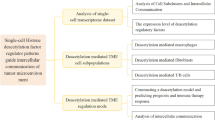
Abstract
Background
The diversity of histologic composition reflects the inter- and intra-tumor heterogeneity of lung adenocarcinomas (LUADs) macroscopically. Insights into the oncological characteristics and tumor microenvironment (TME) of different histologic subtypes of LUAD at the single-cell level can help identify potential therapeutic vulnerabilities and combinational approaches to improve the survival of LUAD patients.
Methods
Through comparative profiling of cell communities defined by scRNA-seq data, we characterized the TME of LUAD samples of distinct histologic subtypes, with relevant results further confirmed in multiple bulk transcriptomic, proteomic datasets and an independent immunohistochemical validation cohort.
Results
We find that the hypoxic and acidic situation is the worst in the TME of solid LUADs compared to other histologic subtypes. Besides, the tumor metabolic preferences vary across histologic subtypes and may correspondingly im**e on the metabolism and function of immune cells. Remarkably, tumor cells from solid LUADs upregulate energy and substance metabolic activities, particularly the folate-mediated one-carbon metabolism and the key gene MTHFD2, which could serve as a potential therapeutic target. Additionally, ubiquitination modifications may also be involved in the progression of histologic patterns. Immunologically, solid LUADs are characterized by a predominance of exhausted T cells and immunosuppressive myeloid cells, where the hypoxic, acidified and nutrient-deprived TME has a non-negligible impact. Discrepancies in stromal cell function, evidenced by varying degrees of stromal remodeling and fibrosis, may also contribute to the specific immune phenotype of solid LUADs.
Conclusions
Overall, our research proposes several potential entry points to improve the immunosuppressive TME of solid LUADs, thereby synergistically potentiating their immunotherapeutic efficacy, and may provide precise therapeutic strategies for LUAD patients of distinct histologic subtype constitution.
Similar content being viewed by others
Background
Invasive lung adenocarcinomas (LUADs) account for almost 70–90% of all surgically resected lung cancers [1]. The morphologic manifestations of invasive LUADs have been well characterized microscopically and are mainly differentiated into lepidic, papillary, acinar, micropapillary, and solid growth patterns [1, 2]. The diversity of histologic composition macroscopically reflects the inter-tumor and intra-tumor heterogeneity of LUADs, with most LUADs manifesting as a successive tissue transition between two or more histologic patterns [2]. Solid and acinar are two histologic subtypes of LUAD with high frequency, the solid type was identified as a histologic pattern stronger in aggressiveness, higher in grade, and worse in prognosis than the acinar type [2]. Identification of patients who may benefit from additional treatment after curative surgery for early LUAD has been a focus in the field of adjuvant therapy. An early study suggested that the solid type benefited from adjuvant chemotherapy in terms of disease-free survival (DFS) and specific DFS, while the acinar type did not [3]. Moreover, the latest grading system, introduced with 20% or more of high-grade patterns (including the solid pattern) as the cut-off for histologically high-grade LUADs, consistently demonstrated that those patients with high-grade LUADs could benefit from adjuvant chemotherapy [Immunohistochemistry Tissues were fixed in 4% paraformaldehyde, embedded in paraffin, cut into sections, and placed on adhesion microscope slides. Sections were subjected to immunohistochemical (IHC) staining according to standard procedures. We performed the IHC by using the MTHFD2 mouse anti-human antibody (Abcam, ab56772). The primary antibody was incubated at 4 °C overnight followed by using the BOND™ Polymer Refine Detection Kit (Leica, DS9800) according to the manufacturer’s instructions. Whole slide scanning was performed using panoramic MIDI under a 40 × objective lens. For each slide, the histologic patterns were firstly identified according to the cellular structure of the tumor, then three to five non-overlap** fields of view for each histologic region were randomly captured at 100 × magnification, and the staining intensity of MTHFD2 was finally semi-quantified using the Image J software (1.53q) by transforming it into mean optical density [32]. The statistical difference in staining intensity of MTHFD2 between solid and lepidic/acinar was determined by the Wilcoxon rank-sum test. The statistical analyses involved in this study were described in the corresponding method section. All statistical analyses and data presentations were performed by the R program (versions 3.6.3 and 4.0.2). All reported P values were two-tailed, and P < 0.05 was considered statistically significant.Statistics
Results
Analysis of scRNA-seq data from histologically annotated LUAD samples
The present study was a repurposing of scRNA-seq data from two of our previously published researches [12, 13]. All surgically excised samples came from patients with untreated, primary non-metastatic LUADs. The histologic constituents of each tumor sample were assessed and recorded semi-quantitatively [1, 2]. We attempted to single out tumor samples with high histologic purity for the purpose of dissecting subtype-specific oncological and immunological characteristics. Collectively, four solid-type, four acinar-type LUAD samples, and five adjacent normal lung samples were enrolled in this study. The representative hematoxylin–eosin (HE) stained images clearly visualized the microscopic structure of the acinar and solid patterns (Additional file 7: Fig. S1A–B). There was a "near-pure" tumor with the solid pattern covering more than 70% of the whole tumor in each solid LUAD sample [33]. With regards to the acinar type, the proportion of acinar pattern in each sample was greater than 50%, with the content of solid/micropapillary patterns limited to less than 10%, allowing to minimize the impacts of high-grade histologic components. The clinicopathological information for all enrolled samples was summarized in Additional file 1: Table S1.
