Abstract
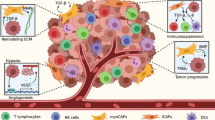
Metastasis is the leading cause of death in colorectal cancer (CRC) patients, and the liver is the most common site of metastasis. Tumor cell metastasis can be thought of as an invasion-metastasis cascade and metastatic organotropism is thought to be a process that relies on the intrinsic properties of tumor cells and their interactions with molecules and cells in the microenvironment. Many studies have provided new insights into the molecular mechanism and contributing factors involved in CRC liver metastasis for a better understanding of the organ-specific metastasis process. The purpose of this review is to summarize the theories that explain CRC liver metastasis at multiple molecular dimensions (including genetic and non-genetic factors), as well as the main factors that cause CRC liver metastasis. Many findings suggest that metastasis may occur earlier than expected and with specific organ-anchoring property. The emergence of potential metastatic clones, the timing of dissemination, and the distinct routes of metastasis have been explained by genomic studies. The main force of CRC liver metastasis is also thought to be epigenetic alterations and dynamic phenotypic traits. Furthermore, we review key extrinsic factors that influence CRC cell metastasis and liver tropisms, such as pre-niches, tumor stromal cells, adhesion molecules, and immune/inflammatory responses in the tumor microenvironment. In addition, biomarkers associated with early diagnosis, prognosis, and recurrence of liver metastasis from CRC are summarized to enlighten potential clinical practice, including some markers that can be used as therapeutic targets to provide new perspectives for the treatment strategies of CRC liver metastasis.
Similar content being viewed by others
Background
Colorectal cancer (CRC) is the world’s second leading cause of cancer deaths, and liver metastasis from CRC accounts for the majority of fatalities in CRC patients [1, 2]. CRC’s most common target metastatic sites are the liver, lung, bone, and brain, known as organ tropism, and the liver is the most common site of CRC metastasis [3]. Up to 50% of CRC patients have liver metastasis, and approximately 15–23% of patients have metastasis at the time of diagnosis. Hepatic resection combined with modern adjuvant systemic regimens is only effective in 20% of colorectal cancer liver metastasis (CRLM) patients. Even after curative hepatic resection, the 5-year overall survival rate is around 48% [4]. In practice, however, approximately 80% of CRLM patients have unresectable metastatic lesions [5, 6].They are typically downstaged by systemic and local therapy (including stereotactic radiotherapy, transarterial radioembolization, and transarterial chemoembolization) to achieve metastatic liver lesion resection, improving patients’ long-term survival and prognosis. Although the development of adjuvant systemic therapy has significantly improved the clinical outcome of patients with stage IV CRC liver metastases [7], early detection of CRLM remains critical to good results. Despite the high resolution of computed tomography (CT) as the most commonly used detection modality, up to 30% of liver metastasis cannot be detected in their early stages. MRI and PET-CT may have higher sensitivity and specificity, but the costs are prohibitive [2].
Many studies have found that cancer metastasis is a complex selective process influenced by anatomical, biological, and microenvironmental factors [8]. Theories such as “seed-soil,” pre-niche, and the crosstalk between tumor cells and immune cells provide direct evidence for metastatic propensity and organ-specific tropism of metastatic cancer cells, implying that metastasis is a process that is dependent on organ-targeted anchoring characteristics [9,10,11]. Thus, investigating how metastatic CRC appears, when it appears, and the underlying mechanism provides clues for the treatment of CRC and its liver metastasis.
Theories underlying organ-specific metastasis process
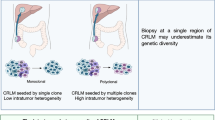
According to a traditional model of metastatic spread, cancer cells undergo the following general steps known as metastatic cascade, which can be divided into two major phases: (1) dissemination from the primary lesion to distant organs by entering systemic circulation and adapting to a new microenvironment (intravasation, extravasation), and (2) colonization followed by expanding growth [12]. The multi-step process typically involves an invasion-metastasis cascade. According to Hanahan and Weinberg, the hallmark of “activating invasion and metastasis” is one of the six important features of cancer. Invading cancer cells pass through or collaborate with stroma to avoid elimination by immune system cells such as neutrophils, monocytes/macrophages, and endothelial cells. Epithelial-mesenchymal transition (EMT) program can be activated by carcinoma cells and orchestrate most steps of invasion and metastasis, except for colonization. Disseminated tumor cells act in dormancy in circulation and new environment tissue to avoid immune surveillance, and then they interact with the tissue microenvironment to be awakened from dormancy. Moreover, the development of metastatic colonization needs multiple biological programs, and these adaptations require intrinsic capabilities of cancer cells and a permissive tumor microenvironment with stromal support cells. The invasion-metastasis process, which appears to be a linear progression from the primary tumor to metastatic colonies, operates under the guidance of a specific paradigm [13,14,15]. However, several studies have suggested that metastasis cannot be defined solely by chronological order, as many interdigitated and mutually exclusive metastasis events do not appear to follow a linear progression model [16]. In our understanding of CRC metastasis to the liver, the general “seed and soil” theory and the “mechanistic theory” are highly complementary [17]. Anatomically, CRC metastasis is thought to occur in a stepwise fashion, with the majority of venous drainage from the intestine entering the portal system to flow into the liver, and then the disseminated cells in the bloodstream are arrested by the first available liver capillary beds with endothelial cells and basement membrane [18]. Although the physical characteristics influence organ tropism and affect the non-random organ-specificity, the fact is that organs receiving similar blood volumes have distinct metastatic-formation efficacy. The special ability of circulating tumor cells to form secondary growth cannot be explained as purely mechanistic [19, 20]. Cancer cells entering the circulation disperse in various directions, but their anchorage to specific metastatic sites is determined by various factors [21]. Stephen Paget proposed a hypothesis in 1889 that described cancer cells as “seeds” and receptive microenvironments as “soils,” both of which are required for rate-limiting steps in the formation of micrometastasis [9]. Recently, growing evidence for the process of pre-metastatic niche formation adds new insights to the “seed-soil” theory. For example, metastasis-initiating cells co-opt the metastatic microenvironment to facilitate colonization. Before cancer cells disseminate from the primary tumor, a subpopulation of primary tumor cells has the “prime” potential and reprograms the distant microenvironments [22]. Pre-metastatic niche formation is important in CRLM. By recruiting various cellular components (Kupffer cells, macrophages, and fibroblasts), producing CRC-derived factors (chemokines and cytokines) and exosomes, primary CRC prepares a favorable microenvironment in the liver. It mediates liver-target metastasis [23, 24]. As a result, the concept of pre-metastasis niches may trump the chronological metastasis pattern that is dependent on circulation. Recently, many discoveries have clarified the key molecules and cells involved in liver-specific metastasis of CRC. For example, LINC00485 is a newly discovered class of Long non-coding RNAs (lncRNAs), and low expression of LINC00485 predicts a poor prognosis for CRC patients. LINC00485 attenuated CRC cell invasion and liver metastasis by directly modulating the miR-581/EDEM1 axis. Overexpression of LINC00485 enhanced the expression of epithelium markers E-cadherin and significantly down-regulated the expression of mesenchymal markers N-cadherin, indicating a loss of malignant phenotype in cancer cells [25]. Higher levels of tumor suppressor microRNAs (miR-25-3p, miR-130b-3p, miR-425-5p, miR-934) in the exosomes were secreted by CRC cells in more advanced disease, and these exosomal miRNAs induced macrophages M2 polarization to promote liver metastasis of CRC. CXCL13 secreted by M2-polarized macrophages promoted the transcription of exosomal miR-934 in CRC cells, forming a positive feedback loop to foster CRLM [35]. The relative timing of metastatic spread is determined by comparing the genetic divergence between the primary tumors and metastasis. In the classic “linear evolution model,” the metastasis-initiating clone(s) emerge late in the primary tumor and seed at the metastatic sites as a byproduct of tumor development. Instead, in the “parallel evolution model,” the metastatic subclone(s) spread from the primary tumor to distant sites early, and both the primary and metastatic subclones evolve concurrently. As a result, compared with the “linear evolution model,” a greater degree of Primary-Metastasis genetic divergence is expected in the “parallel evolution model” [36]. In addition to the time of metastasis, studies of the clonal relationship between primary tumors and metastases explained metastatic seeding patterns based on genomic data analysis: identification of monoclonal/polyclonal metastasis and monophyletic/polyphyletic metastasis may provide information for treatment improvement. In metastatic CRC, both monoclonal/polyclonal metastasis and monophyletic/polyphyletic seeding patterns were observed. Polyclonal metastasis appeared to be the most common type of CRC metastasis. A polyphyletic seeding pattern was observed in the case of CRC with liver metastasis followed by lung metastasis [37, 38]. For the distinct routes of metastasis, Hai-ning et al. investigated the genomic evolution for the clonal origin and revealed three metastatic models (sequential, branch-off, and diaspora) by phylogenetic reconstruction using Treeomics. The results of the genomic analysis showed that liver and lung metastasis might originate from primary tumors independently rather than subsequently, providing genomic evidence for the organotropisms of metastatic CRC cells. However, the relationship between the characteristics of primary site subclones and their potential for liver metastasis has not been thoroughly investigated [38].
