Abstract
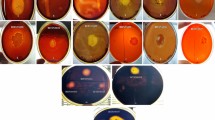
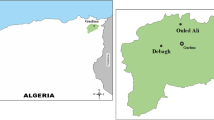
This study aims to determine the diversity of culturable thermophilic bacteria isolated from eight terrestrial hot springs in Northeastern of Algeria using the conventional methods, SDS-PAGE fingerprinting of whole-cell proteins and 16S rRNA gene sequencing. In addition, their hydrolytic enzyme activities were also investigated. A total of 293 strains were isolated from the hot springs’ water and sediment using different culture media. Overall, five distinct bacterial groups were characterized by whole-cell protein pattern analysis. Based on the 16S rRNA gene sequencing of 100 selected strains, the isolates were assigned to the following three major phyla: Firmicutes (93%), Deinococcus-Thermus (5%), and Actinobacteria (2%), which included 27 distinct species belonging to 12 different phylotypes, Aeribacillus, Aneurinibacillus, Anoxybacillus, Bacillus, Brevibacillus, Geobacillus, Laceyella, Meiothermus, Saccharomonospora, Thermoactinomyces, Thermobifida, and Thermus. The screening for nine extracellular enzymes showed that 65.87% of the isolates presented at least five types of enzyme activities, and 6.48% of strains combined all tested enzymes (amylase, cellulase, pectinase, esculinase, protease, gelatinase, lipase, lecithinase, and nuclease). It was found that Bacillus, Anoxybacillus, Aeribacillus, and Aneurinibacillus were the genera showing the highest activities. Likewise, the study showed an abundant and diverse thermophilic community with novel taxa presenting a promising source of thermozymes with important biotechnological applications. This study showed that a combined identification method using SDS-PAGE profiles of whole-cell proteins and subsequent 16S rRNA gene sequence analysis could successfully differentiate thermophilic bacteria from Algerian hot springs.









Similar content being viewed by others
References
Kumar S, Dangi AK, Shukla P, Baishya D, Khare SK (2019) Thermozymes: adaptive strategies and tools for their biotechnological applications. Bioresour Technol 278:372–382. https://doi.org/10.1016/j.biortech.2019.01.088
Panda AK, Bisht SS, De Mandal S, Kumar NS (2018) Microbial diversity of thermophiles through the lens of next generation sequencing, Microbial diversity in the genomic era. Elsevier Inc
Mahajan GB, Balachandran L (2017) Sources of antibiotics: hot springs. Biochem Pharmacol 134:35–41. https://doi.org/10.1016/j.bcp.2016.11.021
Mehetre GT, Dastager SG, Dharne MS (2019) Biodegradation of mixed polycyclic aromatic hydrocarbons by pure and mixed cultures of biosurfactant producing thermophilic and thermo-tolerant bacteria. Sci Total Environ 679:52–60. https://doi.org/10.1016/j.scitotenv.2019.04.376
Wang J, Salem DR, Sani RK (2019) Extremophilic exopolysaccharides: a review and new perspectives on engineering strategies and applications. Carbohydr Polym 205:8–26. https://doi.org/10.1016/j.carbpol.2018.10.011
Ilyas S, Lee J, Kim B (2014) Bioremoval of heavy metals from recycling industry electronic waste by a consortium of moderate thermophiles: process development and optimization. J Clean Prod 70:194–202. https://doi.org/10.3906/biy-1003-75
Rajkumari J, Bhuyan B, Das N, Pandey P (2019) Environmental applications of microbial extremophiles in the degradation of petroleum hydrocarbons in extreme environments. Environ Sustain 2:311–328. https://doi.org/10.1007/s42398-019-00065-1
Urbieta MS, Donati ER, Chan KG, Shahar S, Sin LL, Goh KM (2015) Thermophiles in the genomic era: biodiversity, science, and applications. Biotechnol Adv 33:633–647. https://doi.org/10.1016/j.biotechadv.2015.04.007
