Abstract
Porcine deltacoronavirus (PDCoV) is one of the most important enteropathogenic pathogens, and it causes enormous economic losses to the global commercial pork industry. PDCoV was initially reported in Hong Kong (China) in 2012 and subsequently emerged in swine herds with diarrhea in Ohio (USA) in 2014. Since then, it has spread to Canada, South Korea, mainland China, and several Southeast Asian countries. Information about the epidemiology, evolution, prevention, and control of PDCoV and its prevalence in China has not been comprehensively reported, especially in the last five years. This review is an update of current information on the general characteristics, epidemiology, geographical distribution, and evolutionary relationships, and the status of PDCoV vaccine development, focusing on the prevalence of PDCoV in China and vaccine research in particular. Together, this information will provide us with a greater understanding of PDCoV infection and will be helpful for establishing new strategies for controlling this virus worldwide.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronaviruses (CoVs) are a family of enveloped viruses with a positive-stranded RNA genome, and they can be genetically divided into four genera (Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus) that belong to the family Coronaviridae of the order Nidovirales [1].
CoVs are distributed widely among mammals and birds [2,3,4,5]. Individual CoVs usually infect their hosts in a species-specific manner, with alpha- and betacoronaviruses mainly infecting mammals, gammacoronaviruses normally infecting avian species, and deltacoronaviruses infecting both mammals and avian species. Attention to coronaviruses has increased in recent years because of the emergence of severe acute respiratory syndrome (SARS), Middle-East respiratory syndrome (MERS), and the newly emerging coronavirus disease 2019 (COVID-19), all of which cause acute respiratory illness, with mortality rates up to 9.5% in SARS and 35% in MERS, and with an inferred infection fatality rate varying from 0.00% to 1.63%, with corrected values varying from 0.00% to 1.54% for COVID-19 [6,7,8,9,10].
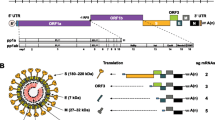
Porcine deltacoronavirus (PDCoV), a member of the genus Deltacoronavirus, is a novel swine enteropathogenic coronavirus that causes acute diarrhea, vomiting, and dehydration in neonatal piglets [11,12,13,14,15]. PDCoV was initially identified in 2012 during a molecular surveillance study in Hong Kong and emerged later in swine herds with diarrhea in Ohio (USA) in 2014 [2, 16]. Subsequently, the outbreak exhibited a global spread, and the virus has been detected in fecal samples from piglets in South Korea, mainland China, Thailand, Vietnam, and Laos [17,28, 29]. Downstream of ORF1a and ORF1b, there are several additional ORFs that code for the structural proteins: spike (S), envelope (E), membrane (M), nonstructural protein 6 (NS6), nucleocapsid (N), and nonstructural protein 7 (NS7) [17, 30]. The S protein forms peplomers on the virion surface and plays an important role in receptor attachment and viral and host cell membrane fusion [31,2]. Li et al. observed that PDCoV can efficiently infect cells with an unusually broad species range, including human and chicken cells [58]. Similar results were also reported in the comparative transcriptome profiling of human and pig intestinal epithelial cells after PDCoV infection [57]. Therefore, it appears inevitable that similar zoonotic events will occur again in the future, so more epidemiological studies need to be conducted to clarify the origin, epidemiology, and interspecific transmission mechanisms of coronaviruses.
Evolution
To date, 104 complete genome sequences of PDCoV isolates from the United States, China, Korea, Laos, Vietnam, Thailand, and Japan are available in the GenBank database. To investigate the molecular origin and evolution of PDCoV in mainland China, we performed a detailed phylogenetic analysis at the genome level using strains that have been reported in countries for the first time or for which epidemiological data show a high prevalence.
A total of 22 strains of porcine deltacoronavirus and seven avian isolates from seven countries were selected for phylogenetic analysis. Of these, 14 PDCoV isolates were from 13 different provinces of mainland China (Fig. 3).
Phylogenetic analysis of the complete genome sequences of 29 members of the genus Deltacoronavirus. The tree was constructed using the distance-based neighbor-joining method in MEGA7.0. Bootstrap analysis was carried out on 1000 replicate data sets, and values are shown adjacent to the branching points. Red represents the Chinese PDCoVs, blue represents the Vietnamese, Laotian, and Thai PDCoVs, yellow represents the US, Korean, and Japanese PDCoVs, green represents the Hong Kong (China) PDCoVs, and grey represents the avian deltacoronaviruses.
