Abstract
Type III secretion (T3S) systems are complex bacterial structures used by many pathogens to inject proteins directly into the cytosol of the host cell. These secretion machines evolved from the bacterial flagella and they have been grouped into families by phylogenetic analysis. The T3S system is composed of more than 20 proteins grouped into five complexes: the cytosolic platform, the export apparatus, the basal body, the needle, and the translocon complex. While the proteins located inside the bacterium are conserved, those exposed to the external media present high variability among families. This suggests that the T3S systems have adapted to interact with different cells or tissues in the host, and/or have been subjected to the evolutionary pressure of the host immune defenses. Such adaptation led to changes in the sequence of the T3S needle tip and translocon suggesting differences in the mechanism of assembly and structure of this complex.
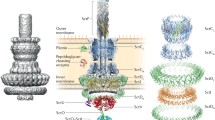
Graphical abstract


adapted from data of the Ysc family (Broz et al. 2007). PM host plasma membrane, OM outer membrane, IM inner membrane






Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Abby SS, Rocha EPC (2012) The Non-Flagellar Type III Secretion System Evolved from the bacterial flagellum and diversified into host-cell adapted systems. PLOS Genetics 8(9):e1002983
Adams W, Morgan J, Kwuan L, Auerbuch V (2015) Yersinia pseudotuberculosis YopD mutants that genetically separate effector protein translocation from host membrane disruption. Mol Microbiol 96:764–778
Akeda Y, Galan JE (2005) Chaperone release and unfolding of substrates in type III secretion. Nature 437:911–915
Akopyan K, Edgren T, Wang-Edgren H, Rosqvist R, Fahlgren A, Wolf-Watz H, Fallman M (2011) Translocation of surface-localized effectors in type III secretion. Proc Nat Acad Sci 108:1639–1644
Allison SE, Tuinema BR, Everson ES, Sugiman-Marangos S, Zhang K, Junop MS, Coombes BK (2014) Identification of the docking site between a type III secretion system ATPase and a chaperone for effector cargo. J Biol Chem 289:23734–23744
Barth H, Aktories K, Popoff MR, Stiles BG (2004) Binary bacterial toxins: biochemistry, biology, and applications of common Clostridium and Bacillus proteins. Microbiol Mol Biol Rev 68:373–402
Broz P, Mueller CA, Muller SA, Philippsen A, Sorg I, Engel A, Cornelis GR (2007) Function and molecular architecture of the Yersinia injectisome tip complex. Mol Microbiol 65:1311–1320
Campanella J, Bitincka L, Smalley J (2003) MatGAT: an application that generates similarity/identity matrices using protein or DNA sequences. BMC Bioinformatics 4:29
Chaudhury S, Battaile KP, Lovell S, Plano GV, De Guzman RN (2013) Structure of the Yersinia pestis tip protein LcrV refined to 1.65Å resolution. Acta Crystallogr Sect F Struct Biol Cryst Commun 69:477–481
Chen L, Ai X, Portaliou AG, Minetti CA, Remeta DP, Economou A, Kalodimos CG (2013) Substrate-activated conformational switch on chaperones encodes a targeting signal in type III secretion. Cell Rep 3:709–715
Cheung M, Shen D-K, Makino F, Kato T, Roehrich AD, Martinez-Argudo I, Walker ML, Murillo I, Liu X, Pain M, Brown J, Frazer G, Mantell J, Mina P, Todd T, Sessions RB, Namba K, Blocker AJ (2015) Three-dimensional electron microscopy reconstruction and cysteine-mediated crosslinking provide a model of the type III secretion system needle tip complex. Mol Microbiol 95:31–50
Cornelis GR (2002) The Yersinia Ysc-Yop “Type III” weaponry. Nat Rev Mol Cell Biol 3:742–754
Cornelis GR (2006) The type III secretion injectisome. Nat Rev Microbiol 4:811–825
Daniell SJ, Takahashi N, Wilson R, Friedberg D, Rosenshine I, Booy FP, Shaw RK, Knutton S, Frankel G, Aizawa S (2001) The filamentous type III secretion translocon of enteropathogenic Escherichia coli. Cell Microbiol 3:865–871