Sahay H, Yadav AN, Singh AK, Singh S, Kaushik R, Saxena AK (2017) Hot springs of Indian Himalayas: potential sources of microbial diversity and thermostable hydrolytic enzymes. 3 Biotech 7:1–11
Li L, Ma Z (2019) Global microbiome diversity scaling in hot springs with DAR (diversity-area relationship) profiles. Front Microbiol 10:118. https://doi.org/10.3389/fmicb.2019.00118
Stone ET, Murray R, College H, Moran MD (2018) Microbial diversity in the thermal springs within Hot Springs National Park. 72:31–37. https://scholarworks.uark.edu/jaas/vol72/iss1/9. Accessed 12 April 2020
Power JF, Carere CR, Lee CK, Evans DW, Button M, White D, Climo MD, Hinze AM, Morgan XC, Mcdonald IR, Cary SC, Stott MB (2018) Microbial biogeography of 1,000 geothermal springs in New Zealand. Nat Commun 9:2876. https://doi.org/10.1101/247759
Narsing Rao MP, Liu L, Jiao JY, **ao M, Li WJ (2018) Hot springs of India: occurrence and microbial diversity, pp 29–55. https://doi.org/10.1007/978-981-13-0329-6_2
Wilkins LGE, Ettinger CL, Jospin G, Eisen JA (2019) Metagenome-assembled genomes provide new insight into the microbial diversity of two thermal pools in Kamchatka, Russia. Sci Rep 9:1–15. https://doi.org/10.1038/s41598-019-39576-6
Ward LM, Idei A, Terajima S, Kakegawa T, Fischer WW, McGlynn SE (2017) Microbial diversity and iron oxidation at Okuoku-hachikurou Onsen, a Japanese hot spring analog of Precambrian iron formations. Geobiology 15:817–835. https://doi.org/10.1111/gbi.12266
Tang J, Liang Y, Jiang D, Li L, Luo Y, Shah MMR, Daroch M (2018) Temperature-controlled thermophilic bacterial communities in hot springs of western Sichuan, China. BMC Microbiol 18:1–14. https://doi.org/10.1186/s12866-018-1271-z
Tobler DJ, Benning LG (2011) Bacterial diversity in five Icelandic geothermal waters: temperature and sinter growth rate effects. Extremophiles 15:473–485. https://doi.org/10.1007/s00792-011-0378-z
Inan K, Çanakçi S, Beldü AO (2011) Isolation and characterization of xylanolytic new strains of Anoxybacillus from some hot springs in Turkey. Turk J Biol 35:529–542. https://doi.org/10.3906/biy-1003-75
Saibi H (2009) Geothermal resources in Algeria. Renew Sust Energ Rev 13:2544–2552. https://doi.org/10.1016/j.rser.2009.06.019
Stambouli AB, Khiat Z, Flazi S, Kitamura Y (2012) A review on the renewable energy development in Algeria: current perspective, energy scenario and sustainability issues. Renew Sust Energ Rev 16:4445–4460. https://doi.org/10.1016/j.rser.2012.04.031
Boukhenfouf W, Boucenna A (2019) Comparison of the radio-isotopic composition between hot spring water of Hammam Debagh and its associated deposits. Radiat Prot Dosim 187:369–377. https://doi.org/10.1093/rpd/ncz177
Fekraoui A, Kedaid F (2005) Geothermal resources and uses in Algeria: a country update report. Proc World Geothermal Congress:24–29. https://doi.org/10.1037/10320-002
Liang J, Kang D, Wang Y, Yu Y, Fan J, Takashi E (2015) Carbonate ion-enriched hot spring water promotes skin wound healing in nude rats. PLoS One 10:e0117106. https://doi.org/10.1371/journal.pone.0117106
Amarouche-Yala S, Benouadah A, El Ouahab BA, Moulla AS, Ouarezki SA, Azbouche A (2015) Physicochemical, bacteriological, and radiochemical characterization of some Algerian thermal spring waters. Water Qual Expo Health 7:233–249. https://doi.org/10.1007/s12403-014-0130-x
Benammar L, Menasria T, Chergui A, Benfiala S, Ayachi A (2017) Indoor fungal contamination of traditional public baths (Hammams). Int Biodeterior Biodegrad 117:115–122. https://doi.org/10.1016/j.ibiod.2016.12.004