The single-cell transcriptomic profiles generated by each sample were then combined for integrated analysis. Following strict quality control procedures, a sparse matrix with 97,875 cells and 25,233 genes was obtained (Methods). Before performing unsupervised graph-based clustering analysis, potential batch effects between samples were assessed and eliminated. Subsequently, all cells were labeled preliminary based on the expression of canonical cell markers (roughly, PTPRC for immune cells, EPCAM for epitheliums, VWF and COL1A2 for stromal cells; Additional file 7: Fig. S1C-D). Among these cells, 30,208 (30.86%) originated from solid samples, 25,250 (25.80%) originated from acinar samples, and 42,417 (43.34%) originated from adjacent lung tissues.
Tumor cells from solid LUADs create a more anoxic and acidic TME
We then committed to comparing the transcriptional characteristics of tumor cells derived from solid or acinar samples. By inferring large-scale copy number variations from transcriptome information, extensive chromosomal aberrations were observed in tumor-derived epitheliums relative to stromal cells (Additional file 7: Fig S1E). Comparing solid and acinar samples using gene set variation analyses (GSVA) [34] revealed that hallmarks associated with aggressiveness and metabolic activity, such as G2M checkpoint, angiogenesis, epithelial-mesenchymal transition (EMT), MYC targets V1 and PI3K/AKT/mTOR signaling, were up-regulated in tumor cells from solid samples (Fig. 1A, Additional file 2: Table S2), which was consistent with a more aggressive histopathological phenotype of solid LUADs. Notably, immune response-related hallmarks (such as TNFα signaling via NF-κB, IL2-STAT5 signaling, and IL6-JAK-STAT3 signaling) were also significantly enriched in solid samples. These findings emphasized the invasiveness of tumor cells from solid LUADs as well as their adept immune evasion capabilities.
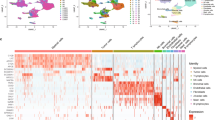
Tumor cells from solid LUADs create a more anoxic and acidic tumor microenvironment. A. Differentially enriched hallmarks (top) and KEGG pathways (bottom) between tumor cells from solid and acinar samples revealed by GSVA. B. Violin plots showing enrichment scores of tumor proliferating rate and hypoxia signatures by histologic subtypes in the TCGA LUAD cohort. Global differences were measured by the Kruskal–Wallis test. C. Violin plots of upregulated genes in solid LUAD tumor cells. D. Boxplots showing mRNA expression of HIF1A, LDHA, UBE2S and UBE2C by histologic subtypes in the TCGA LUAD cohort. Box centerlines, median; box limits, the 25th and 75th percentiles; box whiskers, 1.5 × the interquartile range. Comparisons were performed using two-sided Wilcoxon rank-sum test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, n s not significant). E–H. Violin plots showing enrichment scores of glycolysis (E), lactate transmembrane transporter activity (F), checkpoint molecules (G) and ubiquitin mediated proteolysis (H) signatures by histologic subtypes in the TCGA LUAD cohort. Global differences were measured by the Kruskal–Wallis test
It has been well established that hypoxia and acidification characterized the tumor microenvironment [35]. When comparing tumor cells from solid and acinar samples, hypoxia and glycolysis hallmarks were found to be more prominent in the former (Additional file 2: Table S2). Consistently, by applying single sample enrichment analysis (ssGSEA) in bulk RNA-seq data from the TCGA LUAD cohort, we found the enrichment scores of the tumor proliferating rate [36] and hypoxia [25] signatures were increased stepwise with histologic progression (Fig. 1B; Additional file 2: Table S2). Moreover, the expression levels of hypoxia-inducible factor-1 alpha (HIF1A) and lactate dehydrogenase A (LDHA), were observably upregulated in tumor cells from solid LUADs (Fig. 1C, Additional file 2: Table S2). As the key mediator of hypoxic response, HIF1A was intimately linked to multiple aspects of antitumor immunobiological processes [73]. A concomitant concern is whether the progression of histologic patterns is accompanied by the transformation in metabolic profiles of LUADs. Here we did find a gradient of metabolic alterations and relatively specific metabolic preferences between histologic subtypes, these metabolic properties coincided with the malignant potential of the histologic subtypes and might have a direct or indirect impact on intra-tumoral immune function. Energetically, tumor cells from solid LUADs upregulated glycolytic activity, confronting immune cells, which also relied on glycolysis for effector functions, with a scarcer energy source [10]. Notably, the reasons for the spatial distribution and functional differences of immune cells in the solid histologic region remain elusive. Here