The phylogenetic analysis showed that the PDCoV isolates clustered together, while the avian deltacoronaviruses formed a separate cluster, and the evolutionary relationship between the two is distant. All porcine deltacoronavirus strains are closely related in the genetic evolution and have a high degree of sequence similarity, indicating that they might have originated from a common ancestor. The US, Korean, and Japanese PDCoV isolates grouped in the same branch with up to 99.9% nucleotide sequence identity and therefore might represent the same strain. The isolates from Vietnam, Laos, and Thailand belong to another branch with 98.4–99.8% whole-genome nucleotide sequence identity. It is clear that the Southeast Asian PDCoV isolates are much more closely related to the Chinese strains.
The nucleotide sequence identity of the 14 strains from mainland China ranged from 97.7% to 99.7%, with the isolates from Qinghai and Gansu and the Anhui strains belonging to the same branch with 99.1–99.5% nucleotide sequence identity. The isolates CHN/AH/2004, CHN/GS/2016, and CHN/QH/2017 were found to be more closely related to HKU15-44, while the Jiangsu isolate (CHN/Jiangsu/2014) was more closely related to HKU15-155.
Compared with the US, Korean, and Japanese PDCoV strains, most Chinese strains (except HKU15-44, CHN/AH/2004, CHN/GS/2016, and CHN/QH/2017) have a continuous deletion mutation of three nucleotides (AAT) at one site in the S gene. Previous reports have shown that the mutation rate of the S gene is relatively high, which may lead to altered tissue tropism, virulence, and even host specificity [27]. Whether the deletion of ATT has an effect on the virulence of the virus needs to be studied further.
In general, analysis of genetic evolution based on whole-genome sequencing shows that PDCoV has undergone extensive variation in different regions, and the mutations occur mainly in the S gene. Thus, it is important to monitor genetic variations occurring in the PDCoV S gene as well as to evaluate the impact of these variations on pathogenicity in order to develop an effective vaccine to control the disease.
Virulence and pathogenicity
Coronavirus infection has been documented previously in livestock and companion animals [55, 91]. TGEV and PEDV and the newly reported SADS-CoV mainly cause severe intestinal infections in piglets, leading to high morbidity and mortality and vast economic losses [88, 92, 93]. Bovine CoV, rat CoV, and infectious bronchitis virus (IBV) cause mild to severe respiratory tract infections in cattle, rats, and chickens, respectively [94,95,96]. Feline infectious peritonitis virus (FIPV) causes highly lethal disease in domestic cats [97].
Clinically, PDCoV can cause infection in pigs of various ages but mainly causes infection in newborn piglets, characterized by mild to severe diarrhea, vomiting, dehydration, anorexia, and growth retardation [13, 15, 98]. Inoculation experiments have suggested that although PDCoV exhibits enteropathogenicity in both gnotobiotic and conventional piglets, infected pigs display milder signs of clinical impact and disease severity than those infected with PEDV and TGEV [12, 24, 82].
Due to their underdeveloped immune system, neonatal piglets are highly susceptible to viral infection during their first few weeks. Mortality rates are highest in neonatal piglets, often reaching nearly 100%. Commercial fattening pigs and sows can also exhibit typical clinical features, such as diarrhea, inappetence, and persistent viral shedding in feces, but the symptoms of fattening pigs and sows are relatively mild, and the mortality rate is lower, with the animals gradually recovering [15].
Within 20–48 h postinfection, diarrhea is observed. Diarrhea typically lasts for at least 1 week. Vomiting symptoms were inconsistent in different experiments, which may be due to differences in virulence between strains. Core body temperatures remained within normal limits, and no respiratory signs were observed. Viral shedding peaked on day 7 postinfection, and virus was still detectable in feces and in the ileum at day 21 postinfection, which may enhance the risk for viral transmission [12,13,14, 21, 24, 52].
Pathological changes are characterized by intestinal villous atrophy and shortening, and villous changes are associated with extensive intestinal epithelial degeneration and necrosis. Gross lesions are observed in the small intestines, and no significant lesions are observed in extraintestinal tissues except the lung. PDCoV infection can cause mild interstitial pneumonia in gnotobiotic piglets [12], which has not been reported for PEDV or TGEV.
Vaccine and control strategies
Vaccines remain the most effective means to control coronavirus infections. However, there are no effective vaccines available for PDCoV. Strategies for PDCoV vaccine development include inactivated virus vaccines, subunit vaccines, viral vector vaccines, and live-attenuated virus vaccines, each of which has both advantages and disadvantages. Multiple routes of vaccine research are being evaluated for PDCoV and other enteric coronaviruses.
