Abstract
Background
Bacterial communities are essential to the biogeochemical cycle in riverine ecosystems. However, little is presently known about the integrated biogeography of planktonic and sedimentary bacterial communities in large rivers.
Results
This study provides the first spatiotemporal pattern of bacterial communities in the Yangtze River, the largest river in Asia with a catchment area of 1,800,000 km2. We find that sedimentary bacteria made larger contributions than planktonic bacteria to the bacterial diversity of the Yangzte River ecosystem with the sediment subgroup providing 98.8% of 38,906 operational taxonomic units (OTUs) observed in 280 samples of synchronous flowing water and sediment at 50 national monitoring stations covering a 4300 km reach. OTUs within the same phylum displayed uniform seasonal variations, and many phyla demonstrated autumn preference throughout the length of the river. Seasonal differences in bacterial communities were statistically significant in water, whereas bacterial communities in both water and sediment were geographically clustered according to five types of landforms: mountain, foothill, basin, foothill-mountain, and plain. Interestingly, the presence of two huge dams resulted in a drastic fall of bacterial taxa in sediment immediately downstream due to severe riverbed scouring. The integrity of the biogeography is satisfactorily interpreted by the combination of neutral and species sorting perspectives in meta-community theory for bacterial communities in flowing water and sediment.
Conclusions
Our study fills a gap in understanding of bacterial communities in one of the world’s largest river and highlights the importance of both planktonic and sedimentary communities to the integrity of bacterial biogeographic patterns in a river subject to varying natural and anthropogenic impacts.
Similar content being viewed by others
Background
Rivers play an important part in coupling biogeochemical cycles between continents and oceans [1]. River water flow acts as a carrier of biotic and abiotic substances, whereas sediment serves as a sink or source in the cycling of nutrients. Previous studies on riverine biodiversity mostly focused on invertebrates or fish, and very limited reports were related to bacterial composition along river networks [2,3,4,5]. Bacteria hold key roles in microbial communities and contribute significantly to biogeochemical processes and the cycling of nutrients in river ecosystems [6,7,8]. Recent studies have shown that the biogeography patterns of bacterial communities in complex freshwater networks can be explained by their origins in upstream freshwater [9] and terrestrial sources [10]. Planktonic bacterial taxa arise as the sum of multiple upstream sources of bacteria that grow in rainfall, lakes, groundwater, and soil. The resulting planktonic bacterial community is vulnerable to changes in its composition and structure. Sedimentary bacterial taxa develop from long-term cumulative processes of sediment erosion and deposition under ambient conditions [11]. The spatiotemporal distribution of planktonic and sedimentary bacterial communities in rivers could be quite different. Moreover, bacterial diversity is significantly altered by varying fluvial landforms and severe human interference along a large river. A better understanding of bacterial responses to the changing environment of river ecosystems is useful in the context of riverine cycles of nutrients, e.g., carbon and nitrogen, which are highly relevant to emission or sequestration of greenhouse gases [12,13,14].
Until recently, riverine bacterial communities have proved highly diverse and variable. Spatial and temporal variability of bacterioplankton composition in rivers has been demonstrated [2,3,4,5, 4: Figure S3) and co-occurrence of season-associated OTUs within the same phylum in network analysis (Additional file 5: Figure S4, Fig. 1b, d). In each phylum, positive association (red lines) dominates negative association (blue lines), indicating the uniform seasonality response of season-associated OTUs within a phylum. This uniform response is quantified by the network density value (d) (“Season-associated taxa analysis” section). Phyla that respond consistently are identified by having a higher density value implying a denser distribution of the association among OTUs. For example, Verrucomicrobia and Spirochaetes are the two most tightly correlated clusters (d = 0.433 and 0.455, respectively), whereas Acidobacteria in sediment samples and Actinobacteria and Bacteroidetes in water samples are more dispersed. The co-occurrence network analysis reveals almost exclusively or overwhelmingly positive correlations, consistent with the general pattern of dominant season-associated OTUs in most phyla (Fig. 1a, c).
In summary, the results show that microbial communities in Yangtze River are particularly sensitive to the season and are more prevalent in the autumn.