we propose the following two potential explanations. Firstly, the potential contribution of differences in the spatial distribution of oxygen and nutrients in the tumor regions of solid LUADs; and secondly, the obstruction by ECM components to the migration and movement of immune cells. In the case of the former, we introduce here the tumor model proposed by Lloyd et al. whereby tumor cores tend to maximize their population density and exhibit static, less proliferative phenotypes, while tumor margins are characterized by aggressive proliferative phenotypes [85]. Intriguingly, this model fits highly with the spatial characteristics of the solid pattern of LUAD identified by Tavernari et al. [10]. The harsh metabolic microenvironment created by vicious competition for limited resources in the tumor core may be detrimental to the survival and functional execution of immune cells. Indeed, Lambrechts et al. also suggested that the degree of hypoxia increases progressively from the tumor margin toward the core, whereas most immune cells are inclined to accumulate at the normoxic tumor margin [69]. In the case of the latter, CAFs and their remodeling of the ECM are key factors in structuring the immune infiltration barrier [86]. Based on comparative analysis of transcriptional profiles of the identified fibroblasts, we find that the fibrillar collagen transcriptional level is significantly higher in solid LUAD-derived fibroblasts [28]. And bulk transcriptome-based analysis further confirmed the elevated fibrillar collagen transcription and extracellular matrix remodeling activities in solid LUADs. In addition, it was noteworthy that solid LUADs are often accompanied by substantial intracellular and extracellular mucus production and secretion [1]. This implies that therapeutic regimens targeting CAFs or local ECM potentially promote immune infiltration into the tumor core of solid LUADs, thereby increasing the inter-contact between immune and tumor compartments.
Conclusions
Collectively, we herein proposed some potential entry points to disrupt the immune exclusion and immunosuppressive phenotype and to potentiate immunotherapeutic efficacy for solid LUADs, yet the realization of these notions requires further investigation and validation at different experimental techniques scales, such as microdissection and spatial omics techniques, as well as tumor models. Furthermore, considering the prospect of possible future applications of histologic subtype-directed LUAD treatment, the development of methods to determine the histologic composition or the presence of certain key components in the tumor prior to treatment is crucial.
Availability of data and materials
All data used for this study are publicly published and available with detailed access links described in the Data resources section. No new algorithms were developed for this manuscript. All code generated for analysis is available from the authors upon request.
Abbreviations
- LUAD:
-
Lung adenocarcinoma
- TME:
-
Tumor microenvironment
- TMB:
-
Tumor mutation burden
- ICB:
-
Immune checkpoint blocker
- EMT:
-
Epithelial-mesenchymal transition
- GSVA:
-
Gene set variation analysis
- ssGSEA:
-
Single sample gene set enrichment analysis
- HIF1A:
-
Hypoxia-inducible factor-1 alpha
- LDHA:
-
Lactate dehydrogenase A
- TCGA:
-
The cancer genome atlas
- EAS:
-
The east Asian ancestry
- CPTAC:
-
The clinical proteomic tumor analysis consortium
- MTHFD2:
-
Methylenetetrahydrofolate dehydrogenase 2
- ROS:
-
Reactive oxygen species
- pDCs:
-
Plasmacytoid dendritic cells
- DCs:
-
Dendritic cells
- cDCs:
-
Conventional dendritic cells
- CAFs:
-
Cancer-associated fibroblasts
- ECM:
-
Extracellular matrix
- MHC:
-
Major histocompatibility complex
- FFPE:
-
Formalin-fixed and paraffin-embedded
- IHC:
-
Immunohistochemistry
References
Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger KR, Yatabe Y, et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol O Publ Intern Assoc Study Lung Cancer. 2011;6(2):244–85.
Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM, Beasley MB, et al. The 2015 world health organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol Off Publ Intern Assoc Study of Lung Cancer. 2015;10(9):1243–60.
Tsao MS, Marguet S, Le Teuff G, Lantuejoul S, Shepherd FA, Seymour L, et al. Subtype classification of lung adenocarcinoma predicts benefit from adjuvant chemotherapy in patients undergoing complete resection. J Clin Oncol Off J Am Soc Clin Oncol. 2015;33(30):3439–46.
Moreira AL, Ocampo PSS, **a Y, Zhong H, Russell PA, Minami Y, et al. A grading system for invasive pulmonary adenocarcinoma: a proposal from the international association for the study of lung cancer pathology committee. J Thorac Oncol Off Publ Intern Assoc Study Lung Cancer. 2020;15(10):1599–610.