Ait Ouali A, Issaadi A, Maizi D, Ayadi A, Bouhdjar A (2019) Geothermal potential in the Ouarsenis-Biban-Kabylie (North Central Algeria): hot spring catalogue. Arab J Geosci 12:741. https://doi.org/10.1007/s12517-019-4945-4
Arab M, Bakour S, Lalaoui R, Aissaoui N, Nas F, Hoceini A, Fournier P, Klouche-khelil N (2018) Diversity of aerobic bacilli analysis using molecular and culture-based approaches in Debagh hot spring diversity of aerobic bacilli analysis using molecular and culture-based. Geomicrobiol J 36:1–11. https://doi.org/10.1080/01490451.2018.1520937
Gomri MA, Khaldi TEM, Kharroub K (2018) Analysis of the diversity of aerobic, thermophilic endospore-forming bacteria in two Algerian hot springs using cultural and non-cultural methods. Ann Microbiol 68:915–929. https://doi.org/10.1007/s13213-018-1401-8
Kuisene N, Emantene R, Valiunas D, Chitavichus D (2002) Characterization of thermophilic spore-forming bacteria from a geothermal spring in Lithuania based on 16S rDNA and 16S-23S rDNA intergenic spacers analyses. Mikrobiologiia 71:824–828
Aanniz T, Ouadghiri M, Melloul M, Swings J, Elfahime E, Ibijbijen J, Ismaili M, Amar M (2015) Thermophilic bacteria in Moroccan hot springs, salt marshes and desert soils. Braz J Microbiol 46:443–453. https://doi.org/10.1590/S1517-838246220140219
Yohandini H, Julinar M (2015) Isolation and phylogenetic analysis of thermophile community within Tanjung Sakti Hot Spring, South Sumatera, Indonesia. HAYATI J Biosci 22:143–148. https://doi.org/10.1016/j.hjb.2015.10.006
Ming H, Ji WL, Li S, Zhao ZL, Zhan LY, Meng XL, Zhou EM, Nie GX, Li WJ (2017) Laceyella thermophila sp. nov., a thermophilic bacterium isolated from a hot spring. Int J Syst Evol Microbiol 67:2953–2958. https://doi.org/10.1099/ijsem.0.002057
Kreil DP, Ouzounis CA (2001) Identification of thermophilic species by the amino acid compositions deduced from their genomes. Nucleic Acids Res 29:1608–1615. https://doi.org/10.1093/nar/29.7.1608
Liu B, Li H, Wu S, Zhang X, **e L (2006) A simple and rapid method for the differentiation and identification of thermophilic bacteria. Can J Microbiol 52:753–758. https://doi.org/10.1139/w06-036
Brock TD, Freeze H (1969) Thermus aquaticus gen. n. and sp. n., a nonsporulating extreme thermophile. J Bacteriol 98:289–297
Degryse E, Glansdorff N, Piérard A (1978) A comparative analysis of extreme thermophilic bacteria belonging to the genus Thermus. Arch Microbiol 117:189–196. https://doi.org/10.1007/BF00402307
Oshima T, Imahori K (1974) Description of Thermus thermophilus (Yoshida and Oshima) comb. nov., a nonsporulating thermophilic bacterium from a Japanese thermal spa. Int J Syst Bacteriol 24:102–112. https://doi.org/10.1099/00207713-24-1-102
Adiguzel A, Ozkan H, Baris O, Inan K, Gulluce M, Sahin F (2009) Identification and characterization of thermophilic bacteria isolated from hot springs in Turkey. J Microbiol Methods 79:321–328. https://doi.org/10.1016/j.mimet.2009.09.026
Arya M, Joshi GK, Gupta AK, Kumar A, Raturi A (2015) Isolation and characterization of thermophilic bacterial strains from Soldhar (Tapovan) hot spring in Central Himalayan Region, India. Ann Microbiol 65:1457–1464. https://doi.org/10.1007/s13213-014-0984-y
Murray RGE, Doetsch RN, Robinow CF (1994) Determinative and cytological light microscopy. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 21–41
Gregersen T (1978) Method for the distinction of Gram-negative from Gram-positive bacteria. Eur J Appl Microbiol 5:123–127. https://doi.org/10.1007/BF01385437
Maugeri TL, Gugliandolo C, Caccamo D, Stackebrandt E (2001) A polyphasic taxonomic study of thermophilic bacilli from shallow, marine vents. Syst Appl Microbiol 24:572–587. https://doi.org/10.1078/0723-2020-00054