Delalez NJ, Wadhams GH, Rosser G, Xue Q, Brown MT, Dobbie IM, Berry RM, Leake MC, Armitage JP (2010) Signal-dependent turnover of the bacterial flagellar switch protein FliM. Proc Natl Acad Sci USA 107:11347–11351
Deng W, Marshall NC, Rowland JL, McCoy JM, Worrall LJ, Santos AS, Strynadka N, Finlay BB (2017) Assembly, structure, function and regulation of type III secretion systems. Nat rev Microbiol 15(6):323–337
Derewenda U, Mateja A, Devedjiev Y, Routzahn KM, Evdokimov AG, Derewenda ZS, Waugh DS (2004) The structure of Yersinia pestis V-Antigen, an Essential Virulence Factor and Mediator of immunity against plague. Structure 12:301–306
Dey S, Basu A, Datta S (2012) Characterization of molten globule PopB in absence and presence of its chaperone PcrH. Prot J 31:401–416
Diepold A (2020) Assembly and post-assembly turnover and dynamics in the type III secretion system. Curr Top Microbiol Immunol 427:35–66
Diepold A, Amstutz M, Abel S, Sorg I, Jenal U, Cornelis GR (2010) Deciphering the assembly of the Yersinia type III secretion injectisome. EMBO J 29:1928–1940
Diepold A, Kudryashev M, Delalez NJ, Berry RM, Armitage JP (2015) Composition, formation, and regulation of the cytosolic c-ring, a dynamic component of the type III secretion injectisome. PLoS Biol 13:e1002039
Dietsche T, Tesfazgi Mebrhatu M, Brunner MJ, Abrusci P, Yan J, Franz-Wachtel M, Scharfe C, Zilkenat S, Grin I, Galan JE, Kohlbacher O, Lea S, Macek B, Marlovits TC, Robinson CV, Wagner S (2016) Structural and functional characterization of the bacterial Type III secretion export apparatus. PLoS Pathog 12:e1006071
Discola KF, Forster A, Boulay F, Simorre JP, Attree I, Dessen A, Job V (2014) Membrane and chaperone recognition by the major translocator protein PopB of the type III secretion system of Pseudomonas aeruginosa. J Biol Chem 289:3591–3601
Dohlich K, Zumsteg AB, Goosmann C, Kolbe M (2014) A substrate-fusion protein is trapped inside the type III secretion system channel in Shigella flexneri. PLOS Pathogens 10:e1003881
Faudry E, Vernier G, Neumann E, Forge V, Attree I (2006) Synergistic pore formation by type III toxin translocators of Pseudomonas aeruginosa. Biochemistry 45:8117–8123
Faudry E, Job V, Dessen A, Attree I, Forge V (2007) Type III secretion system translocator has a molten globule conformation both in its free and chaperone-bound forms. FEBS Journal 274:3601–3610
Fujii T, Cheung M, Blanco A, Kato T, Blocker AJ, Namba K (2012) Structure of a type III secretion needle at 7-A resolution provides insights into its assembly and signaling mechanisms. Proc Natl Acad Sci USA 109:4461–4466
Gautier R, Douguet D, Antonny B, Drin G (2008) HELIQUEST: a web server to screen sequences with specific α-helical properties. Bioinformatics 24:2101–2102
Gebus C, Eric F, Bohn Y-ST, Sylvie E, Attree I (2008) Oligomerization of PcrV and LcrV, protective antigens of Pseudomonas aeruginosa and Yersinia pestis. J Biol Chem 283:23940–23949
Goure J, Pastor A, Faudry E, Chabert J, Dessen A, Attree I (2004) The V antigen of Pseudomonas aeruginosa is required for assembly of the functional PopB/PopD translocation pore in host cell membranes. Infect Immun 72:4741–4750
Goure J, Broz P, Attree O, Cornelis GR, Attree I (2005) Protective anti-V antibodies inhibit Pseudomonas and Yersinia translocon assembly within host membranes. J Infect Dis 192:218–225
Guo EZ, Galán JE (2021) Cryo-EM structure of the needle filament tip complex of the Salmonella type III secretion injectisome. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.2114552118
Habenstein B, El Mammeri N, Tolchard J, Lamon G, Tawani A, Berbon M, Loquet A (2020) Structures of type III secretion system needle filaments. Curr Top Microbiol Immunol 427:109–131
Harmon DE, Murphy JL, Davis AJ, Mecsas J (2013) A mutant with aberrant extracellular LcrV-YscF interactions fails to form pores and translocate Yop effector proteins but retains the ability to trigger Yop secretion in response to host cell contact. J Bacteriol 195:2244–2254
