Abstract
Background
Pulpitis is a common disease mainly caused by bacteria. Conventional approaches of diagnosing the state of dental pulp are mainly based on clinical symptoms, thereby harbor deficiencies. The accurate and rapid diagnosis of pulpitis is important for choosing the suitable therapy. The study aimed to identify pulpits related key genes by integrating micro-array data analysis and systems biology network-based methods such as weighted gene co-expression network analysis (WGCNA).
Methods
The micro-array data of 13 inflamed pulp and 11 normal pulp were acquired from Gene Expression Omnibus (GEO). WGCNA was utilized to establish a genetic network and categorize genes into diverse modules. Hub genes in the most associated module to pulpitis were screened out using high module group members (MM) methods. Pulpitis model in rat was constructed and iRoot BP plus was applied to cap pulp. Reverse transcription-quantitative polymerase chain reaction (RT-qPCR) was used for validation of hub genes.
Results
WGCNA was established and genes were categorized into 22 modules. The darkgrey module had the highest correlation with pulpitis among them. A total of 5 hub genes (HMOX1, LOX, ACTG1, STAT3, GNB5) were identified. RT-qPCR proved the differences in expression levels of HMOX1, LOX, ACTG1, STAT3, GNB5 in inflamed dental pulp. Pulp cap** reversed the expression level of HMOX1, LOX, ACTG1.
Conclusion
The study was the first to produce a holistic view of pulpitis, screen out and validate hub genes involved in pulpitis using WGCNA method. Pulp cap** using iRoot BP plus could reverse partial hub genes.
Similar content being viewed by others
Introduction
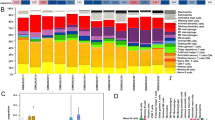
Inflammation of tooth pulp is known as pulpitis. As one of the most common dental diseases, pulpitis is triggered by various stimuli after destruction of the hard dental tissue surrounding pulp. When microbial incursion happens in pulp, balance between inflammation and reparative process damages. A mild inflammation in pulp is considered reversible pulpitis which can resolve and return to normal pulp after removing the etiology. If etiology is not removed, more immune reaction happens and the balance between damage and repair is broken in dental pulp, leading to uncontrollable inflammation and irreversible pulpitis. If treatment isn’t carried out in time, pulpitis could finally cause pulp necrosis, periapical periodontitis and other serious complications, leading to more medical and economic burden [7. The expressing levels of STAT3, HMOX1 and ACTG1 were higher in pulp of inflamed group than that of normal group significantly, but only levels of HMOX1 and ACTG1 were lower in pulp of iRoot group than that of Inflamed group. The expressing levels of LOX and GNB5 were lower in pulp of inflamed group in contrast to normal group significantly. Only LOX was higher in pulp of iRoot group than that of inflamed group.
Validation of hub genes and impact on hub gene expression of pulp cap** using iRoot BP plus. A–C Expressing levels of HMOX1, STAT3, and ACTG1 were considerably upregulated in inflamed pulp and pulp cap** using iRoot BP plus reverse the expressing level of HMOX1 and ACTG1. D, E Expressing levels of LOX and GNB5 were considerably downregulated in inflamed pulp and pulp cap** using iRoot BP plus reverse the expressing level of LOX. Wilcoxon test was applied to compare each two group
Discussion
In previous stuty, intersections of PPI network nodes and pulpitis-related genes were screened out in a merged dataset [38] found ACTG1 regulates the proteins of inflammation-related pathways, which indicated ACTG1 might serve as a new biomarker and treatment target of inflammation. GNB5 has widespread expression and is capable of encoding guanine nucleotide-binding protein sub-unit beta-5 (Gβ5). Central nervous system G-protein signal transmission is downregulated by Gβ5 through the interplay with G-protein–coupled acceptors. GNB5 knockdown mice have damaged neurological, retinal, and cardiac functions [39]. However, to our knowledge ACTG1 and GNB5 have not been studied in pulpitis yet. In present study, changes of expression levels of ACTG1 and GNB5 were observed which indicated they may be associated with the inflammation status of pulp. More in depth researches are needed to interrogate the effects of ACTG1 and GNB5 on pulpitis.