Hou L, Wang T, Chen D, She Y, Deng J, Yang M, et al. Prognostic and predictive value of the newly proposed grading system of invasive pulmonary adenocarcinoma in Chinese patients: a retrospective multicohort study. Mod Pathol. 2022;35(6):749–56.
Caso R, Sanchez-Vega F, Tan KS, Mastrogiacomo B, Zhou J, Jones GD, et al. The underlying tumor genomics of predominant histologic subtypes in lung adenocarcinoma. J Thorac Oncol Off Publ Intern Assoc Study Lung Cancer. 2020;15(12):1844–56.
Dong ZY, Zhang C, Li YF, Su J, **e Z, Liu SY, et al. Genetic and immune profiles of solid predominant lung adenocarcinoma reveal potential immunotherapeutic strategies. J Thorac oncol Off Publ Intern Assoc Study Lung Cancer. 2018;13(1):85–96.
Ci B, Yang DM, Cai L, Yang L, Girard L, Fujimoto J, et al. Molecular differences across invasive lung adenocarcinoma morphological subgroups. Transl lung cancer res. 2020;9(4):1029–40.
Li X, Wenes M, Romero P, Huang SC, Fendt SM, Ho PC. Navigating metabolic pathways to enhance antitumour immunity and immunotherapy. Nat Rev Clin Oncol. 2019;16(7):425–41.
Tavernari D, Battistello E, Dheilly E, Petruzzella AS, Mina M, Sordet-Dessimoz J, et al. Nongenetic evolution drives lung adenocarcinoma spatial heterogeneity and progression. Cancer Discov. 2021;11(6):1490–507.
Nguyen TT, Lee HS, Burt BM, Wu J, Zhang J, Amos CI, et al. A lepidic gene signature predicts patient prognosis and sensitivity to immunotherapy in lung adenocarcinoma. Genome Med. 2022;14(1):5.
Chen J, Tan Y, Sun F, Hou L, Zhang C, Ge T, et al. Single-cell transcriptome and antigen-immunoglobin analysis reveals the diversity of B cells in non-small cell lung cancer. Genome Biol. 2020;21(1):152.
He D, Wang D, Lu P, Yang N, Xue Z, Zhu X, et al. Single-cell RNA sequencing reveals heterogeneous tumor and immune cell populations in early-stage lung adenocarcinomas harboring EGFR mutations. Oncogene. 2021;40(2):355–68.
Cancer Genome Atlas Research N. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511(7511):543–50.
Chen J, Yang H, Teo ASM, Amer LB, Sherbaf FG, Tan CQ, et al. Genomic landscape of lung adenocarcinoma in east Asians. Nat Genet. 2020;52(2):177–86.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–4.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):pl1.
Gillette MA, Satpathy S, Cao S, Dhanasekaran SM, Vasaikar SV, Krug K, et al. Proteogenomic characterization reveals therapeutic vulnerabilities in lung adenocarcinoma. Cell. 2020;182(1):200-25 e35.
Kim N, Kim HK, Lee K, Hong Y, Cho JH, Choi JW, et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat Commun. 2020;11(1):2285.
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM 3rd, et al. Comprehensive integration of single-cell data. Cell. 2019;177(7):1888-902.e21.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184(13):3573-87.e29.
**ng X, Yang F, Huang Q, Guo H, Li J, Qiu M, et al. Decoding the multicellular ecosystem of lung adenocarcinoma manifested as pulmonary subsolid nodules by single-cell RNA sequencing. Sci Adv. 2021. https://doi.org/10.1126/sciadv.abd9738.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1(6):417–25.
Buffa FM, Harris AL, West CM, Miller CJ. Large meta-analysis of multiple cancers reveals a common, compact and highly prognostic hypoxia metagene. Br J Cancer. 2010;102(2):428–35.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–7.
Liu Y, He S, Wang XL, Peng W, Chen QY, Chi DM, et al. Tumour heterogeneity and intercellular networks of nasopharyngeal carcinoma at single cell resolution. Nat Commun. 2021;12(1):741.
Piersma B, Hayward MK, Weaver VM. Fibrosis and cancer: a strained relationship. Biochim Biophys Acta Rev Cancer. 2020;1873(2): 188356.
Tang Z, Kang B, Li C, Chen T, Zhang Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019;47(W1):W556–60.
Wu Y, Yang S, Ma J, Chen Z, Song G, Rao D, et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 2022;12(1):134–53.
DeTomaso D, Jones MG, Subramaniam M, Ashuach T, Ye CJ, Yosef N. Functional interpretation of single cell similarity maps. Nat Commun. 2019;10(1):4376.
Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9(7):671–5.
Yang SM, Chen LW, Wang HJ, Chen LR, Lor KL, Chen YC, et al. Extraction of radiomic values from lung adenocarcinoma with near-pure subtypes in the international association for the study of lung cancer/the american thoracic society/the european respiratory society (IASLC/ATS/ERS) classification. Lung cancer. 2018;119:56–63.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013;14:7.
Wang B, Zhao Q, Zhang Y, Liu Z, Zheng Z, Liu S, et al. Targeting hypoxia in the tumor microenvironment: a potential strategy to improve cancer immunotherapy. J Exp Clin Cancer Res. 2021;40(1):24.
Bagaev A, Kotlov N, Nomie K, Svekolkin V, Gafurov A, Isaeva O, et al. Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell. 2021;39(6):845-65.e7.
Lequeux A, Noman MZ, **ao M, Van Moer K, Hasmim M, Benoit A, et al. Targeting HIF-1 alpha transcriptional activity drives cytotoxic immune effector cells into melanoma and improves combination immunotherapy. Oncogene. 2021;40(28):4725–35.
Martinez-Reyes I, Chandel NS. Cancer metabolism: looking forward. Nat Rev Cancer. 2021;21(10):669–80.
Du X, Song H, Shen N, Hua R, Yang G. The molecular basis of ubiquitin-conjugating enzymes (E2s) as a potential target for cancer therapy. Int J Mol Sci. 2021;22(7):3440.
Pan YH, Yang M, Liu LP, Wu DC, Li MY, Su SG. UBE2S enhances the ubiquitination of p53 and exerts oncogenic activities in hepatocellular carcinoma. Biochem Biophys Res Commun. 2018;503(2):895–902.
Jung CR, Hwang KS, Yoo J, Cho WK, Kim JM, Kim WH, et al. E2-EPF UCP targets pVHL for degradation and associates with tumor growth and metastasis. Nat Med. 2006;12(7):809–16.
Liu W, **n H, Eckert DT, Brown JA, Gnarra JR. Hypoxia and cell cycle regulation of the von Hippel-Lindau tumor suppressor. Oncogene. 2011;30(1):21–31.
Sun T, Liu Z, Yang Q. The role of ubiquitination and deubiquitination in cancer metabolism. Mol Cancer. 2020;19(1):146.
Yang C, Zhang J, Liao M, Yang Y, Wang Y, Yuan Y, et al. Folate-mediated one-carbon metabolism: a targeting strategy in cancer therapy. Drug Discov Today. 2021;26(3):817–25.
Shang M, Yang H, Yang R, Chen T, Fu Y, Li Y, et al. The folate cycle enzyme MTHFD2 induces cancer immune evasion through PD-L1 up-regulation. Nat Commun. 2021;12(1):1940.
Yu C, Yang L, Cai M, Zhou F, **ao S, Li Y, et al. Down-regulation of MTHFD2 inhibits NSCLC progression by suppressing cycle-related genes. J Cell Mol Med. 2020;24(2):1568–77.
Harris IS, DeNicola GM. The complex interplay between antioxidants and ROS in cancer. Trends Cell Biol. 2020;30(6):440–51.
Sahoo BM, Banik BK, Borah P, Jain A. Reactive oxygen species (ROS): key components in cancer therapies. Anticancer Agents Med Chem. 2022;22(2):215–22.
Lee CQE, Kerouanton B, Chothani S, Zhang S, Chen Y, Mantri CK, et al. Coding and non-coding roles of MOCCI (C15ORF48) coordinate to regulate host inflammation and immunity. Nat Commun. 2021;12(1):2130.
Donnelly RP, Finlay DK. Glucose, glycolysis and lymphocyte responses. Mol Immunol. 2015;68(2 Pt C):513–9.
Caushi JX, Zhang J, Ji Z, Vaghasia A, Zhang B, Hsiue EH, et al. Transcriptional programs of neoantigen-specific TIL in anti-PD-1-treated lung cancers. Nature. 2021;596(7870):126–32.
Ren X, Zhang L, Zhang Y, Li Z, Siemers N, Zhang Z. Insights gained from single-cell analysis of immune cells in the tumor microenvironment. Annu Rev Immunol. 2021;39:583–609.
Guo X, Zhang Y, Zheng L, Zheng C, Song J, Zhang Q, et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat Med. 2018;24(7):978–85.
van der Leun AM, Thommen DS, Schumacher TN. CD8(+) T cell states in human cancer: insights from single-cell analysis. Nat Rev Cancer. 2020;20(4):218–32.
Geltink RIK, Kyle RL, Pearce EL. Unraveling the complex interplay between T cell metabolism and function. Annu Rev Immunol. 2018;36:461–88.