Hu B, Morado DR, Margolin W, Rohde JR, Arizmendi O, Picking WL, Picking WD, Liu J (2015) Visualization of the type III secretion sorting platform of Shigella flexneri. Proc Nat Acad Sci 112:1047–1052
Hu B, Lara-Tejero M, Kong Q, Galán JE, Liu J (2017a) In situ molecular architecture of the Salmonella Type III secretion machine. Cell 168:1065–1074
Hu Y, Huang H, Cheng X, Shu X, White AP, Stavrinides J, Köster W, Zhu G, Zhao Z, Wang Y (2017b) A global survey of bacterial type III secretion systems and their effectors. Environ Microbiol 19:3879–3895
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589
Kenjale R, Wilson J, Zenk SF, Saurya S, Picking WL, Picking WD, Blocker A (2005) The needle component of the type III secreton of Shigella regulates the activity of the secretion apparatus. J Biol Chem 280:42929–42937
Kim JF (2001) Revisiting the chlamydial type III protein secretion system: clues to the origin of type III protein secretion. Trends Genet 17:65–69
Kudryashev M, Stenta M, Schmelz S, Amstutz M, Wiesand U, Castaño-Díez D, Degiacomi MT, Münnich S, Bleck CKE, Kowal J, Diepold A, Heinz DW, Dal Peraro M, Cornelis GR, Stahlberg H (2013) In situ structural analysis of the Yersinia enterocolitica injectisome. eLife 2:e00792
Kuhlen L, Abrusci P, Johnson S, Gault J, Deme J, Caesar J, Dietsche T, Mebrhatu MT, Ganief T, Macek B, Wagner S, Robinson CV, Lea SM (2018) Structure of the core of the type III secretion system export apparatus. Nat Struct Mol Biol 25:583–590
Lara-Tejero M (2020) The Type III secretion system sorting platform. Curr Top Microbiol Immunol 427:133–142
Lara-Tejero M, Kato J, Wagner S, Liu X, Galan JE (2011) A sorting platform determines the order of protein secretion in bacterial type III systems. Science 331:1188–1191
Lee PC, Zmina SE, Stopford CM, Toska J, Rietsch A (2014) Control of type III secretion activity and substrate specificity by the cytoplasmic regulator PcrG. Proc Natl Acad Sci USA 111:E2027-2036
Lombardi C, Tolchard J, Bouillot S, Signor L, Gebus C, Liebl D, Fenel D, Teulon J-M, Brock J, Habenstein B, Pellequer J-L, Faudry E, Loquet A, Attrée I, Dessen A, Job V (2019) Structural and functional characterization of the type three secretion system (T3SS) needle of Pseudomonas aeruginosa. Front Microbiol. https://doi.org/10.3389/fmicb.2019.00573
London E, Shahidullah K (2009) Transmembrane vs. non-transmembrane hydrophobic helix topography in model and natural membranes. Curr Opin Struct Biol 19:464–472
Loquet A, Sgourakis NG, Gupta R, Giller K, Riedel D, Goosmann C, Griesinger C, Kolbe M, Baker D, Becker S, Lange A (2012) Atomic model of the type III secretion system needle. Nature 486:276–279
Makino F, Shen D, Kajimura N, Kawamoto A, Pissaridou P, Oswin H, Pain M, Murillo I, Namba K, Blocker AJ (2016) The architecture of the cytoplasmic region of Type III secretion systems. Sci Rep 6:33341
Man P, Montagner C, Vitrac H, Kavan D, Pichard S, Gillet D, Forest E, Forge V (2010) Accessibility changes within diphtheria toxin T domain when in the functional molten globule state, as determined using hydrogen/deuterium exchange measurements. FEBS J 277:653–662
Matteï P-J, Faudry E, Job V, Izoré T, Attree I, Dessen A (2011) Membrane targeting and pore formation by the type III secretion system translocon. FEBS Journal 278:414–426
Miletic S, Fahrenkamp D, Goessweiner-Mohr N, Wald J, Pantel M, Vesper O, Kotov V, Marlovits TC (2021) Substrate-engaged type III secretion system structures reveal gating mechanism for unfolded protein translocation. Nat Commun 12:1546
Minamino T, Kawamoto A, Kinoshita M, Namba K (2020) Molecular organization and assembly of the export apparatus of flagellar type III secretion systems. Curr Top Microbiol Immunol 427:91–107
Mueller CA, Broz P, Muller SA, Ringler P, Erne-Brand F, Sorg I, Kuhn M, Engel A, Cornelis GR (2005) The V-antigen of Yersinia forms a distinct structure at the tip of injectisome needles. Science 310:674–676