The underlying molecular mechanisms and contributing factors involved in CRLM
Genetic and epigenetic changes associated with liver metastasis of CRC
Cancer cells are thought to acquire metastatic capacity due to genetic and epigenetic changes. Genetic mutations can potentially disrupt epigenetic patterns, and the interaction between these two mechanisms can promote metastasis. [33, 39,40,41,42].
Although numerous genetic alterations have been detected between the primary tumor and metastatic sites in CRC [43,44,45], much remains unknown about the interaction between tumor genomic features and metastatic potential and organ-specific metastatic patterns [34, 46]. The clonal relationship observed between paired primary tumors, and metastasis explains at least part of the dissemination of metastasis-competent clones in different temporal patterns and trajectories [36, 47, 48]. This section will will sort out the genetic/epigenetic alterations associated with CRC liver-specific metastasis and summarize recent studies on the metastatic evolution patterns (temporal patterns and routes) observed in the metastatic CRC cohort.
Genetic alterations associated with liver metastasis of CRC
Several whole-genome sequencing analyses on metastatic tumors have been performed in recent years to gain insight into the critical genetic events involved in CRLM. Several studies have been conducted to investigate single nucleotide variations (SNVs), mutated genes, and chromosome copy numbers of CRLM. An analysis of metastatic solid tumor genomes revealed that consistent genetic changes indicate cancer metastasis remains to be further identified [44]. A pan-cancer cohort study of 25,000 patients’ tumor genomic profiling identified the associations between genetic alterations and metastatic patterns in 50 tumor types. The result showed that copy number alterations were not significantly associated with the metastatic burden for CRC. Chromosomal instability may be established early in tumor development and was already high in patients with low metastatic burden [45]. Oga et al. discovered 6855 mutations in primary CRC tumors without liver metastasis, primary metastatic CRC, and paired liver metastasis (LM) lesions using whole-exome sequencing (WES) analysis. The result showed that the somatic genomic profiles of primary CRC tumors and LM lesions were not significantly different; however, LM regions showed an enriched A-to-C nucleotide conversion in the context of “AAG,” an event that may be specific to liver metastases [49]. Li et al. used WES to look for somatic SNVs (sSNVs) in primary tumors and matched liver metastasis samples from 16 CRC patients with liver metastasis. The SNVs data were analyzed using ABSOLUTE software to calculate the proportion of mutational genes in each sample. An average of 34% (8–63%) mutations were shared by both primary tumors and liver metastasis, indicating a common ancestral trunk among them. Furthermore, an average of 34% (12%–88%) mutations were metastasis-private, which may be a result obtained or lost during the tumor metastasis process. The probable timing order of mutation events has been investigated by analyzing the distribution of cancer cell fractions (CCF). A higher median CCF value indicates that the mutation occurred earlier. Data on the median CCF value of TP53 and KRAS showed that TP53 mutations occurred earlier than KRAS in primary tumors but later than KRAS in liver metastasis [50].
Several studies have been conducted to identify frequently mutated genes involved in metastasis. For example, 707 genes have been identified as LM-associated genes, which specifically mutated in the LM regions but not in CRC tumors without liver metastasis, including ADAMTS10, NELL1, and RXFP3, implying their roles in liver metastasis. Furthermore, ADAP1 fusions were discovered in the RNA-seq dataset, indicating that ADAP1 was fused to GET4, SUN1, or NOC4L in an out-of-frame manner in the LM region. Two in-frame fusions of the ADAP1’s ArfGAP domain with proteins from GEMIN4 and TMEM8A have been discovered, which may facilitate metastasis by activating GTPase [49]. A study used targeted sequencing of primary tumors and matched liver metastasis samples to describe the genome landscape of Chinese CRLM patients. The most frequently mutated genes were found to be TP53 (324/396, 82%), PC (302/396, 76%), KRAS (166/396, 42%), SMAD4 (54/396, 14%), FLG (52/396, 13%), and FBXW7 (43/396, 11%). Furthermore, the distribution of genomic changes was related to the time of metastasis (synchronous/metachronous liver metastasis). Alterations in genes of FBXW7, FLT3, XIRP2, TSC2, LATS1, and CREBBP were significantly enriched in metachronous lesions, and alterations in CDK12 were significantly enriched in synchronous LM [51].
The differences in chromosome copy number between primary and secondary tumors revealed that genetic aberrations in liver metastasis are a dynamic process, such as the presence of a focal amplification of chromosome 7p in primary tumors but not in the LM region. The loss or gain of copy number variations (CNVs) most likely allows clones to be more fit in a new environment [49]. Anand and colleagues investigated the link between aneuploidy and CRC metastasis. Aneuploidy is not just a byproduct of chromosomal instability; it has a direct influence on cancer cells’ metastatic capability, either promoting or inhibiting metastasis behavior. HCT116 colon cells with an extra copy of chromosome 5 exhibit increased invasive behavior by activating an EMT program and upregulated matrix metalloproteinases (MMPs) [52]. In addition, CNV alterations, as a common biological event during tumor progression and therapy, usually involve multiple genes. There are potentially complex interactions between co-amplified or co-deleted genes affected by CNV events, acting as a whole. It has been reported that CDK12 and HER2 were frequently co-amplified in CRC, and inhibition of CDK12 can enhance the sensitivity of CRC cells to lapatinib, an anti-HER2 tyrosine kinase inhibitor (TKI) [53].
