Abstract
Background
Prognostic modeling of NK cell marker genes in patients with hepatocellular carcinoma based on single cell sequencing and transcriptome data analysis.
Methods
Marker genes of NK cells were analyzed according to single cell sequencing data of hepatocellular carcinoma. Univariate Cox regression, lasso regression analysis, and multivariate Cox regression were performed to estimate the prognostic value of NK cell marker genes. TCGA, GEO and ICGC transcriptomic data were applied to build and validate the model. Patients were divided into high and low risk groups based on the median risk score. XCELL, timer, quantitative sequences, MCP counter, EPIC, CIBERSORT and CIBERSORT-abs were performed to explore the relationship between risk score and tumor microenvironment in hepatocellular carcinoma. Finally the sensitivity of the model to chemotherapeutic agents was predicted.
Results
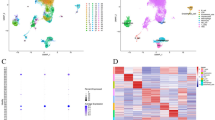
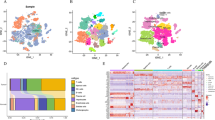
Single-cell sequencing identified 207 marker genes for NK cells in hepatocellular carcinoma. Enrichment analysis suggested that NK cell marker genes were mainly involved in cellular immune function. Eight genes were selected for prognostic modeling after multifactorial COX regression analysis. The model was validated in GEO and ICGC data. Immune cell infiltration and function were higher in the low-risk group than in the high-risk group. The low-risk group was more suitable for ICI and PD-1 therapy. Half-maximal inhibitory concentrations of Sorafenib, Lapatinib, Dabrafenib, and Axitinib were significantly different on the two risk groups.
Conclusion
A new signature of hepatocyte NK cell marker genes possesses a powerful ability to predict prognosis and immunotherapeutic response in patients with hepatocellular carcinoma.









Similar content being viewed by others
Data availability
Data used in this study can be downloaded from TCGA (https://tcga-data.nci.nih.gov/tcga/), GEO (https://www.ncbi.nlm.nih.gov/geo/) and ICGC (https://dcc.icgc.org/).
Abbreviations
- scRNA-seq:
-
Single-cell RNA sequencing
- RNA-seq:
-
RNA sequencing
- TCGA:
-
The Cancer Genome Atlas
- GEO:
-
Gene Expression Omnibus
- ICGC:
-
International Cancer Genome Consortium
- GSVA:
-
Gene set variation analysis
- TME:
-
Tumor microenvironment
- DEGs:
-
Differentially expressed genes
- ICB:
-
Immune checkpoint blockade
- KEGG:
-
Kyoko Gene and Genome Encyclopedia
- GO:
-
Gene ontology
- TMB:
-
Tumor mutational burden
- LASSO:
-
Least absolute shrinkage and selection operator
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
References
Bernardini G, Antonangeli F, Bonanni V, Santoni A (2016) Dysregulation of chemokine/chemokine receptor axes and NK cell tissue localization during diseases. Front Immunol 7:402
Bode D, Cull AH, Rubio-Lara JA, Kent DG (2021) Exploiting single-cell tools in gene and cell therapy. Front Immunol 12:702636
Brauning A, Rae M, Zhu G, Fulton E, Admasu TD, Stolzing A et al (2022) Aging of the immune system: focus on natural killer cells phenotype and functions. Cells 11(6):1017
Chidambaranathan-Reghupaty S, Fisher PB, Sarkar D (2021) Hepatocellular carcinoma (HCC): epidemiology, etiology and molecular classification. Adv Cancer Res 149:1–61
Crispe IN (2014) Immune tolerance in liver disease. Hepatology (baltimore, MD) 60(6):2109–2117
Dimitroulis D, Damaskos C, Valsami S, Davakis S, Garmpis N, Spartalis E et al (2017) From diagnosis to treatment of hepatocellular carcinoma: an epidemic problem for both developed and develo** world. World J Gastroenterol 23(29):5282–5294
Dong S, Guo X, Han F, He Z, Wang Y (2022) Emerging role of natural products in cancer immunotherapy. Acta Pharmaceutica Sinica b 12(3):1163–1185
Forner A, Reig M, Bruix J (2018) Hepatocellular carcinoma. Lancet (london, England) 391(10127):1301–1314
Fu Y, Liu S, Zeng S, Shen H (2019) From bench to bed: the tumor immune microenvironment and current immunotherapeutic strategies for hepatocellular carcinoma. J Exp Clin Cancer Res: CR 38(1):396
Gajewski TF, Schreiber H, Fu YX (2013) Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol 14(10):1014–1022
Ganesan P, Kulik LM (2023) Hepatocellular carcinoma: new developments. Clin Liver Dis 27(1):85–102
He S, Zhang J, Zhang W, Chen F, Luo R (2017) FOXA1 inhibits hepatocellular carcinoma progression by suppressing PIK3R1 expression in male patients. J Exp Clin Cancer Res: CR 36(1):175
Herberman RB, Nunn ME, Holden HT, Lavrin DH (1975) Natural cytotoxic reactivity of mouse lymphoid cells against syngeneic and allogeneic tumors. II. Characterization of effector cells. Int J Cancer 16(2):230–239
Hernaez R, Avila MA (2023) Immunogenomic classification of hepatocellular carcinoma patients for immune check-point inhibitors therapy: cui bono? Gut 72(1):7–9
Jovic D, Liang X, Zeng H, Lin L, Xu F, Luo Y (2022) Single-cell RNA sequencing technologies and applications: a brief overview. Clin Transl Med 12(3):e694