Mueller CA, Broz P, Cornelis GR (2008) The type III secretion system tip complex and translocon. Mol Microbiol 68:1085–1095
Notredame C, Higgins DG, Heringa J (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217
Pastor A, Chabert J, Louwagie M, Garin J, Attree I (2005) PscF is a major component of the Pseudomonas aeruginosa type III secretion needle. FEMS Microbiol Lett 253:95–101
Petty NK, Bulgin R, Crepin VF, Cerdeño-Tárraga AM, Schroeder GN, Quail MA, Lennard N, Corton C, Barron A, Clark L, Toribio AL, Parkhill J, Dougan G, Frankel G, Thomson NR (2010) The Citrobacter rodentium genome sequence reveals convergent evolution with human pathogenic Escherichia coli. J Bacteriol 192:525–538
Poklar N, Lah J, Salobir M, Macek P, Vesnaver G (1997) pH and temperature-induced molten globule-like denatured states of equinatoxin II: a study by UV-melting, DSC, far- and near-UV CD spectroscopy, and ANS fluorescence. Biochemistry 36:14345–14352
Radics J, Königsmaier L, Marlovits TC (2014) Structure of a pathogenic type 3 secretion system in action. Nat Struct Mol Biol 21:82–87
Rathinavelan T, Zhang L, Picking WL, Weis DD, De Guzman RN, Im W (2010) A repulsive electrostatic mechanism for protein export through the Type III secretion apparatus. Biophys J 98:452–461
Roblin P, Dewitte F, Villeret V, Biondi EG, Bompard C (2015) A Salmonella type three secretion effector/chaperone complex adopts a hexameric ring-like structure. J Bacteriol 197:688–698
Romano FB, Rossi KC, Savva CG, Holzenburg A, Clerico EM, Heuck AP (2011) Efficient isolation of Pseudomonas aeruginosa type III secretion translocators and assembly of heteromeric transmembrane pores in model membranes. Biochemistry 50:7117–7131
Romano FB, Tang Y, Rossi KC, Monopoli KR, Ross JL, Heuck AP (2016) Type 3 secretion translocators spontaneously assemble a hexadecameric transmembrane complex. J Biol Chem 291:6304–6315
Russo BC, Duncan JK, Goldberg MB (2019) Topological analysis of the type 3 secretion system translocon pore protein IpaC following its native delivery to the plasma membrane during infection. mBio 10:e00877-00819
Sawa T, Katoh H, Yasumoto H (2014) V-antigen homologs in pathogenic gram-negative bacteria. Microbiol Immunol 58:267–285
Schmidt H, Hensel M (2004) Pathogenicity islands in bacterial pathogenesis. Clin Microbiol Rev 17:14–56
Simossis VA, Kleinjung J, Heringa J (2005) Homology-extended sequence alignment. Nucl Acids Res 33:816–824
Sundin C, Thelaus J, Broms JE, Forsberg A (2004) Polarisation of type III translocation by Pseudomonas aeruginosa requires PcrG, PcrV and PopN. Microb Pathog 37:313–322
Tabor DE, Oganesyan V, Keller AE, Yu L, McLaughlin RE, Song E, Warrener P, Rosenthal K, Esser M, Qi Y, Ruzin A, Stover CK, DiGiandomenico A (2018) Pseudomonas aeruginosa PcrV and Psl, the molecular targets of bispecific antibody MEDI3902, are conserved among diverse global clinical isolates. J Infect Dis 218:1983–1994
Tan YW, Yu HB, Sivaraman J, Leung KY, Mok Y-K (2009) Map** of the chaperone AcrH binding regions of translocators AopB and AopD and characterization of oligomeric and metastable AcrH-AopB-AopD complexes in the type III secretion system of Aeromonas hydrophila. Protein Sci 18:1724–1734
Tang Y, Romano FB, Breña M, Heuck AP (2018) The Pseudomonas aeruginosa type III secretion translocator PopB assists the insertion of the PopD translocator into host cell membranes. J Biol Chem 293:8982–8993
Tang Y, Guo H, Vermeulen AJ, Heuck AP (2021) Topological analysis of type 3 secretion translocons in native membranes. Methods Enzymol 649:397–429
Tejeda-Dominguez F, Huerta-Cantillo J, Chavez-Dueñas L, Navarro-Garcia F (2017) A novel mechanism for protein delivery by the Type 3 Secretion system for extracellularly secreted proteins. mBio 8(2):e00184-00117