Genome events related to metastatic evolution pattern
According to phylogenetic analysis of non-synonymous SNVs from the primary tumor and metastatic liver lesions, there were three main clonal evolution patterns from primary to liver metastases: clonal-clonal pattern (C–C) (early events), subclonal-clonal pattern (S–C) (middle-stage events), none-clonal pattern (0-C) (later events). In terms of CNV events, Chr 20q amp, 17p del, 18q del, and 8p del in clonal- clonal evolution were considered as early events, 8q amp in liver metastasis-specific evolution was considered as later events, and 8q amp, 13q amp, and 8p del in subclonal-clonal evolution were considered as middle-stage events. SYNE1 was a mutant gene with S–C clonal evolution characteristics. Its mRNA expression level in normal, CRC primary, and liver metastasis gradually decreased; however, its functional mechanism in CRLM remains unknown [50]. Tumor mutation burden, an immunotherapy biomarker, in conjunction with HLALOH (HLA, Human leukocytes antigens, LOH, Loss of heterozygosity), is used as an indicator to assess the efficacy of immunotherapy [54]. Subclonal mutation loads were higher in primary tumors than in clonal mutation loads. In contrast, the proportion of clonal mutation was increased in metastatic lesions, which is consistent with the S–C evolutionary pattern, indicating the role of selection in metastasis. HLA LOH occurred in samples with recurrent mutations of S–C changing pattern, including KRAS, SYNE1, FBXL2, DNAH11, and CACNA1H, indicating that this mutational clonal pattern promotes CRC cells evading the immune system during liver metastasis [50].
Epigenetic modifications associated with liver metastasis of CRC
No genetic changes have been identified as consensus metastasis-specific drivers in the process of CRC metastasis. However, epigenetic changes may provide an alternative mechanism to induce tumor cells for metastatic phenotypes [55]. The core content of epigenetic modification is the covalent modification of histones and nucleic acids (including methylation, acetylation, ubiquitination, etc.). In addition, epigenetic regulation also includes chromatin remodeling and transcriptional mediators (mainly non-coding RNAs, such as microRNAs and long ncRNAs) of the RNA splicing machinery. They affect gene expression without sequence changes in DNA [56, 57] (Fig. 2). Epigenetic changes play an important role in CRLM (Table 1), but whether there are metastasis-specific epigenetic drivers and their mechanism need to be investigated further [40].
The role of epigenetic modifications in CRC liver metastasis. Epigenetic modification plays a vital role in gene regulation, mainly for various covalent modifications of histones and nucleic acids. The change of nucleic acid is in DNA and RNA. In addition, epigenetic modification also includes chromatin remodeling, non-coding RNA regulation, and other mechanisms. DNA methylation mainly occurs at the C of 5′-CpG-3′ to generate 5-methylcytosine (5mC). Under the action of DNA methyltransferase (DNMT), methyl groups are covalently bonded to the 5' carbon of cytosines of CpG dinucleotide residues. Hypermethylated gene expression is suppressed. Chromatin remodeling can regulate gene expression by regulating chromatin changes in chromatin structure and location, such as PU.1 opening chromatin regions of downstream effector genes and recruiting additional epigenetic modifiers to regulate gene expression. N6-Adenylate methylation (m6A), which inserts a methyl substituent on the N atom at the 6-position of adenosine. During transcription, m6A deposited on RNA transcripts affects gene expression post-transcriptionally by altering the structure of RNA or the specific recognition of m6-binding proteins. Non-coding RNAs are endogenous RNA molecules that cannot be translated into proteins but have particular gene expression regulatory functions, regulating post-transcriptional gene expression by complementary binding to RNA transcripts of the target gene
Dysregulation in DNA methylation is the mainly studied DNA modification in tumor and metastasis [64]. The methylation changes of primary CRC, metastatic CRC, and liver metastases differ between individuals. CRC primary tumors exhibited global hypomethylation and CpG island (CGI) hypermethylation compared to healthy tissues, whereas metastatic colorectal lesions exhibit high-level global methylation but lower CGI methylation [65]. The study by Udali et al. came to the same conclusion. Primary CRC and synchronous liver metastases had similar epigenetic DNA hypomethylation status when compared with homologous cancer-free colon tissues, indicating that these epigenetic mechanisms occurred in the early stages of CRC development and were maintained till the stage of liver metastasis progression [66]. However, the mechanism by which methylation inhibits tumor suppressor gene expression may be compromised during metastasis. Mahdi et al. discovered that the regulatory mechanism of methylation on gene expression might be compromised during the process of CRC tumor cell metastasis and colonization in the liver. The expression levels of three endothelin system genes changed significantly during the liver colonization of CC531 cells. When metastatic cell lines were exposed to Decitabine (DAC, which inhibits DNA methyltransferases), the expression of endothelin system genes did not increase, indicating that these gene expression changes were not caused by DNA methylation. This suggests that the regulatory function of epigenetic alterations may be gradually lost in the late stage of metastasis [67]. The microenvironment-induced epigenetic mutation is an essential mechanism for metastatic tumor cells to grow in their new niche. The hepatic growth factor (HGF) is abundant in the microenvironment of liver metastases. HGF from the metastatic liver microenvironment was shown to activate the c-Met/PI3K/AKT/mTOR axis in CRC cells, activating the SREBP2-dependent cholesterol biosynthesis pathway to promote CRC liver metastasis [68]. PU.1 is a pioneer factor that remodels chromosomes by opening the