Munster MJ, Munster AP, Woodrow JR, Sharp RJ (1986) Isolation and preliminary taxonomic studies of Thermus strains isolated from Yellowstone National Park, USA. Microbiology 132:1677–1683. https://doi.org/10.1099/00221287-132-6-1677
Belduz AO, Dulger S, Demirbag Z (2003) Anoxybacillus gonensis sp. nov., a moderately thermophilic, xylose-utilizing, endospore-forming bacterium. Int J Syst Evol Microbiol 53:1315–1320. https://doi.org/10.1099/ijs.0.02473-0
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261. https://doi.org/10.1099/ijs.0.64915-0
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Marteinsson VT, Birrien JL, Jeanthon C, Prieur D (1996) Numerical taxonomic study of thermophilic Bacillus isolated from three geographically separated deep-sea hydrothermal vents. FEMS Microbiol Ecol 21:255–266. https://doi.org/10.1016/S0168-6496(96)00061-X
Kasana RC, Salwan R, Dhar H, Dutt S, Gulati A (2008) A rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Curr Microbiol 57:503–507. https://doi.org/10.1007/s00284-008-9276-8
Atlas RM (2005) Handbook of media for environmental microbiology, 2nd edn. CRC Press. Taylor & Francis Group
Soares MMCN, Da Silva R, Gomes E (1999) Screening of bacterial strains for pectinolytic activity: characterization of the polygalacturonase produced by Bacillus sp. Rev Microbiol 30:299–303. https://doi.org/10.1590/S0001-37141999000400002
Nabi N, Chaouachi M, Zellama MS, Ben Hafsa A, Mrabet B, Saïd K, Fathia HS (2016) A new QRT-PCR assay designed for the differentiation between elements provided from Agrobacterium sp. in GMOs plant events and natural Agrobacterium sp. bacteria. Food Chem 196:58–65. https://doi.org/10.1016/j.foodchem.2015.09.015
White D, Sharp RJ, Priest FG (1994) A polyphasic taxonomic study of thermophilic bacilli from a wide geographical area. Antonie Van Leeuwenhoek 64:357–386. https://doi.org/10.1007/BF00873093
Frazier WC (1926) A method for the detection of changes in gelatin due to bacteria. J Infect Dis 39:302–309. https://doi.org/10.1093/infdis/39.4.302
Gopinath SCB, Anbu P, Hilda A (2005) Extracellular enzymatic activity profiles in fungi isolated from oil-rich environments. Mycoscience 46:119–126. https://doi.org/10.1007/s10267-004-0221-9
Sierra G (1957) A simple method for the detection of lipolytic activity of micro-organisms and some observations on the influence of the contact between cells and fatty substrates. Antonie Van Leeuwenhoek 23:15–22. https://doi.org/10.1007/BF02545855
Priest FG, Goodfellow M, Todd C (1988) A numerical classification of the genus Bacillus. Microbiology 134:1847–1882. https://doi.org/10.1099/00221287-134-7-1847
Elder BL, Trujillo I, Blazevic DJ (1977) Rapid deoxyribonuclease test with methyl green. J Clin Microbiol 6:312–313
Magurran AE (2004) Measuring biological diversity. Wiley-Blackwell, Oxford
Cuecas A, Portillo MC, Kanoksilapatham W, Gonzalez JM (2014) Bacterial distribution along a 50 °C temperature gradient reveals a parceled out hot spring environment. Microb Ecol 68:729–739. https://doi.org/10.1007/s00248-014-0437-y
Pala C, Molari M, Nizzoli D, Bartoli M, Viaroli P, Manini E (2018) Environmental drivers controlling bacterial and archaeal abundance in the sediments of a Mediterranean lagoon ecosystem. Curr Microbiol 75:1147–1155. https://doi.org/10.1007/s00284-018-1503-3
Inan K, Ozer A, Guler HI et al (2016) Brevibacillus gelatini sp. nov., isolated from a hot spring. Int J Syst Evol Microbiol 66:712–718. https://doi.org/10.1099/ijsem.0.00078
Narayan VV, Hatha MA, Morgan HW, Rao D (2008) Isolation and characterization of aerobic thermophilic bacteria from the Savusavu Hot Springs in Fiji. Microbes Environ 23:350–352. https://doi.org/10.1264/jsme2.ME08105