Biogeography patterns of bacterial communities
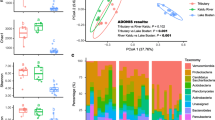
Non-metric multidimensional scaling (NMDS) was applied using unweighted UniFrac distance to identify the community compositions of all samples (Fig. 2). The first axis revealed that the bacterial communities of water samples were different from the corresponding sediment samples regardless of sampling sites and season. Bacterial communities of water samples demonstrated clear seasonal groups. However, bacterial communities of sediment samples did not form two separated clusters by season. The consistency of the results was confirmed by using the analysis of similarity (ANOSIM) statistic test of pairwise Bray-Curtis dissimilarities (Additional file 6: Figure S5). No significant difference was found between spring and autumn (global r = 0.122, P = 0.001) for sedimentary bacterial communities, whereas two seasonally distinct groups (global r = 0.525, P = 0.001) were observed for the water samples.
Overall, the spatial similarity of bacterial communities is better described by mean dendritic distance rather than cumulative dendritic distance or site catchment area (Additional file 7: Table S2). Closer correlation was demonstrated between the bacterial community similarity matrix and mean dendritic distance (Fig. 3b (mantel r = 0.3324, P = 0.001), d (mantel r = 0.3480, P = 0.001)) in terms of sediment samples than those based on water samples (Fig. 3a (mantel r = 0.2843, P = 0.001), c (mantel r = 0.2415, P = 0.001)). From the results, the mean dendritic distance appears more appropriate to describe bacterial community similarity in autumn sediment samples (Fig. 3d) and spring water samples (Fig. 3a). The distance decay analysis suggests that geographical distance could be of importance in structuring the bacterial assembly and determining the spatial similarity between different sites along the Yangtze River.
The Yangtze River flows through various landform types, including plateau, mountain, basin, foothill, and plain. Mainstream samples were used to study the effects of spatial variation, such as the river continuum and landform on bacterial communities. For water bacterial communities, NMDS (Additional file 8: Figure S6 (a) and (b)) gave similar results in both spring and autumn, with five separate groups: group 1 (between stations 1 and 2), group 2 (between stations 3 and 5), group 3 (from stations 6 to 9), group 4 (from stations 10 to 13), and group 5 (from stations 14 to 24), corresponding to the local landforms: mountain, foothill, basin, foothill-mountain, and plain. Moving window analysis (Additional file 9: Figure S7) was used to characterize the change rate of bacterial communities along the mainstream based on a comparison of results between two consecutive sampling sites. Higher change rates were always found at sampling sites where the landform type changed. Meanwhile, ANOSIM analysis (Additional file 10: Figure S8) further illustrated that taxonomic compositions of microbial communities significantly varied by landform type (P = 0.001). For the sediment samples, a similar clustering result (Additional file 8: Figure S6 (c) and (d)) was also obtained; this indicated that bacterial communities from the same landform tended to be similar to each other. The results of these analyses revealed that spatial variation in bacterial compositions across the samples could be partially attributed to the landform.
To further investigate the taxonomic distribution and differentially dominant clades of diverse landform ecosystems in water and sediment, we used the LEfSe biomarker discovery suite [31] to compare the abundance of bacterial compositions at each taxonomic level and determine taxa differentially abundant in at least one landform. Figure 4 depicts cladograms that visualize all detected bacterial compositions (relative abundance > 0.5%) from domain to genus level. Thirty and 70 differentially abundant taxa (i.e., colored circles in Fig. 4) were detected in water and sediment, respectively. These significantly enriched taxa provide a good indication of the primary characteristics of bacterial community structures in the Yangtze River, corresponding to the five landform types.
LEfSe cladogram of microbial community obtained for five landform types in water (a) and sediment (b). All detected taxa, with relative abundance ≥ 0.5% in at least one sample, assigned to domain (innermost), phylum, class, order, family, and genus (outermost), are used to determine the taxa or clades most likely to explain differences between landform types. Differentially abundant taxa (biomarkers) are colored according to their most abundant landform habitats; red, green, orange, purple, and blue circles stand for taxa that are abundant in plains, mountains, foothill-mountains, basins, and foothills, respectively. The color intensity of the outmost ring is proportional to the taxa abundance (genus level) at the landform type of greatest prevalence
Influential factors on bacterial community compositions
Even though canonical correspondence analysis (CCA) of bacterial communities in water and sediment indicated weak correlation to environmental factors (Additional file 11: Table S3, and Additional file 12: Figure S9), water temperature was found to be the primary factor in structuring bacterial community assemblages in both water and sediment of the Yangtze River. In addition, DO (dissolved oxygen) influenced the bacterial community in water, and pH, Mdd (mean dendritic distance), and TN (total nitrogen) influenced the bacterial community in sediment.