Pulp-cap** agents ought to harbor antibacteria, nontoxicity, antiinflammation and excellent sealing abilities, and ought to induce dentin mineralization [40]. The pulp cap** using Mineral Trioxide Aggregate (MTA) could reduce inflammation cells, decrease vascular density, and regulate IL-6 expression in mandibular first molars from Wistar rats [41]. iRoot BP Plus has shown similar properties and result with MTA in the pulpectomy in dog teeth [42] and displays a superior clinical handling property compared with MTA. Morepver, iRoot BP Plus possesses splendid compatibility, the capability of eliciting odontoblast differentiation and mineralization [43]. It's deemed as an appropriate alternative for calcium hydroxide in the pulpectomy of permanent teeth [43]. Clinically, iRoot BP Plus is quite prospective as a pulp-blocking agent. Therefore, we used iRoot BP plus as blocking agent to construct the model for VPT in the treatment of pulpitis. In present study, pulp cap** of iRoot BP plus could successfully rescue the inflammation state and reverse the expression level of some hub genes (HMOX1, ACTG1, LOX). It indicated that pulp cap** using iRoot BP plus could regulate inflammation and promote the ability of angiogenesis and odontogenesis. However, in our study, the levels of STAT3 and GNB5 were not reversed, and these two genes were less studied in pulpitis. Therefore, the role of STAT3 and GNB5 in pulpitis and material modification to regulate these two genes should be further investigated.
Conclusion
WGCNA was completed based on a pulpitis data set. Amongst the 22 modules, the darkgray module was determined as the most pivotal module for pulpitis, from which 5 core genes, A HMOX1, LOX, ACTG1, STAT3 and GNB5, were selected, and assumed to exert pivotal effects on the pathophysiologic causal links of pulpitis. The darkgray module was determined to be related to methyltransferase activity, triglyceride metabolism and positive modulation of pri-miRNA transcription from RNA polymerase II promotor. Those outcomes could foster further experiment researches on the function of the genes in pulpitis pathogenesis, which haven't been described yet. Moreover, the identified genes may be considered new treatment targets for the therapies of pulpitis, and help to further reveal the latent causal links regarding pulpitis.
RT-PCR proved the differences in expression levels of HMOX1, LOX, ACTG1, STAT3, GNB5 in inflamed dental pulp compared to healthy dental pulp. Pulp cap** reversed the expression level of HMOX1, LOX, ACTG1. More modification of the material properties is required to regulate the expression level of STAT3, GNB5 and restore the pulp to its normal physiological state.
Availability of data and materials
The datasets used for the current study are available from the corresponding author on reasonable request.
References
Zheng J, Wu Z, Niu K, **e Y, Hu X, Fu J, Tian D, Fu K, Zhao B, Kong W, et al. Microbiome of deep dentinal caries from reversible pulpitis to irreversible pulpitis. J Endod. 2019;45(3):302-309.e301.
Mejàre IA, Axelsson S, Davidson T, Frisk F, Hakeberg M, Kvist T, Norlund A, Petersson A, Portenier I, Sandberg H, et al. Diagnosis of the condition of the dental pulp: a systematic review. Int Endod J. 2012;45(7):597–613.
Zanini M, Meyer E, Simon S. Pulp inflammation diagnosis from clinical to inflammatory mediators: a systematic review. J Endod. 2017;43(7):1033–51.
Bunney PE, Zink AN, Holm AA, Billington CJ, Kotz CM. Orexin activation counteracts decreases in nonexercise activity thermogenesis (NEAT) caused by high-fat diet. Physiol Behav. 2017;176:139–48.
Guan X, Zhou Y, Yang Q, Zhu T, Chen X, Deng S, Zhang D. Vital pulp therapy in permanent teeth with irreversible pulpitis caused by caries: a prospective cohort study. J Person Med. 2021;11(11):1125.
Ricucci D, Siqueira JF Jr, Abdelsayed RA, Lio SG, Rôças IN. Bacterial invasion of pulp blood vessels in teeth with symptomatic irreversible pulpitis. J Endod. 2021;47(12):1854–64.
Ricucci D, Siqueira JF Jr, Li Y, Tay FR. Vital pulp therapy: histopathology and histobacteriology-based guidelines to treat teeth with deep caries and pulp exposure. J Dent. 2019;86:41–52.
Chen M, Zeng J, Yang Y, Wu B. Diagnostic biomarker candidates for pulpitis revealed by bioinformatics analysis of merged microarray gene expression datasets. BMC Oral Health. 2020;20(1):279.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 2008;9:559.
Yao Q, Song Z, Wang B, Qin Q, Zhang JA. Identifying key genes and functionally enriched pathways in sjögren’s syndrome by weighted gene co-expression network analysis. Front Genet. 2019;10:1142.
