consent to publish manuscript
Competing Interests
The authors declare no competing interests.
Supplementary Information
ESM 1
(DOCX 164 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, B., Wang, Y., Zhang, H. et al. The Variations of Microbial Diversity and Community Structure Along Different Stream Orders in Wuyi Mountains. Microb Ecol 86, 2330–2343 (2023). https://doi.org/10.1007/s00248-023-02240-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02240-8