Thomas DR, Morgan DG, DeRosier DJ (1999) Rotational symmetry of the C ring and a mechanism for the flagellar rotary motor. Proc Natl Acad Sci USA 96:10134–10139
Troisfontaines P, Cornelis GR (2005) Type III secretion: more systems than you think. Physiology (Bethesda, Md.) 20:326–339
van der Goot FG, Gonzalez-Manas JM, Lakey JH, Pattus F (1991) A “molten-globule” membrane-insertion intermediate of the pore-forming domain of colicin A. Nature 354:408–410
Van Nguyen, S, Jobichen C, Tan Kang W, Tan Yih W, Chan Siew L, Ramesh K, Yuan Y, Hong Y, Seetharaman J, Leung Ka Y, Sivaraman J, Mok YuK (2015) Structure of AcrH–AopB chaperone-translocator complex reveals a role for membrane hairpins in type III secretion system translocon assembly. Structure 23:2022–2031
Vanden Bergh P, Frey J (2014) Aeromonas salmonicida subsp. salmonicida in the light of its type-three secretion system. Microbial Biotechnol 7:381–400
Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, Yuan D, Stroe O, Wood G, Laydon A, Žídek A, Green T, Tunyasuvunakool K, Petersen S, Jumper J, Clancy E, Green R, Vora A, Lutfi M, Figurnov M, Cowie A, Hobbs N, Kohli P, Kleywegt G, Birney E, Hassabis D, Velankar S (2022) AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res 50:D439-d444
Vermeulen AJ, Tang Y, Heuck AP (2016) Translocation of toxins by gram-negative pathogens using the type III secretion system. In: Gopalakrishnakone P, Stiles B, Alape-Girón A, Dubreuil JD, Mandal M (eds) Microbial Toxins. Springer, Dordrecht, pp 1–18
Wagner S, Diepold A (2020) A Unified nomenclature for injectisome-Type Type III secretion systems. Curr Top Microbiol Immunol 427:1–10
Wagner S, Konigsmaier L, Lara-Tejero M, Lefebre M, Marlovits TC, Galan JE (2010) Organization and coordinated assembly of the type III secretion export apparatus. Proc Natl Acad Sci USA 107:17745–17750
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25:1189–1191
Worrall LJ, Hong C, Vuckovic M, Deng W, Bergeron JR, Majewski DD, Huang RK, Spreter T, Finlay BB, Yu Z, Strynadka NC (2016) Near-atomic-resolution cryo-EM analysis of the Salmonella T3S injectisome basal body. Nature. https://doi.org/10.1038/nature20576
Worrall LJ, Hu J, Strynadka NCJ (2020) Aligning the symmetry of the Type III secretion system needle complex. J Chem Inform Model 60:2430–2435
Zarivach R, Vuckovic M, Deng W, Finlay BB, Strynadka NC (2007) Structural analysis of a prototypical ATPase from the type III secretion system. Nat Struct Mol Biol 14:131–137
Zilkenat S, Franz-Wachtel M, Stierhof YD, Galan JE, Macek B, Wagner S (2016) Determination of the stoichiometry of the complete bacterial type III secretion needle complex using a combined quantitative proteomic approach. Mol Cell Proteomics 15:1598–1609
Acknowledgments
We thank former and current members of the Heuck lab for their contributions and insightful discussions and Dr. Eugenia M. Clerico for critically reading of the manuscript and design of the figures. We also thank the UMass College of Natural Sciences Dean’s Bridge and Seed Fund support and the UMass Faculty Research Grant/Healey Endowment Grant (to AH).
Funding
This work was supported by the National Institutes of Health Grant No. R41 AI149922.
Author information
Authors and Affiliations
Contributions
AH wrote the main manuscript, prepared Figures 3 and 4, performed the amino acid identity calculations, performed the multisequence alignments, and did the conservation calculations. MB retrieved protein sequences from GenBank, prepared Caption Abstract and Table S2. AH and MB analyzed the data and prepared the rest of the figures and tables. Both authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Heuck, A.P., Brovedan, M.A. Evolutionary Conservation, Variability, and Adaptation of Type III Secretion Systems. J Membrane Biol 255, 599–612 (2022). https://doi.org/10.1007/s00232-022-00247-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-022-00247-9