enclosed chromatin and enlisting the help of additional epigenetic modifiers. According to one study, HGF caused PU.1 phosphorylation in metastatic cells. The phosphorylated PU.1 regulated downstream regulatory elements to activate the effector gene DPP4. The HGF/PU.1/DPP4 axis was activated, which promoted the growth of CRC tumor cells at the site of the liver. Targeting the chromatin remodeling pathway in the future may provide additional treatment options for metastatic cancer [ All data included in this study are available upon request by contact with the corresponding author. Colorectal cancer Colorectal cancer liver metastasis Single nucleotide variations Somatic single nucleotide variations Liver metastasis Cancer cell fractions Epithelial-mesenchymal transition Copy number variations Matrix metalloproteinases Tyrosine kinase inhibitor Clonal- Clonal Subclonal- Clonal None- Clonal Human leukocytes antigens Loss of heterozygosity CpG island Hepatic growth factor N6-methyladenosine Circular RNAs Long non-coding RNAs Cytoskeleton regulator RNA Cancer stem cell Migrating CSCs Transient amplification Tumor- initiating cells LT-TICs: long term tumor- initiating cells G-protein-coupled receptor 5 Intestinal stem cell Myeloid- derived suppressor cells Sphingosine 1-phosphate receptor 1 Signal transducer and activator of transcription 3 Cancer-related fibroblasts Bone morphogenetic protein Nucleotide-binding oligomerization domain 1 LPS-stimulated Monocyte Conditioned Medium Inflammatory monocytes Dendritic cell Array-based comparative genomic hybridization 3'-Untranslated region Long non-coding RNAs Circulating tumor cell Circular RNAs Pre-miRNAs Fms-related tyrosine kinase 3 ligand Riihimaki M, Hemminki A, Sundquist J, Hemminki K. Patterns of metastasis in colon and rectal cancer. Sci Rep. 2016;6:29765. Sawatzki M, Guller U, Gusewell S, Husarik DB, Semela D, Brand S. Contrast-enhanced ultrasound can guide the therapeutic strategy by improving the detection of colorectal liver metastases. J Hepatol. 2021;74(2):419–27. Tamas K, Walenkamp AM, de Vries EG, van Vugt MA, Beets-Tan RG, van Etten B, de Groot DJ, Hospers GA. Rectal and colon cancer: not just a different anatomic site. Cancer Treat Rev. 2015;41(8):671–9. Zarour LR, Anand S, Billingsley KG, Bisson WH, Cercek A, Clarke MF, Coussens LM, Gast CE, Geltzeiler CB, Hansen L, et al. Colorectal cancer liver metastasis: evolving paradigms and future directions. Cell Mol Gastroenterol Hepatol. 2017;3(2):163–73. Baba K. Successful treatment of conversion chemotherapy for initially unresectable synchronous colorectal liver metastasis. World J Gastroenterol. 2015. https://doi.org/10.3748/wjg.v21.i6.1982. Cherradi S, Ayrolles-Torro A, Vezzo-Vie N, Gueguinou N, Denis V, Combes E, Boissiere F, Busson M, Canterel-Thouennon L, Mollevi C, et al. Antibody targeting of claudin-1 as a potential colorectal cancer therapy. J Exp Clin Cancer Res. 2017;36(1):89. Adam R, Kitano Y. Multidisciplinary approach of liver metastases from colorectal cancer. Ann Gastroenterol Surg. 2019;3(1):50–6. Urosevic J, Garcia-Albeniz X, Planet E, Real S, Cespedes MV, Guiu M, Fernandez E, Bellmunt A, Gawrzak S, Pavlovic M, et al. Colon cancer cells colonize the lung from established liver metastases through p38 MAPK signalling and PTHLH. Nat Cell Biol. 2014;16(7):685–94. Langley RR, Fidler IJ. The seed and soil hypothesis revisited–the role of tumor-stroma interactions in metastasis to different organs. Int J Cancer. 2011;128(11):2527–35. Lin Q, Ren L, Jian M, Xu P, Li J, Zheng P, Feng Q, Yang L, Ji M, Wei Y, et al. The mechanism of the premetastatic niche facilitating colorectal cancer liver metastasis generated from myeloid-derived suppressor cells induced by the S1PR1-STAT3 signaling pathway. Cell Death Dis. 2019;10(10):693. Wang M, Su Z, Amoah Barnie P. Crosstalk among colon cancer-derived exosomes, fibroblast-derived exosomes, and macrophage phenotypes in colon cancer metastasis. Int Immunopharmacol. 2020;81: 106298. Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331(6024):1559–64. Suhail Y, Cain MP, Vanaja K, Kurywchak PA, Levchenko A, Kalluri R. Kshitiz: systems biology of cancer metastasis. Cell Syst. 2019;9(2):109–27. Ganesh K, Massague J. Targeting metastatic cancer. Nat Med. 2021;27(1):34–44. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74. Bhattacharya R, Panda CK, Nandi S, Mukhopadhyay A. An insight into metastasis: random or evolving paradigms? Pathol Res Pract. 2018;214(8):1064–73. Zheng W, Wu F, Fu K, Sun G, Sun G, Li X, Jiang W, Cao H, Wang H, Tang W. Emerging mechanisms and treatment progress on liver metastasis of colorectal cancer. Onco Targets Ther. 2021;14:3013–36. Weidle UH, Birzele F, Kruger A. Molecular targets and pathways involved in liver metastasis of colorectal cancer. Clin Exp Metastasis. 2015;32(6):623–35. Obenauf AC, Massague J. Surviving at a Distance: Organ-Specific Metastasis. Trends Cancer. 2015;1(1):76–91. Budczies J, von Winterfeld M, Klauschen F, Bockmayr M, Lennerz JK, Denkert C, Wolf T, Warth A, Dietel M, Anagnostopoulos I, et al. The landscape of metastatic progression patterns across major human cancers. Oncotarget. 2015;6(1):570–83. Rubin MA. Insights into the mechanism of organ-specific cancer metastasis. Cancer Discov. 2014;4(11):1262–4. Klein CA. Selection and adaptation during metastatic cancer progression. Nature. 2013;501(7467):365–72. Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M, Molina H, Kohsaka S, Di Giannatale A, Ceder S, et al. Tumour exosome integrins determine organotropic metastasis. Nature. 2015;527(7578):329–35. Yang L, Li T, Shi H, Zhou Z, Huang Z, Lei X. The cellular and molecular components involved in pre-metastatic niche formation in colorectal cancer liver metastasis. Expert Rev Gastroenterol Hepatol. 2021;15(4):389–99. Li C, Pan B, Liu X, Qin J, Wang X, He B, Pan Y, Sun H, Xu T, Xu X, et al. Long intergenic non-coding RNA LINC00485 exerts tumor-suppressive activity by regulating miR-581/EDEM1 axis in colorectal cancer. Aging. 