Inactivated virus vaccines
Inactivated virus vaccines use chemicals or radiation to render the virus noninfectious while preserving its antigenicity. The most recent PDCoV vaccine was developed by the State Key Laboratory of Veterinary Biotechnology in China. The vaccine is based on inactivated virus formulated with an adjuvant. When administered to seronegative sows using a prime/boost strategy 20 and 40 days before delivery, high levels of spike (S)-specific IgG and neutralizing antibody against PDCoV were found in colostrum and milk, as well as in the serum of piglets born to vaccinated sows [12]. Piglets were infected orally at 5 days of life with 105 TCID50 of PDCoV. The experiment showed that 87.1% of all piglets (n = 31) born to immunized sows were protected against lethal infection, and the infected piglets showed milder diarrhea, less viral shedding, and only minor damage to intestinal villi. In contrast, piglets from unimmunized sows had moderate diarrhea, which quickly worsened at 2 days postinfection and remained severe until the end of the experiment.
Live-attenuated vaccines
Nonreplicating vaccines (inactivated vaccines, subunit vaccines) usually generate short-lived neutralizing antibody responses with comparatively low titers. In contrast, live-attenuated vaccines are generally more immunogenic than nonreplicating vaccines; they can induce long-lasting immunity, produce a comprehensive spectrum of native viral antigens, and present antigens to the immune system in the same manner as in natural infections.
Zhang et al. [99] generated a full-length infectious cDNA clone of PDCoV, which they manipulated by replacing the NS6 gene with a green fluorescent protein (GFP) to generate rPDCoV-ΔNS6-GFP. Growth kinetics studies suggested that rPDCoV-ΔNS6-GFP showed a substantial reduction in viral replication in cell cultures and was highly attenuated in neonatal piglets, indicating that PDCoV lacking NS6 might be an ideal live-attenuated vaccine candidate.
Generally, live-attenuated virus vaccines are promising candidates for use against coronavirus infections, but they also have decreased safety and stability compared to inactivated vaccines, and some live-attenuated virus vaccines have the potential to spontaneously revert to virulence post-vaccination.
Vectored vaccines
Vectored vaccines function as viral gene delivery systems that rely on a host viral genome from a different virus, such as adenovirus, poxvirus, measles virus, parainfluenza virus, rabies virus, or vesicular stomatitis virus, and they have been used in the development of vaccines for CoVs [100,101,102,103,104,105,106].
Porcine adenovirus was used to deliver the core neutralizing epitope of PEDV, and this resulted in robust humoral and mucosal immune responses in piglets [100]. A recombinant vesicular stomatitis virus expressing the PEDV spike protein was developed, and sows immunized with this recombinant vaccine provided protective lactogenic immunity against a virulent G2b PEDV challenge to their piglets [104]. In addition, Yuan et al. used swinepox virus to express an epitope of the S protein of TGEV and a truncated spike protein of PEDV [101, 102].
Virus-like particles (VLPs)
Virus-like particles (VLPs) have drawn increasing attention in recent years. VLPs containing one or more viral structural proteins structurally resemble the native virus, can be easily recognized by antigen-presenting cells and B cells, and are capable of eliciting robust humoral and cell-mediated immune responses that are comparable to those achieved with inactivated or live-attenuated virus vaccines. There have been few reports about PDCoV VLPs, but studies of other animal coronavirus VLPs can provide a reference for PDCoV vaccine research.
Wang et al. [107] produced PEDV virus-like particles (VLPs) composed of S, M, and E proteins using a baculovirus expression system and showed that they induced a high level of anti-PEDV-neutralizing antibodies in mice. Xu et al. [108] developed chimeric IBV VLPs expressing M, E, and a recombinant S protein in baculoviruses. These induced a high level of IBV-specific antibodies and neutralizing antibodies that were comparable to those induced by an inactivated M41 virus via subcutaneous inoculation.
Moreover, other vaccine approaches for expressing the S, E, M, and N genes of two or more coronaviruses as well as other viral genes in bacteria, yeast, plants, and nanoparticles have been assessed [109,110,111,112,113,114,115,116]. However, the efficacy of these VLPs against lethal infection has not been tested in piglets, and further studies need to be performed.
Currently, most commercial vaccines for enteric coronaviruses are designed to induce lactogenic immunity by vaccinating the sow during pregnancy, and antibodies are passively transferred from sows to neonatal piglets via colostrum and milk.