**a L, Oyang L, Lin J, Tan S, Han Y, Wu N, et al. The cancer metabolic reprogramming and immune response. Mol Cancer. 2021;20(1):28.
Borst J, Ahrends T, Babala N, Melief CJM, Kastenmuller W. CD4(+) T cell help in cancer immunology and immunotherapy. Nat Rev Immunol. 2018;18(10):635–47.
Mould KJ, Moore CM, McManus SA, McCubbrey AL, McClendon JD, Griesmer CL, et al. Airspace macrophages and monocytes exist in transcriptionally distinct subsets in healthy adults. Am J Respir Crit Care Med. 2021;203(8):946–56.
Cheng S, Li Z, Gao R, **ng B, Gao Y, Yang Y, et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell. 2021;184(3):792-809.e23.
Tie Y, Zheng H, He Z, Yang J, Shao B, Liu L, et al. Targeting folate receptor beta positive tumor-associated macrophages in lung cancer with a folate-modified liposomal complex. Signal Transduct Target Ther. 2020;5(1):6.
Chen J, Yao Y, Gong C, Yu F, Su S, Chen J, et al. CCL18 from tumor-associated macrophages promotes breast cancer metastasis via PITPNM3. Cancer Cell. 2011;19(4):541–55.
Cardoso AP, Pinto ML, Castro F, Costa AM, Marques-Magalhaes A, Canha-Borges A, et al. The immunosuppressive and pro-tumor functions of CCL18 at the tumor microenvironment. Cytokine Growth Factor Rev. 2021;60:107–19.
Jhunjhunwala S, Hammer C, Delamarre L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer. 2021;21(5):298–312.
Grauel AL, Nguyen B, Ruddy D, Laszewski T, Schwartz S, Chang J, et al. TGFbeta-blockade uncovers stromal plasticity in tumors by revealing the existence of a subset of interferon-licensed fibroblasts. Nat Commun. 2020;11(1):6315.
Chen Y, McAndrews KM, Kalluri R. Clinical and therapeutic relevance of cancer-associated fibroblasts. Nat Rev Clin Oncol. 2021;18(12):792–804.
Hussain A, Voisin V, Poon S, Karamboulas C, Bui NHB, Meens J, et al. Distinct fibroblast functional states drive clinical outcomes in ovarian cancer and are regulated by TCF21. J Exp Med. 2020. https://doi.org/10.1084/jem.20191094.
Biffi G, Tuveson DA. Diversity and biology of cancer-associated fibroblasts. Physiol Rev. 2021;101(1):147–76.
Carman CV, Martinelli R. T Lymphocyte-endothelial interactions: emerging understanding of trafficking and antigen-specific immunity. Front Immunol. 2015;6:603.
Lambrechts D, Wauters E, Boeckx B, Aibar S, Nittner D, Burton O, et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat Med. 2018;24(8):1277–89.
Liang P, Ballou B, Lv X, Si W, Bruchez MP, Huang W, et al. Monotherapy and combination therapy using anti-angiogenic nanoagents to fight cancer. Adv Mater. 2021;33(15): e2005155.
Huinen ZR, Huijbers EJM, van Beijnum JR, Nowak-Sliwinska P, Griffioen AW. Anti-angiogenic agents - overcoming tumour endothelial cell anergy and improving immunotherapy outcomes. Nat Rev Clin Oncol. 2021;18(8):527–40.
Li C, Guo L, Li S, Hua K. Single-cell transcriptomics reveals the landscape of intra-tumoral heterogeneity and transcriptional activities of ECs in CC. Mol Ther Nucleic Acids. 2021;24:682–94.
Faubert B, Solmonson A, DeBerardinis RJ. Metabolic reprogramming and cancer progression. Science. 2020;368(6487):eaaw5473.
Chang CH, Qiu J, O’Sullivan D, Buck MD, Noguchi T, Curtis JD, et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell. 2015;162(6):1229–41.
Vitale I, Manic G, Coussens LM, Kroemer G, Galluzzi L. Macrophages and metabolism in the tumor microenvironment. Cell Metab. 2019;30(1):36–50.
Leone RD, Powell JD. Metabolism of immune cells in cancer. Nat Rev Cancer. 2020;20(9):516–31.
Martinez-Usatorre A, Kadioglu E, Boivin G, Cianciaruso C, Guichard A, Torchia B, et al. Overcoming microenvironmental resistance to PD-1 blockade in genetically engineered lung cancer models. Sci Transl Med. 2021. https://doi.org/10.1126/scitranslmed.abd1616.
Zappasodi R, Serganova I, Cohen IJ, Maeda M, Shindo M, Senbabaoglu Y, et al. CTLA-4 blockade drives loss of T(reg) stability in glycolysis-low tumours. Nature. 2021;591(7851):652–8.