2021;13(3):3866–85. Zhou H, Liu Z, Wang Y, Wen X, Amador EH, Yuan L, Ran X, **ong L, Ran Y, Chen W, et al. Colorectal liver metastasis: molecular mechanism and interventional therapy. Signal Transduct Target Ther. 2022;7(1):70. Zhao S, Mi Y, Guan B, Zheng B, Wei P, Gu Y, Zhang Z, Cai S, Xu Y, Li X, et al. Tumor-derived exosomal miR-934 induces macrophage M2 polarization to promote liver metastasis of colorectal cancer. J Hematol Oncol. 2020;13(1):156. Wang D, Sun H, Wei J, Cen B, DuBois RN. CXCL1 Is critical for premetastatic niche formation and metastasis in colorectal cancer. Cancer Res. 2017;77(13):3655–65. Jiang K, Chen H, Fang Y, Chen L, Zhong C, Bu T, Dai S, Pan X, Fu D, Qian Y, et al. Exosomal ANGPTL1 attenuates colorectal cancer liver metastasis by regulating Kupffer cell secretion pattern and impeding MMP9 induced vascular leakiness. J Exp Clin Cancer Res. 2021;40(1):21. Sun H, Meng Q, Shi C, Yang H, Li X, Wu S, Familiari G, Relucenti M, Aschner M, Wang X, et al. Hypoxia-inducible exosomes facilitate liver-tropic premetastatic niche in colorectal cancer. Hepatology. 2021;74(5):2633–51. Shao S, Zhu Y, Meng T, Liu Y, Hong Y, Yuan M, Yuan H, Hu F. Targeting high expressed alpha5beta1 integrin in liver metastatic lesions to resist metastasis of colorectal cancer by rpm peptide-modified chitosan-stearic micelles. Mol Pharm. 2018;15(4):1653–63. Ganesh K, Basnet H, Kaygusuz Y, Laughney AM, He L, Sharma R, O’Rourke KP, Reuter VP, Huang YH, Turkekul M, et al. L1CAM defines the regenerative origin of metastasis-initiating cells in colorectal cancer. Nat Cancer. 2020;1(1):28–45. Patel SA, Rodrigues P, Wesolowski L, Vanharanta S. Genomic control of metastasis. Br J Cancer. 2021;124(1):3–12. Kim TM, Jung SH, An CH, Lee SH, Baek IP, Kim MS, Park SW, Rhee JK, Lee SH, Chung YJ. Subclonal genomic architectures of primary and metastatic colorectal cancer based on intratumoral genetic heterogeneity. Clin Cancer Res. 2015;21(19):4461–72. Saito T, Niida A, Uchi R, Hirata H, Komatsu H, Sakimura S, Hayashi S, Nambara S, Kuroda Y, Ito S, et al. A temporal shift of the evolutionary principle sha** intratumor heterogeneity in colorectal cancer. Nat Commun. 2018;9(1):2884. Turajlic S, Swanton C. Metastasis as an evolutionary process. Science. 2016;352(6282):169–75. Leung ML, Davis A, Gao R, Casasent A, Wang Y, Sei E, Vilar E, Maru D, Kopetz S, Navin NE. Single-cell DNA sequencing reveals a late-dissemination model in metastatic colorectal cancer. Genome Res. 2017;27(8):1287–99. Chen HN, Shu Y, Liao F, Liao X, Zhang H, Qin Y, Wang Z, Luo M, Liu Q, Xue Z, et al. Genomic evolution and diverse models of systemic metastases in colorectal cancer. Gut. 2022;71(2):322–32. Chiang AC, Massague J. Molecular basis of metastasis. N Engl J Med. 2008;359(26):2814–23. Grady WM, Carethers JM. Genomic and epigenetic instability in colorectal cancer pathogenesis. Gastroenterology. 2008;135(4):1079–99. Testa U, Castelli G, Pelosi E. Genetic alterations of metastatic colorectal cancer. Biomedicines. 2020. https://doi.org/10.3390/biomedicines8100414. Alderton GK. Tumour evolution: epigenetic and genetic heterogeneity in metastasis. Nat Rev Cancer. 2017;17(3):141. Vignot S, Lefebvre C, Frampton GM, Meurice G, Yelensky R, Palmer G, Capron F, Lazar V, Hannoun L, Miller VA, et al. Comparative analysis of primary tumour and matched metastases in colorectal cancer patients: evaluation of concordance between genomic and transcriptional profiles. Eur J Cancer. 2015;51(7):791–9. Priestley P, Baber J, Lolkema MP, Steeghs N, de Bruijn E, Shale C, Duyvesteyn K, Haidari S, van Hoeck A, Onstenk W, et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature. 2019;575(7781):210–6. Nguyen B, Fong C, Luthra A, Smith SA, DiNatale RG, Nandakumar S, Walch H, Chatila WK, Madupuri R, Kundra R, et al. Genomic characterization of metastatic patterns from prospective clinical sequencing of 25,000 patients. Cell. 2022;185(3):563–75. Joung JG, Oh BY, Hong HK, Al-Khalidi H, Al-Alem F, Lee HO, Bae JS, Kim J, Cha HU, Alotaibi M, et al. Tumor heterogeneity predicts metastatic potential in colorectal cancer. Clin Cancer Res. 2017;23(23):7209–16. Sottoriva A, Barnes CP, Graham TA. Catch my drift? Making sense of genomic intra-tumour heterogeneity. Biochim Biophys Acta Rev Cancer. 2017;1867(2):95–100. Loeb LA, Kohrn BF, Loubet-Senear KJ, Dunn YJ, Ahn EH, O’Sullivan JN, Salk JJ, Bronner MP, Beckman RA. Extensive subclonal mutational diversity in human colorectal cancer and its significance. Proc Natl Acad Sci USA. 2019;116(52):26863–72. Oga T, Yamashita Y, Soda M, Kojima S, Ueno T, Kawazu M, Suzuki N, Nagano H, Hazama S, Izumiya M, et al. Genomic profiles of colorectal carcinoma with liver metastases and newly identified fusion genes. Cancer Sci. 2019;110(9):2973–81. Li C, Xu J, Wang X, Zhang C, Yu Z, Liu J, Tai Z, Luo Z, Yi X, Zhong Z. Whole exome and transcriptome sequencing reveal clonal evolution and exhibit immune-related features in metastatic colorectal tumors. Cell Death Discov. 2021;7(1):222. Wang HW, Yan XL, Wang LJ, Zhang MH, Yang CH, Wei L, ** KM, Bao Q, Li J, Wang K, et al. Characterization of genomic alterations in Chinese colorectal cancer patients with liver metastases. J Transl Med. 2021;19(1):313. Vasudevan A, Baruah PS, Smith JC, Wang Z, Sayles NM, Andrews P, Kendall J, Leu J, Chunduri NK, Levy D, et al. Single-Chromosomal gains can function as metastasis suppressors and promoters in colon cancer. Dev Cell. 2020;52(4):413–28. Li H, Wang J, Yi Z, Li C, Wang H, Zhang J, Wang T, Nan P, Lin F, Xu D, et al. CDK12 inhibition enhances sensitivity of HER2+ breast cancers to HER2-tyrosine kinase inhibitor via suppressing PI3K/AKT. Eur J Cancer. 2021;145:92–108. Pancione M, Giordano G, Remo A, Febbraro A, Sabatino L, Manfrin E, Ceccarelli M, Colantuoni V. Immune escape mechanisms in colorectal cancer pathogenesis and liver metastasis. J Immunol Res. 2014;2014: 686879. Banerjee R, Smith J, Eccles MR, Weeks RJ, Chatterjee A. Epigenetic basis and targeting of cancer metastasis. Trends Cancer. 2022;8(3):226–41. Hogg SJ, Beavis PA, Dawson MA, Johnstone RW. Targeting the epigenetic regulation of antitumour immunity. Nat Rev Drug Discov. 