Menasria T, Monteoliva-Sánchez M, Benammar L, Benhadj M, Ayachi A, Hacène H, Gonzalez-Paredes A, Aguilera M (2019) Culturable halophilic bacteria inhabiting Algerian saline ecosystems: a source of promising features and potentialities. World J Microbiol Biotechnol 35:132. https://doi.org/10.1007/s11274-019-2705-y
Eribe ERK, Olsen I (2002) SDS-PAGE of whole-cell proteins and random amplified polymorphic DNA (RAPD) analyses of Leptotrichia isolates. Microb Ecol Health Dis 14:193–202. https://doi.org/10.1080/08910600310002073
Kim TW, Kim YH, Kim SE, Lee JH, Park CS, Kim HY (2010) Identification and distribution of Bacillus species in Doenjang by whole-cell protein patterns and 16s rRNA gene sequence analysis. J Microbiol Biotechnol 20:1210–1214. https://doi.org/10.4014/jmb.1002.02008
Nicholson WL, Munakata N, Horneck G, Melosh HJ, Setlow P (2000) Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments. Microbiol Mol Biol Rev 64:548–572. https://doi.org/10.1128/mmbr.64.3.548-572.2000
Kumar M, Yadav AN, Tiwari R, Prasanna R, Saxena AK (2014) Deciphering the diversity of culturable thermotolerant bacteria from Manikaran hot springs. Ann Microbiol 64:741–751. https://doi.org/10.1007/s13213-013-0709-7
Spanevello MD, Patel BKC (2004) The phylogenetic diversity of Thermus and Meiothermus from microbial mats of an Australian subsurface aquifer runoff channel. FEMS Microbiol Ecol 50:63–73. https://doi.org/10.1016/j.femsec.2004.05.008
Tindall BJ, Sikorski J, Lucas S, Goltsman E, Copeland A, Glavina del Rio T, Nolan M, Tice H, Cheng JF, Han C, Pitluck S, Liolios K, Ivanova N, Mavromatis K, Ovchinnikova G, Pati A, Fähnrich R, Goodwin L, Chen A, Palaniappan K, Land M, Hauser L, Chang YJ, Jeffries CD, Rohde M, Göker M, Woyke T, Bristow J, Eisen JA, Markowitz V, Hugenholtz P, Kyrpides NC, Klenk HP, Lapidus A (2010) Complete genome sequence of Meiothermus ruber type strain (21T). Stand Genomic Sci 3:26–36. https://doi.org/10.4056/sigs.1032748
Liu L, Salam N, Jiao JY, Jiang HC, Zhou EM, Yin YR, Ming H, Li WJ (2016) Diversity of culturable thermophilic actinobacteria in hot springs in Tengchong, China and studies of their biosynthetic gene profiles. Microb Ecol 72:150–162. https://doi.org/10.1007/s00248-016-0756-2
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0
Kecha M, Benallaoua S, Touzel JP, Bonaly R, Duchiron F (2007) Biochemical and phylogenetic characterization of a novel terrestrial hyperthermophilic archaeon pertaining to the genus Pyrococcus from an Algerian hydrothermal hot spring. Extremophiles 11:65–73. https://doi.org/10.1007/s00792-006-0010-9
Bouanane-Darenfed A, Fardeau ML, Grégoire P, Joseph M, Kebbouche-Gana S, Benayad T, Hacene H, Cayol JL, Ollivier B (2011) Caldicoprobacter algeriensis sp. nov. a new thermophilic anaerobic, xylanolytic bacterium isolated from an Algerian hot spring. Curr Microbiol 62:826–832. https://doi.org/10.1007/s00284-010-9789-9
Mokrane S, Bouras N, Meklat A, Lahoum A, Zitouni A, Verheecke C, Mathieu F, Schumann P, Spröer C, Sabaou N, Klenk HP (2016) Thermoactinomyces khenchelensis sp. nov., a filamentous bacterium isolated from soil sediment of a terrestrial hot spring. Antonie van Leeuwenhoek. Int J Gen Mol Microbiol 109:311–317. https://doi.org/10.1007/s10482-015-0634-9
Tang YW, Ellis NM, Hopkins MK, Smith DH, Dodge DE, Persing DH (1998) Comparison of phenotypic and genotypic techniques for identification of unusual aerobic pathogenic gram-negative bacilli. J Clin Microbiol 36:3674–3679. https://doi.org/10.1128/JCM.36.12.3674-3679.1998