Impacts of the large dams
The Yangtze River contains a cascade of large dams, including two of the world’s largest dams, the Three Gorges Dam and the ** prokaryote community structure. Environ Microbiol. 2006;8:732–40." href="/article/10.1186/s40168-017-0388-x#ref-CR35" id="ref-link-section-d167081282e1150">35] neutral model was used to interpret the biogeographic distribution of bacterial communities in both water and sediment of the Yangtze River. The results showed that the neutral interpretation gave an excellent fit to the bacterial community distribution in the large riverine system considered (R2 > 0.7756), and an even higher correlation than previously achieved for smaller systems, such as coastal lakes in Antarctic (R2 ≤ 0.50) [36], and zebrafish (R2 ∈ [0.39, 0.81]) [37] (Fig. 6). Furthermore, the estimated immigration rate (m) originating from sediment communities (spring sediment 0.1640, autumn sediment 0.1463) was much lower than from water communities (spring water 0.1997, autumn water 0.1814), suggesting there were much more serious dispersal limits experienced by sediment communities. In general, the Sloan et al.’s [35] neutral model predicts the occurrence frequency to be > 85.44% in water communities, whereas only 66.08 and 61.96% taxa could be described in spring sediment and autumn sediment samples, respectively (Fig. 6, Additional file 14: Table S4).
Fit the occurrence frequency of different OTUs as a function of mean relative abundance using Sloan et al.’s [35] neutral model, for a spring-water, b spring-sediment, c autumn-water, and d autumn-sediment communities. Orange and green dots indicate the OTUs that occur more and less frequently than given by the model. Dashed lines represent 95% confidence intervals around the model prediction (red line)
The neutral model did not interpret 100% of the community species distribution, indicating that other community assembly mechanisms were perhaps operating at the same time. There is some evidence of species sorting in the present results. For example, the LEfSe analysis indicated that different abundant species or clades (biomarkers) occurred in five landform types (Fig. 4). Canonical correspondence analysis (CCA) of the bacterial communities in water and sediment showed a weak dependence on environmental factors. A significant discrepancy in the abundance of OTUs (Fig. 5) was observed upstream and downstream of the Three Gorges Dam, with most OTUs having higher abundance upstream of the dam except for several OTUs belonging to the genera Anaerolinea and Flavobacterium (Fig. 5). This implies that each sampling site possesses specific species due to its particular environment. The non-random distributions of bacteria are ascribed to the heterogeneous environment that affects their natural habitats and nutrients, influencing selection.
Discussion
A rapidly increasing number of studies based on high-throughput sequencing technologies have revealed a tremendous diversity of bacterial communities residing in the aquatic environment [38,39,40]. Most previous investigations on the variability and diversity of bacterial communities in rivers focused on a single dimension, i.e., either a long-term time series or a small-scale spatial dimension across environmental gradients [4, 19]. Here, we describe spatiotemporal patterns of lotic bacterial communities over a 4300 km river continuum for both water and sediment during spring and autumn seasons. Although changing slightly across the seasons, bacterial communities in sediment provide the main contribution to the bacterial diversity of the Yangtze River, and only 1.2% of the total OTUs are unique to water samples. The bacterial population fluctuation in water samples is higher than in sediment samples, as expected. Few previous studies have compared planktonic and sedimentary bacterial diversity in river reaches and coastal areas [35] neutral model was fitting to describe the relationship between the observed occurrence frequency of OTUs (the proportion of local communities in which each OTU is detected) and their abundance (the mean relative abundance across all local communities) [66]. The model is an adaptation of Hubbell’s neutral community model adjusted to bacterial populations analyzed with molecular tools [24]. This model emphasizes the effects of stochastic dispersal and drift (birth-death immigration process) but ignores the ecological difference between species and their response to the surrounding environment. In this model, the random loss of an individual is immediately replaced by immigration from the meta-community, with probability m, or reproduction within the local community, with probability 1-m [35]. The immigration rate was determined using non-linear least squares fitting in minpack.lm package of R-3.2. To further assess the deviations from the neutral model fitting, OTUs were subsequently sorted into three partitions depending on whether they occurred more frequently than “above” partition, less frequently than “below” partition, or within “neutral” partition the 95% confidence interval of the neutral model predictions.