Coronavirus infections are generally initiated at mucosal surfaces, and it is critical to induce localized intestinal sIgA and T cell immune responses to mucosal infections. For maternal immunity, oral vaccines or intentional infection of the sow may initiate the gut-mammary sIgA axis [117, 118].
A previous study showed that oral inoculation of sows with attenuated TGEV, followed by intramuscular injection with a recombinant subunit vaccine expressing the S protein of TGEV as a booster generated high titers of sIgA antibodies and neutralizing antibodies in colostrum and milk [119]. Similar prime/boost strategies can be applied to PDCoV vaccines to induce active immunity in newborn piglets.
Coronaviruses are an important group of pathogens that can have a devastating impact on humans and animals. New zoonotic coronaviruses are continually emerging or reemerging. In addition to good production management and strict biosecurity measures, the most effective way to control PDCoV is vaccination. Consequently, new vaccine development platforms and technologies are highly desirable, and further research will provide a better understanding of PDCoV replication and pathogenesis, a prerequisite for the development of new and promising vaccines to prevent, control, and ultimately eliminate the virus.
Availability of data and material
All data generated or analysed during this review are included in this published article.
References
De Groot R, Ziebuhr J, Poon L (2008) Revision of the family Coronaviridae
Woo PC, Lau SK, Lam CS, Lau CC, Tsang AK, Lau JH, Bai R, Teng JL, Tsang CC, Wang M, Zheng BJ, Chan KH, Yuen KY (2012) Discovery of seven novel mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J Virol 86(7):3995–4008. https://doi.org/10.1128/JVI.06540-11
Jung K, Hu H, Saif LJ (2017) Calves are susceptible to infection with the newly emerged porcine deltacoronavirus, but not with the swine enteric alphacoronavirus, porcine epidemic diarrhea virus. Arch Virol 162(8):2357–2362. https://doi.org/10.1007/s00705-017-3351-z
Vlasova AN, Wang Q, Jung K, Langel SN, Malik YS, Saif LJ (2020) Porcine coronaviruses. In: Malik YS, Singh RK, Yadav MP (eds) Emerging and transboundary animal viruses livestock diseases and management. Springer, Singapore, pp 79–110
Boley PA, Alhamo MA, Lossie G, Yadav KK, Vasquez-Lee M, Saif LJ, Kenney SP (2020) Porcine deltacoronavirus infection and transmission in poultry, United States. Emerg Infect Dis 26(2):255–265. https://doi.org/10.3201/eid2602.190346
Hilgenfeld R, Peiris M (2013) From SARS to MERS: 10 years of research on highly pathogenic human coronaviruses. Antiviral Res 100(1):286–295. https://doi.org/10.1016/j.antiviral.2013.08.015
Perlman S, Netland J (2009) Coronaviruses post-SARS: update on replication and pathogenesis. Nat Rev Microbiol 7(6):439–450. https://doi.org/10.1038/nrmicro2147
Chafekar A, Fielding BC (2018) MERS-CoV: understanding the latest human coronavirus threat. Viruses 10(2). https://doi.org/10.3390/v10020093
Giwa AL, Desai A, Duca A (2020) Novel 2019 coronavirus SARS-CoV-2 (COVID-19): an updated overview for emergency clinicians. Emerg Med Pract 22(5):1–28
Ioannidis JPA (2021) Infection fatality rate of COVID-19 inferred from seroprevalence data. 99:19-33F. WHO, Bulletin of the World Health Organization. https://doi.org/10.2471/BLT.20.265892
Xu Z, Zhong H, Zhou Q, Du Y, Chen L, Zhang Y, Xue C, Cao Y (2018) A highly pathogenic strain of porcine deltacoronavirus caused watery diarrhea in newborn piglets. Virol Sin 33(2):131–141. https://doi.org/10.1007/s12250-018-0003-8