2020;19(11):776–800. Jung G, Hernandez-Illan E, Moreira L, Balaguer F, Goel A. Epigenetics of colorectal cancer: biomarker and therapeutic potential. Nat Rev Gastroenterol Hepatol. 2020;17(2):111–30. Wang L, Wang E, Prado Balcazar J, Wu Z, **ang K, Wang Y, Huang Q, Negrete M, Chen KY, Li W, et al. Chromatin remodeling of colorectal cancer liver metastasis is mediated by an HGF-PU 1-DPP4 Axis. Adv Sci. 2021;8(19):e2004673. Wang S, Gao S, Zeng Y, Zhu L, Mo Y, Wong CC, Bao Y, Su P, Zhai J, Wang L, et al. N6-Methyladenosine reader YTHDF1 Promotes ARHGEF2 translation and RhoA signaling in colorectal cancer. Gastroenterology. 2022;162(4):1183–96. Chen RX, Chen X, **a LP, Zhang JX, Pan ZZ, Ma XD, Han K, Chen JW, Judde JG, Deas O, et al. N(6)-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10(1):4695. Bleau AM, Redrado M, Nistal-Villan E, Villalba M, Exposito F, Redin E, de Aberasturi AL, Larzabal L, Freire J, Gomez-Roman J, et al. miR-146a targets c-met and abolishes colorectal cancer liver metastasis. Cancer Lett. 2018;414:257–67. Zheng X, Chen L, Zhou Y, Wang Q, Zheng Z, Xu B, Wu C, Zhou Q, Hu W, Wu C, et al. A novel protein encoded by a circular RNA circPPP1R12A promotes tumor pathogenesis and metastasis of colon cancer via Hippo-YAP signaling. Mol Cancer. 2019;18(1):47. Yue B, Liu C, Sun H, Liu M, Song C, Cui R, Qiu S, Zhong M. A positive feed-forward loop between LncRNA-CYTOR and Wnt/beta-catenin signaling promotes metastasis of colon cancer. Mol Ther. 2018;26(5):1287–98. Su R, Wu X, Tao L, Wang C. The role of epigenetic modifications in colorectal cancer metastasis. Clin Exp Metastasis. 2022;39(4):521–39. Ili C, Buchegger K, Demond H, Castillo-Fernandez J, Kelsey G, Zanella L, Abanto M, Riquelme I, Lopez J, Viscarra T, et al. Landscape of genome-wide DNA methylation of colorectal cancer metastasis. Cancers. 2020. https://doi.org/10.3390/cancers12092710. Udali S, De Santis D, Ruzzenente A, Moruzzi S, Mazzi F, Beschin G, Tammen SA, Campagnaro T, Pattini P, Olivieri O, et al. DNA methylation and hydroxymethylation in primary colon cancer and synchronous hepatic metastasis. Front Genet. 2017;8:229. Mahdi MR, Georges RB, Ali DM, Bedeer RF, Eltahry HM, Gabr AHZ, Berger MR. Modulation of the endothelin system in colorectal cancer liver metastasis: influence of epigenetic mechanisms? Front Pharmacol. 2020;11:180. Zhang KL, Zhu WW, Wang SH, Gao C, Pan JJ, Du ZG, Lu L, Jia HL, Dong QZ, Chen JH, et al. Organ-specific cholesterol metabolic aberration fuels liver metastasis of colorectal cancer. Theranostics. 2021;11(13):6560–72. Chen Y, Lin Y, Shu Y, He J, Gao W. Interaction between N(6)-methyladenosine (m(6)A) modification and noncoding RNAs in cancer. Mol Cancer. 2020;19(1):94. Lee J, Hong HK, Peng SB, Kim TW, Lee WY, Yun SH, Kim HC, Liu J, Ebert PJ, Aggarwal A, et al. Identifying metastasis-initiating miRNA-target regulations of colorectal cancer from expressional changes in primary tumors. Sci Rep. 2020;10(1):14919. Mouillet JF, Donker RB, Mishima T, Cronqvist T, Chu T, Sadovsky Y. The unique expression and function of miR-424 in human placental trophoblasts. Biol Reprod. 2013;89(2):25. Zhu ED, Li N, Li BS, Li W, Zhang WJ, Mao XH, Guo G, Zou QM, **ao B. miR-30b, down-regulated in gastric cancer, promotes apoptosis and suppresses tumor growth by targeting plasminogen activator inhibitor-1. PLoS ONE. 2014;9(8): e106049. Chen T, You Y, Jiang H, Wang ZZ. Epithelial-mesenchymal transition (EMT): A biological process in the development, stem cell differentiation, and tumorigenesis. J Cell Physiol. 2017;232(12):3261–72. Smith AG, Macleod KF. Autophagy, cancer stem cells and drug resistance. J Pathol. 2019;247(5):708–18. Miyazaki T, Chung S, Sakai H, Ohata H, Obata Y, Shiokawa D, Mizoguchi Y, Kubo T, Ichikawa H, Taniguchi H, et al. Stemness and immune evasion conferred by the TDO2-AHR pathway are associated with liver metastasis of colon cancer. Cancer Sci. 2022;113(1):170–81. Dieter SM, Ball CR, Hoffmann CM, Nowrouzi A, Herbst F, Zavidij O, Abel U, Arens A, Weichert W, Brand K, et al. Distinct types of tumor-initiating cells form human colon cancer tumors and metastases. Cell Stem Cell. 2011;9(4):357–65. Malanchi I, Santamaria-Martinez A, Susanto E, Peng H, Lehr HA, Delaloye JF, Huelsken J. Interactions between cancer stem cells and their niche govern metastatic colonization. Nature. 2011;481(7379):85–9. Walter RJ, Sonnentag SJ, Orian-Rousseau V, Munoz-Sagredo L. Plasticity in colorectal cancer: why cancer cells differentiate. Cancers. 2021. https://doi.org/10.3390/cancers13040918. Batlle E, Clevers H. Cancer stem cells revisited. Nat Med. 2017;23(10):1124–34. Walcher L, Kistenmacher AK, Suo H, Kitte R, Dluczek S, Strauss A, Blaudszun AR, Yevsa T, Fricke S, Kossatz-Boehlert U. Cancer Stem Cells-Origins and Biomarkers: Perspectives for Targeted Personalized Therapies. Front Immunol. 2020;11:1280. de SousaSousa e F, Kurtova AV, Harnoss JM, Kljavin N, Hoeck JD, Hung J, Anderson JE, Storm EE, Modrusan Z, Koeppen H, et al. A distinct role for Lgr5(+) stem cells in primary and metastatic colon cancer. Nature. 2017;543(7647):676–80. Todaro M, Gaggianesi M, Catalano V, Benfante A, Iovino F, Biffoni M, Apuzzo T, Sperduti I, Volpe S, Cocorullo G, et al. CD44v6 is a marker of constitutive and reprogrammed cancer stem cells driving colon cancer metastasis. Cell Stem Cell. 2014;14(3):342–56. Gao W, Chen L, Ma Z, Du Z, Zhao Z, Hu Z, Li Q. Isolation and phenotypic characterization of colorectal cancer stem cells with organ-specific metastatic potential. Gastroenterology. 2013;145(3):636–46. Merlos-Suarez A, Barriga FM, Jung P, Iglesias M, Cespedes MV, Rossell D, Sevillano M, Hernando-Momblona X, da Silva-Diz V, Munoz P, et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell. 2011;8(5):511–24. Qiu L, Yang X, Wu J, Huang C, Miao Y, Fu Z. HIST2H2BF potentiates the propagation of cancer stem cells via notch signaling to promote malignancy and liver metastasis in colorectal carcinoma. Front Oncol. 