Canale FP, Basso C, Antonini G, Perotti M, Li N, Sokolovska A, et al. Metabolic modulation of tumours with engineered bacteria for immunotherapy. Nature. 2021;598(7882):662–6.
Kory N, Wyant GA, Prakash G, Uit de Bos J, Bottanelli F, Pacold ME, et al. SFXN1 is a mitochondrial serine transporter required for one-carbon metabolism. Science. 2018. https://doi.org/10.1126/science.aat9528.
Nilsson R, Nicolaidou V, Koufaris C. Mitochondrial MTHFD isozymes display distinct expression, regulation, and association with cancer. Gene. 2019;716: 144032.
Bonagas N, Gustafsson NMS, Henriksson M, Marttila P, Gustafsson R, Wiita E, et al. Pharmacological targeting of MTHFD2 suppresses acute myeloid leukemia by inducing thymidine depletion and replication stress. Nat Cancer. 2022;3(2):156–72.
Mo J, Gao Z, Zheng L, Yan M, Xue M, Xu J, et al. Targeting mitochondrial one-carbon enzyme MTHFD2 together with pemetrexed confers therapeutic advantages in lung adenocarcinoma. Cell Death Discov. 2022;8(1):307.
Scaletti ER, Gustafsson Westergren R, Andersson Y, Wiita E, Henriksson M, Homan EJ, et al. The first structure of human mthfd2l and its implications for the development of isoform-selective Inhibitors. ChemMedChem. 2022. https://doi.org/10.1002/cmdc.202200274.
Lloyd MC, Cunningham JJ, Bui MM, Gillies RJ, Brown JS, Gatenby RA. Darwinian dynamics of intratumoral heterogeneity: not solely random mutations but also variable environmental selection forces. Can Res. 2016;76(11):3136–44.
Chen X, Song E. Turning foes to friends: targeting cancer-associated fibroblasts. Nat Rev Drug Discov. 2019;18(2):99–115.
Acknowledgements
We thank all the participants in this study.
Funding
This research was funded by the National Science Foundation of China (Grant No.82125001, No.81972172), Clinical Research Plan of SHDC (Grant No. SHDC2020CR2020B), the Shanghai Science and Technology Committee (Grant No. 201409001000), the Shanghai Sailing Program (21YF1438300) and Funding of Shanghai Pulmonary Hospital (Grant No.FKCX1904, No.FKLY20004).
Author information
Authors and Affiliations
Contributions
PZ and DL conceived the study. JH and YY contributed to the data collection. DL and JH conducted the data analyses and interpreted the results. GJ, LZ, and HY provided administrative support and gave critical advice. LS provided help for statistic tests. LH and SL collected clinical samples and conducted the immunohistochemical analysis. DL and SL contributed to the design and writing of the manuscript. HY, JH, LZ, and PZ participated in the revision of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The study was conducted in accordance with the principles of the Declaration of Helsinki, and the study protocol was approved by the ethics committee of Shanghai Pulmonary Hospital. Because of the retrospective nature of the study, patient consent for inclusion was waived.
Consent for publication
Not applicable.
Competing interests
The authors declare no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1.
Characteristics of the LUAD samples included in this study.
Additional file 2: Table S2.
Data of epithelial cells analysis. (1) Differential expression genes between epithelial cells from solid and acinar samples; (2) Differentially enriched HALLMARK signatures in epithelial cells from solid and acinar samples; (3) Differentially enriched KEGG pathways in epithelial cells from solid and acinar samples; (4) Gene list of signatures for ssGSEA in bulk RNA-seq dataset of the TCGA LUAD cohort.
Additional file 3: Table S3.
Markers used for differing between primary immune cell types.
Additional file 4: Table S4.
Data of T/NK cells analysis. (1) Differentially expressed genes between CD8+ T cells from solid and acinar samples; (2) Differentially enriched KEGG pathways between CD8+ T cells from solid and acinar samples; (3) Gene list of signatures for T/NK cells.
Additional file 5: Table S5.
Data of myeloid cells analysis. (1) Gene list of signatures for macrophages; (2) Differentially expressed genes between macrophages from solid and acinar samples; (3) Gene list of signatures for DCs; (4) Differentially expressed genes between mregDCs from solid and acinar samples.
Additional file 6: Table S6.
Gene list of signatures for fibroblasts.
Additional file 7: Fig. S1.