Season-associated taxa analysis
OTUs with occurrence in more than 30% of all sediment or water samples were defined as persistent bacterial OTUs. An occupancy criterion was employed in order to generate the overall trend for taxonomic dendrograms (Fig. 1a, c) in water and sediment sample. Persistent bacterial OTUs that differed significantly between spring and autumn (P < 0.05) were further characterized as season-associated OTUs. According to their abundance in two seasons, season-associated OTUs were classified as autumn-associated OTUs (with significantly higher abundance in autumn) or spring-associated OTUs (with significantly higher abundance in spring). The network density value (d) was determined as the number of significant co-correlations divided by the number of all nodes, that is, a higher value represents a more intensive or dense response. Only persistent bacterial OTUs were displayed in taxonomic dendrograms (Fig. 1a, c) to avoid unstable associations and inconsistent trend caused by transient OTUs. To obtain comprehensive correlation among season-associated OTUs and non-associated OTUs, association networks were applied to both persistent and transient OTUs for each dominant phylum using edge-weighted spring-embedded layout algorithm (Fig. 1b, d).
References
Aufdenkampe AK, Mayorga E, Raymond PA, Melack JM, Doney SC, Alin SR, et al. Riverine coupling of biogeochemical cycles between land, oceans, and atmosphere. Front Ecol Environ. 2011;9:53–60.
Staley C, Unno T, Gould TJ, Jarvis B, Phillips J, Cotner JB, et al. Application of Illumina next-generation sequencing to characterize the bacterial community of the Upper Mississippi River. J Appl Microbiol. 2013;115:1147–58.
Savio D, Sinclair L, Ijaz UZ, Parajka J, Reischer GH, Stadler P, et al. Bacterial diversity along a 2600 km river continuum. Environ Microbiol. 2015;12:4994–5007.
Kolmakova OV, Gladyshev MI, Rozanov AS, Peltek SE, Trusova MY. Spatial biodiversity of bacteria along the largest Arctic river determined by next-generation sequencing. FEMS Microbiol Ecol. 2014;89:442–50.
Read DS, Gweon HS, Bowes MJ, Newbold LK, Field D, Bailey MJ, et al. Catchment-scale biogeography of riverine bacterioplankton. ISME J. 2015;9:516–26.
Battin TJ, Luyssaert S, Kaplan LA, Aufdenkampe AK, Richter A, Tranvik LJ. The boundless carbon cycle. Nat Geosci. 2009;2:598–600.
Findlay S. Stream microbial ecology. J North Am Benthol Soc. 2010;29:170–81.
Madsen EL. Microorganisms and their roles in fundamental biogeochemical cycles. Curr Opin Biotechnol. 2011;22:456–64.
Hauptmann AL, Markussen TN, Stibal M, Olsen NS, Elberling B, Bælum J, et al. Upstream freshwater and terrestrial sources are differentially reflected in the bacterial community structure along a small Arctic River and its estuary. Front Microbiol. 2016;7:1474.
Ruiz-González C, Niño-García JP, del Giorgio PA. Terrestrial origin of bacterial communities in complex boreal freshwater networks. Ecol Lett. 2015;18:1198–206.
Qian N, Zhang R, De Zhou Z. The fluvial process. Bei**g: Science Press (in Chinese); 1987.
Raymond PA, Hartmann J, Lauerwald R, Sobek S, McDonald C, Hoover M, et al. Global carbon dioxide emissions from inland waters. Nature. 2013;503:355–9.
Ward ND, Keil RG, Medeiros PM, Brito DC, Cunha AC, Dittmar T, et al. Degradation of terrestrially derived macromolecules in the Amazon River. Nat Geosci. 2013;6:530–3.
Borges AV, Darchambeau F, Teodoru CR, Marwick TR, Tamooh F, Geeraert N, et al. Globally significant greenhouse-gas emissions from African inland waters. Nat Geosci. 2015;8:637–42.
Feng BW, Li XR, Wang JH, Hu ZY, Meng H, **ang LY, et al. Bacterial diversity of water and sediment in the Changjiang estuary and coastal area of the East China Sea. FEMS Microbiol Ecol. 2009;70:80–92.
**a N, **a X, Liu T, Hu L, Zhu B, Zhang X, et al. Characteristics of bacterial community in the water and surface sediment of the Yellow River, China, the largest turbid river in the world. J Soils Sediments. 2014;14:1894–904.
