Abstract
Background
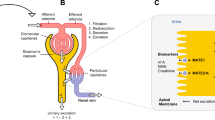
Endogenous biomarkers are promising tools to assess transporter-mediated drug–drug interactions early in humans.
Methods
We evaluated on a common and validated in vitro system the selectivity of 4-pyridoxic acid (PDA), homovanillic acid (HVA), glycochenodeoxycholate-3-sulphate (GCDCA-S) and taurine towards different renal transporters, including multidrug resistance-associated protein, and assessed the in vivo biomarker sensitivity towards the strong organic anion transporter (OAT) inhibitor probenecid at 500 mg every 6 h to reach close to complete OAT inhibition.
Results
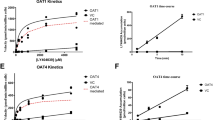
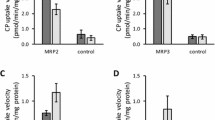
PDA and HVA were substrates of the OAT1/2/3, OAT4 (PDA only) and multidrug resistance-associated protein 4; GCDCA-S was more selective, having affinity only towards OAT3 and multidrug resistance-associated protein 2. Taurine was not a substrate of any of the investigated transporters under the in vitro conditions tested. Plasma exposure of PDA and HVA significantly increased and the renal clearance of GCDCA-S, PDA and HVA decreased; the magnitude of these changes was comparable to those of known clinical OAT probe substrates. PDA and GCDCA-S were the most promising endogenous biomarkers of the OAT pathway activity: PDA plasma exposure was the most sensitive to probenecid inhibition, and, in contrast, GCDCA-S was the most sensitive OAT biomarker based on renal clearance, with higher selectivity towards the OAT3 transporter.
Conclusions
The current findings illustrate a clear benefit of measuring PDA plasma exposure during phase I studies when a clinical drug candidate is suspected to be an OAT inhibitor based on in vitro data. Subsequently, combined monitoring of PDA and GCDCA-S in both urine and plasma is recommended to tease out the involvement of OAT1/3 in the inhibition interaction.
Clinical Trial Registration
EudraCT number: 2016-003923-49.





Similar content being viewed by others
Change history
17 February 2022
A Correction to this paper has been published: https://doi.org/10.1007/s40262-022-01109-2
References
US Food and Drug Administration. In vitro metabolism- and transporter-mediated drug-drug interaction studies. Guidance for industry. 2020. https://www.fda.gov/media/134582/download. Accessed 17 Feb 2021.
Chu X, Liao M, Shen H, Yoshida K, Zur AA, Arya V, et al. Clinical probes and endogenous biomarkers as substrates for transporter drug-drug interaction evaluation: perspectives from the International Transporter Consortium. Clin Pharmacol Ther. 2018;104(5):836–64.
EMEA. Guideline on the investigation of drug interactions. 2012 (cited 2020). https://www.ema.europa.eu/en/documents/scientific-guideline/guideline-investigation-drug-interactions_en.pdf. Accessed 18 Feb 2020.
Guest EJ, Rowland-Yeo K, Rostami-Hodjegan A, Tucker GT, Houston JB, Galetin A. Assessment of algorithms for predicting drug-drug interactions via inhibition mechanisms: comparison of dynamic and static models. Br J Clin Pharmacol. 2011;71(1):72–87.
Peters SA, Schroeder PE, Giri N, Dolgos H. Evaluation of the use of static and dynamic models to predict drug-drug interaction and its associated variability: impact on drug discovery and early development. Drug Metab Dispos. 2012;40(8):1495–507.
Rodrigues AD, Taskar KS, Kusuhara H, Sugiyama Y. Endogenous probes for drug transporters: balancing vision with reality. Clin Pharmacol Ther. 2018;103(3):434–48.
Taskar KS, Pilla Reddy V, Burt H, Posada MM, Varma M, Zheng M, et al. Physiologically-based pharmacokinetic models for evaluating membrane transporter mediated drug-drug interactions: current capabilities, case studies, future opportunities, and recommendations. Clin Pharmacol Ther. 2020;107(5):1082–115.