Wang P, Wang B, Zhang Z, Wang Z. Identification of inflammation-related DNA methylation biomarkers in periodontitis patients based on weighted co-expression analysis. Aging. 2021;13(15):19678–95.
** X, Ma Y, Xu Y, Ogbuehi AC, Liu X, Deng Y, ** J, Pan H, Lin Q, Li B, et al. The genetic and epigenetic mechanisms involved in irreversible pulp neural inflammation. Dis Markers. 2021;2021:8831948.
Dai Y, Sun X, Wang C, Li F, Zhang S, Zhang H, Li G, Yuan L, Chen G, Sun R, et al. Gene co-expression network analysis reveals key pathways and hub genes in Chinese cabbage (Brassica rapa L.) during vernalization. BMC Genomics. 2021;22(1):236.
Gagnon-Bartsch JA, Speed TP. Using control genes to correct for unwanted variation in microarray data. Biostatistics (Oxford, Engl). 2012;13(3):539–52.
Carvalho BS, Irizarry RA. A framework for oligonucleotide microarray preprocessing. Bioinformatics (Oxford, Engl). 2010;26(19):2363–7.
Smyth GK. Limma: linear models for microarray data. In: Gentleman R, Carey VJ, Huber W, Irizarry RA, Dudoit S, editors. Bioinformatics and computational biology solutions using R and bioconductor. New York: Springer; 2005. p. 397–420.
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57.
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8(Suppl 4):S11.
**n B, Lin Y, Tian H, Song J, Zhang L, Lv J. Identification of pulpitis-related potential biomarkers using bioinformatics approach. Comput Math Methods Med. 2021;2021:1808361.
Kodonas K, Fardi A, Gogos C, Economides N. Scientometric analysis of vital pulp therapy studies. Int Endod J. 2021;54(2):220–30.
Lundy T, Stanley HR. Correlation of pulpal histopathology and clinical symptoms in human teeth subjected to experimental irritation. Oral Surg Oral Med Oral Pathol. 1969;27(2):187–201.
Hahn CL, Liewehr FR. Relationships between caries bacteria, host responses, and clinical signs and symptoms of pulpitis. J Endod. 2007;33(3):213–9.
Galler KM, Weber M, Korkmaz Y, Widbiller M, Feuerer M. Inflammatory response mechanisms of the dentine-pulp complex and the periapical tissues. Int J Mol Sci. 2021;22(3):1480.
Lin LM, Ricucci D, Saoud TM, Sigurdsson A, Kahler B. Vital pulp therapy of mature permanent teeth with irreversible pulpitis from the perspective of pulp biology. Aust Endod J. 2020;46(1):154–66.
Emilia E, Neelakantan P. Biomarkers in the dentin-pulp complex: role in health and disease. J Clin Pediatr Dent. 2015;39(2):94–9.
Galicia JC, Henson BR, Parker JS, Khan AA. Gene expression profile of pulpitis. Genes Immun. 2016;17(4):239–43.
Huang X, Chen K. Differential expression of long noncoding RNAs in normal and inflamed human dental pulp. J Endod. 2018;44(1):62–72.
Bei Y, Tianqian H, Fanyuan Y, Haiyun L, Xueyang L, **g Y, Chenglin W, Ling Y. ASH1L suppresses matrix metalloproteinase through mitogen-activated protein kinase signaling pathway in pulpitis. J Endod. 2017;43(2):306-314.e302.
Welty FK. How do elevated triglycerides and low HDL-cholesterol affect inflammation and atherothrombosis? Curr Cardiol Rep. 2013;15(9):400.
Yu L, Zhang B, Deochand D, Sacta MA, Coppo M, Shang Y, Guo Z, Zeng X, Rollins DA, Tharmalingam B, et al. Negative elongation factor complex enables macrophage inflammatory responses by controlling anti-inflammatory gene expression. Nat Commun. 2020;11(1):2286.
Cintra LT, da Silva Facundo AC, Azuma MM, Sumida DH, Astolphi RD, Bomfim SR, Narciso LG, Gomes-Filho JE. Pulpal and periodontal diseases increase triglyceride levels in diabetic rats. Clin Oral Investig. 2013;17(6):1595–9.
Dzeletovic B, Stratimirovic DJ, Stojic D, Djukic LJ. Linear and nonlinear analysis of dental pulp blood flow oscillations in ageing. Int Endod J. 2020;53(8):1033–9.