Janda JM, Abbott SL (2007) 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J Clin Microbiol 45:2761–2764. https://doi.org/10.1128/JCM.01228-07
Zeigler DR, Perkins JB (2008) The genus Bacillus. In: Goldman E, Green LH (eds) Practical handbook of microbiology, 2nd edn. CRC Press, New York, pp 309–337
Goh KM, Gan HM, Chan KG, Chan GF, Shahar S, Chong CS, Kahar UM, Chai KP (2014) Analysis of Anoxybacillus genomes from the aspects of lifestyle adaptations, prophage diversity, and carbohydrate metabolism. PLoS One 9:e90549. https://doi.org/10.1371/journal.pone.0090549
Burgess SA, Flint SH, Lindsay D, Cox MP, Biggs PJ (2017) Insights into the Geobacillus stearothermophilus species based on phylogenomic principles. BMC Microbiol 17:1–12. https://doi.org/10.1186/s12866-017-1047-x
Colak DN, Inan K, Karaoglu H, Canakcı S, Belduz AO (2012) Molecular analysis of the genus Anoxybacillus based on sequence similarity of the genes recN, flaA, and ftsY. Folia Microbiol (Praha) 57:61–69. https://doi.org/10.1007/s12223-011-0094-1
Panda AK, Bisht SS, De Mandal S, Kumar NS (2018) Microbial diversity of thermophiles through the lens of next generation sequencing. Elsevier Inc.
Singh AK, Tripathi BM, Sahay H, Singh RN, Kaushik R, Saxena AK, Arora DK (2010) Biochemical and molecular characterization of thermo-alkali tolerant xylanase producing bacteria from thermal springs of Manikaran. Indian J Microbiol 50:2–9. https://doi.org/10.1007/s12088-010-0071-4
Goh KM, Kahar UM, Chai YY, Chong CS, Chai KP, Ranjani V, Illias RM, Chan KG (2013) Recent discoveries and applications of Anoxybacillus. Appl Microbiol Biotechnol 97:1475–1488. https://doi.org/10.1007/s00253-012-4663-2
Menasria T, Aguilera M, Hocine H, Benammar L, Ayachi A, Si Bachir A, Dekak A, Monteoliva-Sánchez M (2018) Diversity and bioprospecting of extremely halophilic archaea isolated from Algerian arid and semi-arid wetland ecosystems for halophilic-active hydrolytic enzymes. Microbiol Res 207:289–298. https://doi.org/10.1016/j.micres.2017.12.011
Schallmey M, Singh A, Ward OP (2004) Developments in the use of Bacillus species for industrial production. Can J Microbiol 50:1–17. https://doi.org/10.1139/w03-076
Contesini FJ, de Melo RR, Sato HH (2018) An overview of Bacillus proteases: from production to application. Crit Rev Biotechnol 38:321–334. https://doi.org/10.1080/07388551.2017.1354354
de Souza PM, de Oliveira MP (2010) Application of microbial α-amylase in industry—a review. Braz J Microbiol 41:850–861. https://doi.org/10.1590/s1517-83822010000400004
Viksø-Nielsen A, Andersen C, Hoff T, Pedersen S (2006) Development of new α-amylases for raw starch hydrolysis. Biocatal Biotransform 24:121–127. https://doi.org/10.1080/10242420500519191
Han H, Ling Z, Khan A, Virk AK, Kulshrestha S, Li X (2019) Improvements of thermophilic enzymes: from genetic modifications to applications. Bioresour Technol 279:350–361. https://doi.org/10.1016/j.biortech.2019.01.087
Niehaus F, Bertoldo C, Kähler M, Antranikian G (1999) Extremophiles as a source of novel enzymes for industrial application. Appl Microbiol Biotechnol 51:711–729. https://doi.org/10.1007/s002530051456
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible Editor: Vania M.M. Melo
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 7.87 mb)
Rights and permissions
About this article
Cite this article
Benammar, L., İnan Bektaş, K., Menasria, T. et al. Diversity and enzymatic potential of thermophilic bacteria associated with terrestrial hot springs in Algeria. Braz J Microbiol 51, 1987–2007 (2020). https://doi.org/10.1007/s42770-020-00376-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-020-00376-0