Guo Y, Chu X, Parrott NJ, Brouwer KLR, Hsu V, Nagar S, et al. Advancing predictions of tissue and intracellular drug concentrations using in vitro, imaging and physiologically based pharmacokinetic modeling approaches. Clin Pharmacol Ther. 2018;104(5):865–89.
Lai Y, Mandlekar S, Shen H, Holenarsipur VK, Langish R, Rajanna P, et al. Coproporphyrins in plasma and urine can be appropriate clinical biomarkers to recapitulate drug-drug interactions mediated by organic anion transporting polypeptide inhibition. J Pharmacol Exp Ther. 2016;358(3):397–404.
Kunze A, Njumbe Ediage E, Dillen L, Monshouwer M, Snoeys J. Clinical investigation of coproporphyrins as sensitive biomarkers to predict mild to strong OATP1B-mediated drug-drug interactions. Clin Pharmacokinet. 2018;57(12):1559–70.
Mori D, Kimoto E, Rago B, Kondo Y, King-Ahmad A, Ramanathan R, et al. Dose-dependent inhibition of OATP1B by rifampicin in healthy volunteers: comprehensive evaluation of candidate biomarkers and OATP1B probe drugs. Clin Pharmacol Ther. 2019;107(4):1004–13.
Barnett S, Ogungbenro K, Ménochet K, Shen H, Humphreys WG, Galetin A. Comprehensive evaluation of the utility of 20 endogenous molecules as biomarkers of OATP1B inhibition compared with rosuvastatin and coproporphyrin I. J Pharmacol Exp Ther. 2019;368(1):125–35.
Barnett S, Ogungbenro K, Ménochet K, Shen H, Lai Y, Humphreys WG, et al. Gaining mechanistic insight into coproporphyrin I as endogenous biomarker for OATP1B-mediated drug-drug interactions using population pharmacokinetic modeling and simulation. Clin Pharmacol Ther. 2018;104(3):564–74.
Shen H, Holenarsipur VK, Mariappan TT, Drexler DM, Cantone JL, Rajanna P, et al. Evidence for the validity of pyridoxic acid (PDA) as a plasma-based endogenous probe for OAT1 and OAT3 function in healthy subjects. J Pharmacol Exp Ther. 2019;368(1):136–45.
Shen H, Nelson DM, Oliveira RV, Zhang Y, Mcnaney CA, Gu X, et al. Discovery and validation of pyridoxic acid and homovanillic acid as novel endogenous plasma biomarkers of organic anion transporter (OAT) 1 and OAT3 in cynomolgus monkeys. Drug Metab Dispos. 2017;46:178–88.
Tsuruya Y, Kato K, Sano Y, Imamura Y, Maeda K, Kumagai Y, et al. Investigation of endogenous compounds applicable to drug-drug interaction studies involving the renal organic anion transporters, OAT1 and OAT3, in humans. Drug Metab Dispos. 2016;44(12):1925–33.
US Food and Drug Administration. Clinical drug interaction studies: study design, data analysis, and clinical implications. Guidance for industry. 2017. https://www.fda.gov/media/82734/download. Accessed 17 Feb 2020.
Takehara I, Terashima H, Nakayama T, Yoshikado T, Yoshida M, Furihata K, et al. Investigation of glycochenodeoxycholate sulfate and chenodeoxycholate glucuronide as surrogate endogenous probes for drug interaction studies of OATP1B1 and OATP1B3 in healthy Japanese volunteers. Pharm Res. 2017;34(8):1601–14.
Mori D, Kashihara Y, Yoshikado T, Kimura M, Hirota T, Matsuki S, et al. Effect of OATP1B1 genotypes on plasma concentrations of endogenous OATP1B1 substrates and drugs, and their association in healthy volunteers. Drug Metab Pharmacokinet. 2019;34(1):78–86.