Crump BC, Peterson BJ, Raymond PA, RMW A, Rinehart A, JW MC, et al. Circumpolar synchrony in big river bacterioplankton. Proc Natl Acad Sci U S A. 2009;106:21208–12.
Nelson CE. Phenology of high-elevation pelagic bacteria: the roles of meteorologic variability, catchment inputs and thermal stratification in structuring communities. ISME J. 2009;3:13–30.
Fortunato CS, Herfort L, Zuber P, Baptista AM, Crump BC. Spatial variability overwhelms seasonal patterns in bacterioplankton communities across a river to ocean gradient. ISME J. 2012;6:554–63.
Cavender-Bares J, Kozak KH, Fine PVA, Kembel SW. The merging of community ecology and phylogenetic biology. Ecol Lett. 2009;12:693–715.
Leibold MA, Holyoak M, Mouquet N, Amarasekare P, Chase JM, Hoopes MF, et al. The metacommunity concept: a framework for multi-scale community ecology. Ecol Lett. 2004;7:601–13.
Logue JB, Lindström ES. Biogeography of bacterioplankton in inland waters. Fr Rev. 2008;1:99–114.
Vanwonterghem I, Jensen PD, Dennis PG, Hugenholtz P, Rabaey K, Tyson GW. Deterministic processes guide long-term synchronised population dynamics in replicate anaerobic digesters. ISME J. 2014;8:2015–28.
Hubbell SP. The unified neutral theory of biodiversity and biogeography. Princeton: Princeton University Press; 2001.
Battin TJ, Besemer K, Bengtsson MM, Romani AM, Packmann AI. The ecology and biogeochemistry of stream biofilms. Nat Rev Microbiol. 2016;14:251–63.
Logue JB, Lindström ES. Species sorting affects bacterioplankton community composition as determined by 16S rDNA and 16S rRNA fingerprints. ISME J. 2010;4:729–38.
Dumbrell AJ, Nelson M, Helgason T, Dytham C, Fitter AH. Relative roles of niche and neutral processes in structuring a soil microbial community. ISME J. 2010;4:337–45.
Niño-García JP, Ruiz-González C, del Giorgio PA. Interactions between hydrology and water chemistry shape bacterioplankton biogeography across boreal freshwater networks. ISME J. 2016;10:1755–66.
Langenheder S, Székely AJ. Species sorting and neutral processes are both important during the initial assembly of bacterial communities. ISME J. 2011;5:1086–94.
Van der Gucht K, Cottenie K, Muylaert K, Vloemans N, Cousin S, Declerck S, et al. The power of species sorting: local factors drive bacterial community composition over a wide range of spatial scales. Proc Natl Acad Sci. 2007;104:20404–9.
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12:R60.
Liu H, Wang Z, Lu Y. Self-adjustment mechanism of bed structures under hydrology and sediment regimes. Hydrol Res. 2016;47:136–48.
Wang Z-Y, Li Y, He Y. Sediment budget of the Yangtze River. Water Resour Res. 2007;43:W04401.
Zhang W, Yuan J, Han J, Huang C, Li M. Impact of the Three Gorges Dam on sediment deposition and erosion in the middle Yangtze River: a case study of the Shashi Reach. Hydrol Res. 2016;47:175–86.
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP. Quantifying the roles of immigration and chance in sha** prokaryote community structure. Environ Microbiol. 2006;8:732–40.
Logares R, Lindström ES, Langenheder S, Logue JB, Paterson H, Laybourn-Parry J, et al. Biogeography of bacterial communities exposed to progressive long-term environmental change. ISME J. 2013;7:937–48.
Burns AR, Stephens WZ, Stagaman K, Wong S, Rawls JF, Guillemin K, et al. Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J. 2016;10:655–64.
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR, et al. Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc Natl Acad Sci U S A. 2006;103:12115–20.
Galand PE, Casamayora EO, Kirchmand DL, Lovejoye C. Ecology of the rare microbial biosphere of the Arctic Ocean. Proc Natl Acad Sci U S A. 2009;106:22427–32.
Andersson AF, Riemann L, Bertilsson S. Pyrosequencing reveals contrasting seasonal dynamics of taxa within Baltic Sea bacterioplankton communities. ISME J. 2010;4:171–81.