2021;11: 677646. Wang J, **ng Y, Wang Y, He Y, Wang L, Peng S, Yang L, **e J, Li X, Qiu W, et al. A novel BMI-1 inhibitor QW24 for the treatment of stem-like colorectal cancer. J Exp Clin Cancer Res. 2019;38(1):422. Oura K, Morishita A, Tani J, Masaki T. Tumor immune microenvironment and immunosuppressive therapy in hepatocellular carcinoma: a review. Int J Mol Sci. 2021. https://doi.org/10.3390/ijms22115801. Milette S, Sicklick JK, Lowy AM, Brodt P. Molecular pathways: targeting the microenvironment of liver metastases. Clin Cancer Res. 2017;23(21):6390–9. Pretzsch E, Bosch F, Neumann J, Ganschow P, Bazhin A, Guba M, Werner J, Angele M. Mechanisms of metastasis in colorectal cancer and metastatic organotropism: hematogenous versus peritoneal spread. J Oncol. 2019;2019:7407190. Zhu X, Wang F, Wu X, Li Z, Wang Z, Ren X, Zhou Y, Song F, Liang Y, Zeng Z, et al. FBX8 promotes metastatic dormancy of colorectal cancer in liver. Cell Death Dis. 2020;11(8):622. Brodt P. Role of the microenvironment in liver metastasis: from pre- to prometastatic niches. Clin Cancer Res. 2016;22(24):5971–82. Wen XQ, Qian XL, Sun HK, Zheng LL, Zhu WQ, Li TY, Hu JP. MicroRNAs: multifaceted regulators of colorectal cancer metastasis and clinical applications. Onco Targets Ther. 2020;13:10851–66. Vidal-Vanaclocha F, Crende O, de DurangoGarcia C, Herreros-Pomares A, Lopez-Domenech S, Gonzalez A, Ruiz-Casares E, Vilboux T, Caruso R, Duran H, et al. Liver prometastatic reaction: stimulating factors and responsive cancer phenotypes. Semin Cancer Biol. 2021;71:122–33. Zhang Y, Zhao Y, Li Q, Wang Y. Macrophages, as a promising strategy to targeted treatment for colorectal cancer metastasis in tumor immune microenvironment. Front Immunol. 2021;12: 685978. Liu C, Pei H, Tan F. Matrix stiffness and colorectal cancer. Onco Targets Ther. 2020;13:2747–55. Paauwe M, Schoonderwoerd MJA, Helderman R, Harryvan TJ, Groenewoud A, van Pelt GW, Bor R, Hemmer DM, Versteeg HH, Snaar-Jagalska BE, et al. Endoglin expression on cancer-associated fibroblasts regulates invasion and stimulates colorectal cancer metastasis. Clin Cancer Res. 2018;24(24):6331–44. Ma R, Feng Y, Lin S, Chen J, Lin H, Liang X, Zheng H, Cai X. Mechanisms involved in breast cancer liver metastasis. J Transl Med. 2015;13:64. Hu JL, Wang W, Lan XL, Zeng ZC, Liang YS, Yan YR, Song FY, Wang FF, Zhu XH, Liao WJ, et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol Cancer. 2019;18(1):91. Tan HX, Gong WZ, Zhou K, **ao ZG, Hou FT, Huang T, Zhang L, Dong HY, Zhang WL, Liu Y, et al. CXCR4/TGF-beta1 mediated hepatic stellate cells differentiation into carcinoma-associated fibroblasts and promoted liver metastasis of colon cancer. Cancer Biol Ther. 2020;21(3):258–68. Garner H, de Visser KE. Immune crosstalk in cancer progression and metastatic spread: a complex conversation. Nat Rev Immunol. 2020;20(8):483–97. Jiang HY, Najmeh S, Martel G, MacFadden-Murphy E, Farias R, Savage P, Leone A, Roussel L, Cools-Lartigue J, Gowing S, et al. Activation of the pattern recognition receptor NOD1 augments colon cancer metastasis. Protein Cell. 2020;11(3):187–201. **ng W, **ao Y, Lu X, Zhu H, He X, Huang W, Lopez ES, Wong J, Ju H, Tian L, et al. GFI1 downregulation promotes inflammation-linked metastasis of colorectal cancer. Cell Death Differ. 2017;24(5):929–43. Li M, Lai X, Zhao Y, Zhang Y, Li M, Li D, Kong J, Zhang Y, **g P, Li H, et al. Loss of NDRG2 in liver microenvironment inhibits cancer liver metastasis by regulating tumor associate macrophages polarization. Cell Death Dis. 2018;9(2):248. Wang D, Wang X, Si M, Yang J, Sun S, Wu H, Cui S, Qu X, Yu X. Exosome-encapsulated miRNAs contribute to CXCL12/CXCR4-induced liver metastasis of colorectal cancer by enhancing M2 polarization of macrophages. Cancer Lett. 2020;474:36–52. Grossman JG, Nywening TM, Belt BA, Panni RZ, Krasnick BA, DeNardo DG, Hawkins WG, Goedegebuure SP, Linehan DC, Fields RC. Recruitment of CCR2(+) tumor associated macrophage to sites of liver metastasis confers a poor prognosis in human colorectal cancer. Oncoimmunology. 2018;7(9): e1470729. Liu Y, Zhang Q, **ng B, Luo N, Gao R, Yu K, Hu X, Bu Z, Peng J, Ren X, et al. Immune phenotypic linkage between colorectal cancer and liver metastasis. Cancer Cell. 2022;40(4):424-437.e425. Wu Y, Yang S, Ma J, Chen Z, Song G, Rao D, Cheng Y, Huang S, Liu Y, Jiang S, et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 2022;12(1):134–53. Thomassen I, van Gestel YR, Lemmens VE, de Hingh IH. Incidence, prognosis, and treatment options for patients with synchronous peritoneal carcinomatosis and liver metastases from colorectal origin. Dis Colon Rectum. 2013;56(12):1373–80. Garcia-Carbonero N, Martinez-Useros J, Li W, Orta A, Perez N, Carames C, Hernandez T, Moreno I, Serrano G, Garcia-Foncillas J. KRAS and BRAF mutations as prognostic and predictive biomarkers for standard chemotherapy response in metastatic colorectal cancer: a single institutional study. Cells. 2020. https://doi.org/10.3390/cells9010219. Lang H, Baumgart J, Heinrich S, Tripke V, Passalaqua M, Maderer A, Galle PR, Roth W, Kloth M, Moehler M. Extended molecular profiling improves stratification and prediction of survival after resection of colorectal liver metastases. Ann Surg. 2019;270(5):799–805. Tóth C, Sükösd F, Valicsek E, Herpel E, Schirmacher P, Renner M, Mader C, Tiszlavicz L, Kriegsmann J. Expression of ERCC1, RRM1, TUBB3 in correlation with apoptosis repressor ARC, DNA mismatch repair proteins and p53 in liver metastasis of colorectal cancer. Int J Mol Med. 2017;40(5):1457–65. Olsen LM, Fiehn AK, Hasselby JP. ERCC1 expression in advanced colorectal cancer and matched liver metastases. Pathol Res Pract. 2020;216(3): 152826. Aykut B, Ochs M, Radhakrishnan P, Brill A, Hocker H, Schwarz S, Weissinger D, Kehm R, Kulu Y, Ulrich A, et al. EMX2 gene expression predicts liver metastasis and survival in colorectal cancer. BMC Cancer. 2017;17(1):555. Aust N, Schule S, Altendorf-Hofmann AK, Chen Y, Knosel T, Dirsch O, Settmacher U, Weise A, Mrasek K, Liehr T. Loss of chromosome 4 correlates with better long-term survival and lower relapse rate after R0-resection of colorectal liver metastases. J Cancer Res Clin Oncol. 2013;139(11):1861–7. Rehman AH, Jones RP, Poston G. Prognostic and predictive markers in liver limited stage IV colorectal cancer. Eur J Surg Oncol. 