Njumbe Ediage E, Dillen L, Vroman A, Diels L, Kunze A, Snoeys J, et al. Development of an LC-MS method to quantify coproporphyrin I and III as endogenous biomarkers for drug transporter-mediated drug-drug interactions. J Chromatogr B Analyt Technol Biomed Life Sci. 2018;1073:80–9.
Ahmad A, Ogungbenro K, Kunze A, Jacobs F, Snoeys J, Rostami-Hodjegan A, et al. Population pharmacokinetic modelling and simulation to support qualification of pyridoxic acid as endogenous biomarker of OAT1/3 renal transporters. CPT Pharmacometrics Syst Pharmacol. 2021. https://doi.org/10.1002/psp4.12610.
Emanuelsson BM, Beermann B, Paalzow LK. Non-linear elimination and protein binding of probenecid. Eur J Clin Pharmacol. 1987;32(4):395–401.
Maeda K, Tian Y, Fujita T, Ikeda Y, Kumagai Y, Kondo T, et al. Inhibitory effects of p-aminohippurate and probenecid on the renal clearance of adefovir and benzylpenicillin as probe drugs for organic anion transporter (OAT) 1 and OAT3 in humans. Eur J Pharm Sci. 2014;59:94–103.
US Food and Drug Administration. Drug development and drug interactions: table of substrates, inhibitors and inducers. 2020. https://www.fda.gov/drugs/drug-interactions-labeling/drug-development-and-drug-interactions-table-substrates-inhibitors-and-inducers#table5-2. Accessed 17 Feb 2021.
Thakare R, Gao H, Kosa RE, Bi YA, Varma MVS, Cerny MA, et al. Leveraging of rifampicin-dosed cynomolgus monkeys to identify bile acid 3-O-sulfate conjugates as potential novel biomarkers for organic anion-transporting polypeptides. Drug Metab Dispos. 2017;45(7):721–33.
Chesney RW, Han X, Patters AB. Taurine and the renal system. J Biomed Sci. 2010;17(Suppl 1):S4.
Acknowledgements
The authors thank Syneos Health who provided the bioanalytical method of probenecid and Emmanuel Njumbe Ediage for bioanalysis assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by Janssen Pharmaceutical Companies of Johnson & Johnson. Thomas K. Van Der Made is supported by a PhD studentship funded by Janssen Pharmaceutical Companies of Johnson & Johnson.
Conflict of interest
Marie-Emilie Willemin, Ils Pijpers, Lieve Dillen, Sophie Jonkers, Kathleen Steemans, An Tuytelaars, Frank Jacobs, Mario Monshouwer and Jan Snoeys are full-time employees of Janssen Pharmaceutical Companies of Johnson & Johnson.
Ethics approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent for publication
All research participants provided informed consent for publication.
Availability of data and material
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
Author contributions
MEW, TVDM, IP, FJ and AG wrote the manuscript. MEW, TVDM, AK, FJ, MM, DS, AR, AG and JS designed the research. TVDM, IP, SJ, KS and AT performed the research. MEW, TVDM, IP, LD, AK, SJ, KS, AT, FJ and JS analysed the data.
Additional information
The original Online version of this article was revised: The online supplementary file has been incorrectly published.
Supplementary Information
Below is the link to the electronic supplementary material.
40262_2021_1004_MOESM1_ESM.pdf
Material and Methods of the manuscript “Clinical investigation on endogenous biomarkers to predict strong OAT-mediated drug–drug interactions” (PDF 241 kb)
Rights and permissions
About this article
Cite this article
Willemin, ME., Van Der Made, T.K., Pijpers, I. et al. Clinical Investigation on Endogenous Biomarkers to Predict Strong OAT-Mediated Drug–Drug Interactions. Clin Pharmacokinet 60, 1187–1199 (2021). https://doi.org/10.1007/s40262-021-01004-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-021-01004-2