Khandelwal A, Crowley VM, Blagg BSJ (2016) Natural product inspired N-terminal Hsp90 inhibitors: from bench to bedside? Med Res Rev 36(1):92–118
Kiessling R, Klein E, Wigzell H (1975) “Natural” killer cells in the mouse. I. Cytotoxic cells with specificity for mouse Moloney leukemia cells. Specificity and distribution according to genotype. Eur J Immunol 5(2):112–117
Kulik L, El-Serag HB (2019) Epidemiology and management of hepatocellular carcinoma. Gastroenterology 156(2):477–91.e1
Lei Y, Tang R, Xu J, Wang W, Zhang B, Liu J et al (2021) Applications of single-cell sequencing in cancer research: progress and perspectives. J Hematol Oncol 14(1):91
Li XY, Shen Y, Zhang L, Guo X, Wu J (2022) Understanding initiation and progression of hepatocellular carcinoma through single cell sequencing. Biochim Biophys Acta 1877(3):188720
Liu F, Liu W, Zhou S, Yang C, Tian M, Jia G et al (2020) Identification of FABP5 as an immunometabolic marker in human hepatocellular carcinoma. J Immunother Cancer. 8(2):e000501
Marrero JA, Kulik LM, Sirlin CB, Zhu AX, Finn RS, Abecassis MM et al (2018) Diagnosis, staging, and management of hepatocellular carcinoma: 2018 practice guidance by the American association for the study of liver diseases. Hepatology (baltimore, MD) 68(2):723–750
Melnekoff DT, Laganà A (2022) Single-cell sequencing technologies in precision oncology. Adv Exp Med Biol 1361:269–282
Nagaraju GP, Dariya B, Kasa P, Peela S, El-Rayes BF (2022) Epigenetics in hepatocellular carcinoma. Semin Cancer Biol 86(Pt 3):622–632
Piñeiro Fernández J, Luddy KA, Harmon C, O’Farrelly C (2019) Hepatic tumor microenvironments and effects on NK cell phenotype and function. Int J Mol Sci 20(17):4131
Ramachandran P, Matchett KP, Dobie R, Wilson-Kanamori JR, Henderson NC (2020) Single-cell technologies in hepatology: new insights into liver biology and disease pathogenesis. Nat Rev Gastroenterol Hepatol 17(8):457–472
Regan PL, Jacobs J, Wang G, Torres J, Edo R, Friedmann J et al (2011) Hsp90 inhibition increases p53 expression and destabilizes MYCN and MYC in neuroblastoma. Int J Oncol 38(1):105–112
Robinson MW, Harmon C, O’Farrelly C (2016) Liver immunology and its role in inflammation and homeostasis. Cell Mol Immunol 13(3):267–276
Stabile H, Fionda C, Gismondi A, Santoni A (2017) Role of distinct natural killer cell subsets in anticancer response. Front Immunol 8:293
Stuart T, Satija R (2019) Integrative single-cell analysis. Nat Rev Genet 20(5):257–272
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin 71(3):209–249
Suvà ML, Tirosh I (2019) Single-cell RNA sequencing in cancer: lessons learned and emerging challenges. Mol Cell 75(1):7–12
Vujanovic L, Stahl EC, Pardee AD, Geller DA, Tsung A, Watkins SC et al (2017) Tumor-derived α-fetoprotein directly drives human natural killer-cell activation and subsequent cell death. Cancer Immunol Res 5(6):493–502
**ang X, You XM, Li LQ (2018) Expression of HSP90AA1/HSPA8 in hepatocellular carcinoma patients with depression. Onco Targets Ther 11:3013–3023
Xu Q, Tu J, Dou C, Zhang J, Yang L, Liu X et al (2017) HSP90 promotes cell glycolysis, proliferation and inhibits apoptosis by regulating PKM2 abundance via Thr-328 phosphorylation in hepatocellular carcinoma. Mol Cancer 16(1):178
Xu LX, He MH, Dai ZH, Yu J, Wang JG, Li XC et al (2019) Genomic and transcriptional heterogeneity of multifocal hepatocellular carcinoma. Ann Oncol: off J Eur Soc Med Oncol 30(6):990–997
Yang C, Shao Y, Wang X, Wang J, Wang P, Huang C et al (2023) The effect of the histone chaperones HSPA8 and DEK on tumor immunity in hepatocellular carcinoma. Int J Mol Sci 24(3):2653
Zhang Q, Lou Y, Yang J, Wang J, Feng J, Zhao Y et al (2019) Integrated multiomic analysis reveals comprehensive tumour heterogeneity and novel immunophenotypic classification in hepatocellular carcinomas. Gut 68(11):2019–2031
Zhang M, Yan X, Wen P, Bai W, Zhang Q (2021) CircANKRD52 promotes the tumorigenesis of hepatocellular carcinoma by sponging miR-497-5p and upregulating BIRC5 expression. Cell Transplant 30:9636897211008874
Zhuang W, Sun H, Zhang S, Zhou Y, Weng W, Wu B et al (2021) An immunogenomic signature for molecular classification in hepatocellular carcinoma. Mol Ther Nucleic Acids 25:105–115
Acknowledgements
None.
Funding
This study was funded by the National Key Research Programme of China (2022YFC2407304).
Author information
Authors and Affiliations
Contributions
Conceptualization: DY, FZ, YZ. Data curation: JS, YS. Formal analysis: DY, BY. Writing-original draft: DY, FZ, KZ. Writing—review and editing: YD, KZ.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, D., Zhao, F., Su, Y. et al. Integrated analysis of single-cell and bulk RNA-sequencing identifies a signature based on NK cell marker genes to predict prognosis and immunotherapy response in hepatocellular carcinoma. J Cancer Res Clin Oncol 149, 10609–10621 (2023). https://doi.org/10.1007/s00432-023-04965-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04965-y