2019;45(12):2251–6. Sun L, Liu X, Pan B, Hu X, Zhu Y, Su Y, Guo Z, Zhang G, Xu M, Xu X, et al. Serum exosomal miR-122 as a potential diagnostic and prognostic biomarker of colorectal cancer with liver metastasis. J Cancer. 2020;11(3):630–7. Xu H, Wang C, Song H, Xu Y, Ji G. RNA-Seq profiling of circular RNAs in human colorectal cancer liver metastasis and the potential biomarkers. Mol Cancer. 2019;18(1):8. Zhi Q, Wan D, Ren R, Xu Z, Guo X, Han Y, Liu F, Xu Y, Qin L, Wang Y. Circular RNA profiling identifies circ102049 as a key regulator of colorectal liver metastasis. Mol Oncol. 2021;15(2):623–41. Nassar FJ, Msheik ZS, Itani MM, Helou RE, Hadla R, Kreidieh F, Bejjany R, Mukherji D, Shamseddine A, Nasr RR, et al. Circulating miRNA as biomarkers for colorectal cancer diagnosis and liver metastasis. Diagnostics. 2021. https://doi.org/10.3390/diagnostics11020341. Kong H, Wu Y, Zhu M, Zhai C, Qian J, Gao X, Wang S, Hou Y, Lu S, Zhu H. Long non-coding RNAs: novel prognostic biomarkers for liver metastases in patients with early stage colorectal cancer. Oncotarget. 2016;7(31):50428–36. Yang X, Zhang S, He C, Xue P, Zhang L, He Z, Zang L, Feng B, Sun J, Zheng M. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol Cancer. 2020;19(1):46. Gao W, Chen Y, Yang J, Zhuo C, Huang S, Zhang H, Shi Y. Clinical perspectives on liquid biopsy in metastatic colorectal cancer. Front Genet. 2021;12: 634642. Allen JE, Saroya BS, Kunkel M, Dicker DT, Das A, Peters KL, Joudeh J, Zhu J, El-Deiry WS. Apoptotic circulating tumor cells (CTCs) in the peripheral blood of metastatic colorectal cancer patients are associated with liver metastasis but not CTCs. Oncotarget. 2014;5(7):1753–60. Song Z, Chen E, Qian J, Xu J, Cao G, Zhou W, Wang F, Chen M, Xu D, Wang X, et al. Serum chitinase activity prognosticates metastasis of colorectal cancer. BMC Cancer. 2019;19(1):629. Garcia-Alfonso P, Ferrer A, Gil S, Duenas R, Perez MT, Molina R, Capdevila J, Safont MJ, Castanon C, Cano JM, et al. Neoadjuvant and conversion treatment of patients with colorectal liver metastasis: the potential role of bevacizumab and other antiangiogenic agents. Target Oncol. 2015;10(4):453–65. Filip S, Vymetalkova V, Petera J, Vodickova L, Kubecek O, John S, Cecka F, Krupova M, Manethova M, Cervena K, et al. Distant metastasis in colorectal cancer patients-do we have new predicting clinicopathological and molecular biomarkers? a comprehensive review. Int J Mol Sci. 2020. https://doi.org/10.3390/ijms21155255. Alhumaid A, AlYousef Z, Bakhsh HA, AlGhamdi S, Aziz MA. Emerging paradigms in the treatment of liver metastases in colorectal cancer. Crit Rev Oncol Hematol. 2018;132:39–50. Saad AM, Abdel-Rahman O. Initial systemic chemotherapeutic and targeted therapy strategies for the treatment of colorectal cancer patients with liver metastases. Expert Opin Pharmacother. 2019;20(14):1767–75. Punt CJ, Koopman M, Vermeulen L. From tumour heterogeneity to advances in precision treatment of colorectal cancer. Nat Rev Clin Oncol. 2017;14(4):235–46. Sottoriva A, Kang H, Ma Z, Graham TA, Salomon MP, Zhao J, Marjoram P, Siegmund K, Press MF, Shibata D, et al. A Big Bang model of human colorectal tumor growth. Nat Genet. 2015;47(3):209–16. Singovski G, Bernal C, Kuciak M, Siegl-Cachedenier I, Conod A. Ruiz i Altaba A: in vivo epigenetic reprogramming of primary human colon cancer cells enhances metastases. J Mol Cell Biol. 2016;8(2):157–73. Hu M, Zhou X, Wang Y, Guan K, Huang L. Relaxin-FOLFOX-IL-12 triple combination therapy engages memory response and achieves long-term survival in colorectal cancer liver metastasis. J Control Release. 2020;319:213–21. Poureau P-G, Metges J-P. Fundamentals of digestive cancers immunology, especially gastric and hepatocellular carcinomasfondamentaux de l’immunologie des cancers digestifs (Gastriques et Hépatocellulaires). Oncologie. 2021;23(1):47–59. **ng C, Li H, Li RJ, Yin L, Zhang HF, Huang ZN, Cheng Z, Li J, Wang ZH, Peng HL. The roles of exosomal immune checkpoint proteins in tumors. Mil Med Res. 2021;8(1):56. Zhang Y, Song J, Zhao Z, Yang M, Chen M, Liu C, Ji J, Zhu D. Single-cell transcriptome analysis reveals tumor immune microenvironment heterogenicity and granulocytes enrichment in colorectal cancer liver metastases. Cancer Lett. 2020;470:84–94. Kalanxhi E, Meltzer S, Ree AH. Immune-modulating effects of conventional therapies in colorectal cancer. Cancers. 2020. https://doi.org/10.3390/cancers12082193. Chen DL, Lu YX, Zhang JX, Wei XL, Wang F, Zeng ZL, Pan ZZ, Yuan YF, Wang FH, Pelicano H, et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics. 2017;7(19):4836–49. Not applicable. This work was financially supported by grants from the National Key Research and Development Program of China (2021YFF1201300, 2022YFE0103600), the National Natural Science Foundation of China (No. 81872280, 82073094), the CAMS Innovation Fund for Medical Sciences (CIFMS)(2021-I2M-1-014), the Open Issue of State Key Laboratory of Molecular Oncology (No. SKL-KF-2021–16), and the Independent Issue of State Key Laboratory of Molecular Oncology (No. SKL-2021–16). YN have drafted the work and substantively revised it. WY contributed to the literature review. HQ and YS contributed to the supervision and supported final approval of the article. All authors read and approved the final manuscript. Not applicable. Not applicable. The authors declare that there are no conflicts of interest. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data. Niu, Y., Yang, W., Qian, H. et al. Intracellular and extracellular factors of colorectal cancer liver metastasis: a pivotal perplex to be fully elucidated.
Cancer Cell Int 22, 341 (2022). https://doi.org/10.1186/s12935-022-02766-w Received: Accepted: Published: DOI: https://doi.org/10.1186/s12935-022-02766-wAvailability of data and materials
Abbreviations
References
Acknowledgements
Funding
Author information
Authors and Affiliations
Contributions
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Consent for publication
Competing interests
Additional information
Publisher's Note
Rights and permissions
About this article
Cite this article
Keywords