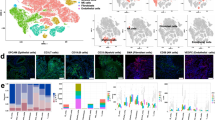
Primary classification of epitheliums, immune cells and stromal cells. A–B. Representative hematoxylin–eosin (HE) staining images of LUAD manifesting as acinar (A) and solid (B) growth pattern. The acinar pattern is predominantly glandular, round to oval in shape, with a central lumen surrounded by tumor cells. While the solid pattern consists of cytoplasm-rich polygonal tumor cells forming dense sheets and lacking any other recognizable patterns. The box regions in the upper panel are shown at higher magnification below. Scale bars, 200 μm (top panels) and 50 μm (lower panels). C. UMAP plots depicting all cells labeled as epitheliums, immune cells or stromal cells and split by sample types. D. UMAP plots of canonical markers for labeling general cell types. E. Heatmaps showing large-scale CNVs for individual epitheliums from tumor samples. Each row represented a cell and the columns represented chromosomal regions. Stromal cells were treated as references (top) and large-scale CNVs were observed in tumor cells (bottom).
Additional file 8: Fig. S2.
Characteristics of epithelial cells from solid and acinar samples. A–B. Boxplots showing HIF1A and LDHA mRNA expression across LUAD histologic subtypes in the EAS (A) and the CPTAC cohorts (B). Box centerlines, median; box limits, the 25th and 75th percentiles; box whiskers, 1.5× the interquartile range. For all comparisons of molecular expression between histologic subtypes, the statistical significance was determined by two-sided Wilcoxon rank-sum test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, n.s not significant). C–D. UBE2S and UBE2C mRNA expression in the the EAS cohort (C), and protein expression in the CPTAC cohort (D). Comparisons were performed using two-sided Wilcoxon rank-sum test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, n.s not significant).
Additional file 9: Fig. S3.
Metabolic differences between epithelial cells from solid and acinar samples. A. The average optical density of MTHFD2 immunohistochemical staining in tumor regions from different histologic patterns as semi-quantified by the Image J software. In the box plot, the centre line represents the median, box edges show the 25th and 75th percentiles, and whiskers extend to 1.5× the interquartile range. The statistical significance was determined by two-sided Wilcoxon rank-sum test. B. Kaplan–Meier survival curves showing the prognostic difference between the low and high MTHFD2 expression groups in the TCGA LUAD cohort. C. Correlation between the expression of MTHFD2 and UBE2S in the TCGA LUAD cohort. P-value was determined by Pearson's correlation test. D–F. Violin plots showing enrichment scores of one-carbon pool by folate (D), pyrimidine metabolism (E) and galactose metabolism (F) signatures by histologic subtypes in the TCGA LUAD cohort. Global differences were measured by the Kruskal-Wallis test.
Additional file 10: Fig. S4.
Primary immune cell types identification and T/NK cells analysis. A–C. UMAP plots of all immune cells colored by major immune types (A), sample origins (B) and histologic subgroups (C). D. UMAP plots of selected canonical markers for identifying major immune types. E–F. UMAP plots of T/NK cells colored by sample origins (E) and histologic subgroups (F). G. Dot plots showing the top marker genes for each T/NK cell cluster.H. Volcano plot showing differential expression genes between CD8+ T cells from solid and acinar samples. I. Differentially enriched KEGG pathways between CD4+ T cells from solid and acinar samples revealed by GSVA.
Additional file 11: Fig. S5.
Myeloid cells analysis. A. UMAP plot of annotated myeloid cells. B. UMAP plots of selected canonical markers for annotating myeloid cells. C. UMAP plot of monocytes and macrophages colored by histologic subgroups. D. Dot plots showing the top marker genes for each monocyte/macrophage cluster. E. Gene ontology annotation of marker genes for the Macro-C2 subset. F. Gene ontology annotation of marker genes for the Macro-C3 subset. G–I. Violin plots showing enrichment scores of macrophage and DC traffic (G), M1 phenotype (H) and immune suppression by myeloid cells (I) signatures by histologic subtypes in the TCGA LUAD cohort. Global differences were measured by the Kruskal-Wallis test.
Additional file 12: Fig. S6.
Fibroblasts analysis. A. UMAP plot of fibroblasts colored by histologic subgroups. B. UMAP plots showing the expression of selected canonical marker genes for fibroblasts. C. Dot plots showing the top marker genes for each fibroblast cluster. D. Gene ontology annotation of marker genes for the Fibro-C1 subset. E–G. Violin plots showing enrichment scores of fibrillar collagens (E), matrix remodeling (F) and EMT (G) signatures by LUAD histologic subtypes in the TCGA cohort. Global differences were measured by the Kruskal-Wallis test.
Additional file 13: Fig. S7.
Endothelial cells analysis. A. UMAP plot of endothelial cells colored by histologic subgroups. B. UMAP plots showing the expression of selected canonical marker genes for endothelial cells. C. Dot plots showing the top marker genes for each endothelium cluster.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, D., Yu, H., Hu, J. et al. Comparative profiling of single-cell transcriptome reveals heterogeneity of tumor microenvironment between solid and acinar lung adenocarcinoma. J Transl Med 20, 423 (2022). https://doi.org/10.1186/s12967-022-03620-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12967-022-03620-3