Aminabadi NA, Parto M, Emamverdizadeh P, Jamali Z, Shirazi S. Pulp bleeding color is an indicator of clinical and histohematologic status of primary teeth. Clin Oral Investig. 2017;21(5):1831–41.
Moreno-Navarrete JM, Ortega F, Rodríguez A, Latorre J, Becerril S, Sabater-Masdeu M, Ricart W, Frühbeck G, Fernández-Real JM. HMOX1 as a marker of iron excess-induced adipose tissue dysfunction, affecting glucose uptake and respiratory capacity in human adipocytes. Diabetologia. 2017;60(5):915–26.
Wang B, Liu P, Huang H, Wang X, Zhang M, Huang J, Lu F, Chen J, Liu Y, Kang Z. Carbon dots up-regulate heme oxygenase-1 expression towards acute lung injury therapy. J Mater Chem B. 2021;9(43):9005–11.
Yu H, Pardoll D, Jove R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat Rev Cancer. 2009;9(11):798–809.
Bae WJ, Yi JK, Park J, Kang SK, Jang JH, Kim EC. Lysyl oxidase-mediated VEGF-induced differentiation and angiogenesis in human dental pulp cells. Int Endod J. 2018;51(3):335–46.
Wu T, Jia X, Feng H, Wu D. ACTG1 regulates intervertebral disc degeneration via the NF-κB-p65 and Akt pathways. Biochem Biophys Res Commun. 2021;545:54–61.
Lodder EM, DeNittis P, Koopman CD, Wiszniewski W, MouradeSouza CF, Lahrouchi N, Guex N, Napolioni V, Tessadori F, Beekman L, et al. GNB5 mutations cause an autosomal-recessive multisystem syndrome with sinus bradycardia and cognitive disability. Am J Hum Genetics. 2016;99(3):704–10.
Li Z, Cao L, Fan M, Xu Q. Direct pulp cap** with calcium hydroxide or mineral trioxide aggregate: a meta-analysis. J Endod. 2015;41(9):1412–7.
Lopes CS, Junqueira MA, Cosme-Silva L, Pegoraro COR, Garbelini CCD, Oliveira TM, Martins NS, Neves JDS, Sakai VT. Initial inflammatory response after the pulpotomy of rat molars with MTA or ferric sulfate. J Appl Oral Sci. 2019;27:e20180550.
Shi S, Bao ZF, Liu Y, Zhang DD, Chen X, Jiang LM, Zhong M. Comparison of in vivo dental pulp responses to cap** with iRoot BP Plus and mineral trioxide aggregate. Int Endod J. 2016;49(2):154–60.
Rao Q, Kuang J, Mao C, Dai J, Hu L, Lei Z, Song G, Yuan G. Comparison of iRoot BP plus and calcium hydroxide as pulpotomy materials in permanent incisors with complicated crown fractures: a retrospective study. J Endod. 2020;46(3):352–7.
Acknowledgements
We would like to thank Weiting Ge for the award of a grant that makes this project possible.
Funding
This study has been supported by the Major Science and Technology Project of Medical and Health of Zhejiang Province of China (WKJ-ZJ-2034) and Basic and Applied Basic Research Project of Affiliated Stomatological Hospital of Zhejiang University (5022299).
Author information
Authors and Affiliations
Contributions
DZ and CZ put forward the conception and design of the study, contributed to the acquisition of data, analysis and interpretation of data and drafted the article. TZ and FY contributed to the acquisition of data and revised it critically for important intellectual content. YZ directed and determined the topic, designed the study, revised and finally approved the manuscript. All authors have read and approved the final article.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The in vivo study was approved by the Ethics Committee for Animal Research at Zhejiang University (ethics approval number: ZJU20220168). In this study, all the methods were carried out in accordance with relevant guidelines and regulations of Declaration of Helsinki. All methods are reported in accordance with ARRIVE guidelines (https://arriveguidelines.org) for the reporting of animal experiments.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1.
Figure S1. PCA results after batch effect removal showed that the inflamed group can be clearly separated from the control group.
Additional file 2.
Table S1. The detailed sequences of the primers for each gene.
Additional file 3.
Detailed process of GO analysis.
Additional file 4.
Detailed process of hub gene detection.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zhang, D., Zheng, C., Zhu, T. et al. Identification of key module and hub genes in pulpitis using weighted gene co-expression network analysis. BMC Oral Health 23, 2 (2023). https://doi.org/10.1186/s12903-022-02638-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12903-022-02638-9