Biers EJ, Sun S, Howard EC. Prokaryotic genomes and diversity in surface ocean waters: interrogating the global ocean sampling metagenome. Appl Environ Microbiol. 2009;75:2221–9.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S, et al. The Sorcerer II Global Ocean Sampling expedition: Northwest Atlantic through eastern tropical Pacific. PLoS Biol. 2007;5:e77.
Zwart G, Crump BC, Agterveld MPK, Hagen F, Han SK. Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquat Microb Ecol. 2002;28:141–55.
Bahr M, Hobbie JE, Sogin ML. Bacterial diversity in an arctic lake: a freshwater SAR11 cluster. Aquat Microb Ecol. 1996;11:271–7.
Gibbons SM, Jones E, Bearquiver A, Blackwolf F, Roundstone W, Scott N, et al. Human and environmental impacts on river sediment microbial communities. PLoS One. 2014;9:e97435.
Green JL, Holmes AJ, Westoby M, Oliver I, David B, Dangerfield M, et al. Spatial scaling of microbial eukaryote diversity. Nature. 2004;432:747–50.
Liu L, Yang J, Yu Z, Wilkinson DM. The biogeography of abundant and rare bacterioplankton in lakes and reservoirs of China. ISME J. 2015;9:2068–77.
Bates ST, Clemente JC, Flores GE, Walters WA, Parfrey LW, Knight R, et al. Global biogeography of highly diverse protistan communities in soil. ISME J. 2013;7:652–9.
Östman Ö, Drakare S, Kritzberg ES, Langenheder S, Logue JB, Lindström ES. Regional invariance among microbial communities. Ecol Lett. 2010;13:118–27.
Liu J, Kuang W, Zhang Z, Xu X, Qin Y, Ning J, et al. Spatiotemporal characteristics, patterns and causes of land use changes in China since the late 1980s. J Geogr Sci. 2014;24:195–210.
Wang J, Fu B, Qiu Y, Chen L. Soil nutrients in relation to land use and landscape position in the semi-arid small catchment on the loess plateau in China. J Arid Environ. 2001;48:537–50.
Reith F, Brugger J, Zammit CM, Gregg AL, Goldfarb KC, Andersen GL, et al. Influence of geogenic factors on microbial communities in metallogenic Australian soils. ISME J. 2012;6:2107–18.
Logue JB, Langenheder S, Andersson AF, Bertilsson S, Drakare S, Lanzén A, et al. Freshwater bacterioplankton richness in oligotrophic lakes depends on nutrient availability rather than on species-area relationships. ISME J. 2012;6:1127–36.
Dai Z, Liu JT. Impacts of large dams on downstream fluvial sedimentation: an example of the Three Gorges Dam (TGD) on the Changjiang (Yangtze River). J Hydrol. 2013;480:10–8.
Janzen T, Haegeman B, Etienne RS. A sampling formula for ecological communities with multiple dispersal syndromes. J Theor Biol. 2015;374:94–106.
Crump BC, Adams HE, Hobbie JE, Kling GW. Biogeography of bacterioplankton in lakes and streams of an arctic tundra catchment. Ecology. 2007;88:1365–78.
Shan J, Ji R, Yu Y, **e Z, Yan X. Biochar, activated carbon, and carbon nanotubes have different effects on fate of 14C-catechol and microbial community in soil. Sci Rep. 2015;5:16000.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Edgar RC. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10:996–8.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Env Microbiol. 2007;73:5261–7.
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012;6:610–8.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Lozupone C, Knight R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol. 2005;71:8228–35.
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, et al. Vegan: community ecology package. R package version 2.4–0. 2016. https://CRAN.R-project.org/package=vegan. Accessed 15 Aug 2016.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Sloan WT, Woodcock S, Lunn M, Head IM, Curtis TP. Modeling taxa-abundance distributions in microbial communities using environmental sequence data. Microb Ecol. 2007;53:443–55.
Acknowledgments
We acknowledge the support from the Majorbio Company (Shanghai, China). We also thank Dr. Alistair Borthwick for his contribution to language editing.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No.91647211 and 51721006).
Availability of data and materials
The datasets supporting the conclusions of this article were deposited into the NCBI’s Sequence Read Archive (SRA) database (Accession Number: SRP079372).
Author information
Authors and Affiliations
Contributions
TL, ANZ, and JWW analyzed the data and prepared the draft of the manuscript. SL, XJ, CD, TM, STL, QC, and SX assisted in the data analysis and provided important suggestions for the manuscript preparation. JRN designed the research. TZ and JRN guided the data analysis and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1: Figure S1.
Map of the Yangtze River basin showing all the sampling sites in this study. Lines indicate the mainstream river and its tributaries, the former having a continuum of 4300 km (i.e., the actual sinuous channel length, equivalent to 2.05 times the straight line distance of 2102 km from start to the end sampling sites). Black dots indicate sampling points in the midstream; red dots represent sampling points in tributaries. (TIFF 5094 kb)
Additional file 2: Figure S2.
Rarefaction curves of bacterial richness of each sample (a) and sub-ecosystems (b) of Yangtze River. The end slope of the rarefaction curve was used to estimate the growth rate of the maximum value of the number of reads sampled. (TIFF 1709 kb)
Additional file 3: Table S1.
Summary of alpha diversity indices, including richness (Ace、Chao) and diversity (Good’s coverage, Shannon, Simpson) indices. (XLSX 34 kb)
Additional file 4: Figure S3.
Correlation distribution in the most populated phyla (coded with different colors) among both persistent and transient bacterial OTUs (> 0.1% relative abundance) with significant associations (P < 0.05) in water samples (a) and sediment samples (b). The Spearman value (> 0.5 or < − 0.5) was plotted to represent the degree of (positive or negative) correlation with higher absolute value as robust correlation. (TIFF 6045 kb)
Additional file 5: Figure S4.
Percentage abundances of prominent bacterial phyla in the rivers (a: water-spring; b: water-autumn; c: sediment-spring; d: sediment-autumn). (TIFF 2766 kb)
Additional file 6: Figure S5.
The ranks of the dissimilarities within and between groups for planktonic (a) and sedimentary (b) bacterial communities were estimated by ANOSIM (analysis of similarity statistics). The samples are grouped by season. (TIFF 1644 kb)
Additional file 7: Table S2.
Spearman relationships between the three kinds of river network distance and Bray-Curtis similarity of bacterial communities. (XLSX 10 kb)
Additional file 8: Figure S6.
Non-metric multidimensional scaling diagram showing the bacterial composition among five landform types in the water-spring (a), water-autumn (b), sediment-spring (c) and sediment-autumn (d) samples. (TIFF 1007 kb)
Additional file 9: Figure S7.
Moving window analysis based on MiSeq sequencing data for water samples. Each data point in the graph provides a comparison between two consecutive sites, as it represents the correlation between the samples of site x and site x-1. (TIFF 1301 kb)
Additional file 10: Figure S8.
The ranks of the dissimilarities within and between groups for planktonic (a) and sedimentary (b) bacterial communities estimated by ANOSIM (analysis of similarity statistics). The samples are grouped by seasons and landform types. (TIFF 6556 kb)
Additional file 11: Table S3.
Canonical correspondence analysis (CCA) indicating the effects of geographical and environmental factors on bacterial community composition. (XLSX 10 kb)
Additional file 12: Figure S9.
Canonical correspondence analysis showing the bacterial community composition of water (a) and sediment (b) in relation to monitored environmental factors. (TIFF 710 kb)
Additional file 13: Figure S10.
OTU richness of bacterial populations in water and sediment along the river. The number of sequence reads in each sample was normalized by randomly subsampling to the least of reads (24,197 sequences for each sample). The gray line between station 11 and station 12 indicates the location of the Three Gorges Dam. The sequencing depth was 24,197 sequences. (TIFF 601 kb)
Additional file 14: Table S4.
Percentages of the “above,” “below,” and “neutral” partitions of bacterial communities obtained by fitting Sloan et al.’s [39] neutral model. The fitting proportion of bacterial communities by the Sloan neutral model. (XLSX 9 kb)
Additional file 15: Table S5.
Dissimilarity values within and between groups for water-spring samples. The samples are grouped by seasons and landform types. (XLSX 9 kb)
Additional file 16: Table S6.
Dissimilarity values within and between groups for water-autumn samples. The samples are grouped by seasons and landform types. (XLSX 9 kb)
Additional file 17: Table S7.
Detailed information on the 50 sampling stations. (XLSX 12 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Liu, T., Zhang, A.N., Wang, J. et al. Integrated biogeography of planktonic and sedimentary bacterial communities in the Yangtze River. Microbiome 6, 16 (2018). https://doi.org/10.1186/s40168-017-0388-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40168-017-0388-x