Zhang J, Chen J, Liu Y, Da S, Shi H, Zhang X, Liu J, Cao L, Zhu X, Wang X, Ji Z, Feng L (2020) Pathogenicity of porcine deltacoronavirus (PDCoV) strain NH and immunization of pregnant sows with an inactivated PDCoV vaccine protects 5-day-old neonatal piglets from virulent challenge. Transbound Emerg Dis 67(2):572–583. https://doi.org/10.1111/tbed.13369
Vitosh-Sillman S, Loy JD, Brodersen B, Kelling C, Doster A, Topliff C, Nelson E, Bai J, Schirtzinger E, Poulsen E, Meadors B, Anderson J, Hause B, Anderson G, Hesse R (2016) Experimental infection of conventional nursing pigs and their dams with porcine deltacoronavirus. J Vet Diagn Invest 28(5):486–497. https://doi.org/10.1177/1040638716654200
Chen Q, Gauger P, Stafne M, Thomas J, Arruda P, Burrough E, Madson D, Brodie J, Magstadt D, Derscheid R, Welch M, Zhang J (2015) Pathogenicity and pathogenesis of a United States porcine deltacoronavirus cell culture isolate in 5-day-old neonatal piglets. Virology 482:51–59. https://doi.org/10.1016/j.virol.2015.03.024
Li B, Zheng L, Li H, Ding Q, Wang Y, Wei Z (2019) Porcine deltacoronavirus causes diarrhea in various ages of field-infected pigs in China. Biosci Rep 39(9). https://doi.org/10.1042/BSR20190676
Wang L, Byrum B (2014) Zhang Y (2014) Detection and genetic characterization of deltacoronavirus in pigs, Ohio, USA. Emerg Infect Dis 20(7):1227–1230. https://doi.org/10.3201/eid2007.140296
Lee S, Lee C (2014) Complete genome characterization of Korean porcine deltacoronavirus strain KOR/KNU14-04/2014. Genome Announc 2(6). https://doi.org/10.1128/genomeA.01191-14
Dong N, Fang L, Zeng S, Sun Q, Chen H, **ao S (2015) Porcine deltacoronavirus in mainland China. Emerg Infect Dis 21(12):2254–2255. https://doi.org/10.3201/eid2112.150283
Janetanakit T, Lumyai M, Bunpapong N, Boonyapisitsopa S, Chaiyawong S, Nonthabenjawan N, Kesdaengsakonwut S (2015) Amonsin A (2016) porcine deltacoronavirus, Thailand. Emerg Infect Dis 22(4):757–759. https://doi.org/10.3201/eid2204.151852
Saeng-Chuto K, Lorsirigool A, Temeeyasen G, Vui DT, Stott CJ, Madapong A, Tripipat T, Wegner M, Intrakamhaeng M, Chongcharoen W, Tantituvanont A, Kaewprommal P, Piriyapongsa J, Nilubol D (2017) Different lineage of porcine deltacoronavirus in Thailand, Vietnam and Lao PDR in 2015. Transbound Emerg Dis 64(1):3–10. https://doi.org/10.1111/tbed.12585
Zhao Y, Qu H, Hu J, Fu J, Chen R, Li C, Cao S, Wen Y, Wu R, Zhao Q, Yan Q, Wen X, Huang X (2019) Characterization and pathogenicity of the porcine deltacoronavirus isolated in Southwest China. Viruses 11(11):1074. https://doi.org/10.3390/v11111074
**a L, Yang Y, Wang J, **g Y, Yang Q (2018) Impact of TGEV infection on the pig small intestine. Virol J 15(1):102. https://doi.org/10.1186/s12985-018-1012-9
Dong N, Fang L, Yang H, Liu H, Du T, Fang P, Wang D, Chen H, **ao S (2016) Isolation, genomic characterization, and pathogenicity of a Chinese porcine deltacoronavirus strain CHN-HN-2014. Vet Microbiol 196:98–106. https://doi.org/10.1016/j.vetmic.2016.10.022
Ma Y, Zhang Y, Liang X, Lou F, Oglesbee M, Krakowka S, Li J (2015) Origin, evolution, and virulence of porcine deltacoronaviruses in the United States. mBio 6(2):e00064. https://doi.org/10.1128/mBio.00064-15
Suzuki T, Shibahara T, Imai N, Yamamoto T, Ohashi S (2018) Genetic characterization and pathogenicity of Japanese porcine deltacoronavirus. Infect Genet Evol 61:176–182. https://doi.org/10.1016/j.meegid.2018.03.030
Saeng-Chuto K, Jermsutjarit P, Stott CJ, Vui DT, Tantituvanont A, Nilubol D (2020) Retrospective study, full-length genome characterization and evaluation of viral infectivity and pathogenicity of chimeric porcine deltacoronavirus detected in Vietnam. Transbound Emerg Dis 67(1):183–198. https://doi.org/10.1111/tbed.13339
Song D, Zhou X, Peng Q, Chen Y, Zhang F, Huang T, Zhang T, Li A, Huang D, Wu Q, He H, Tang Y (2015) Newly emerged porcine deltacoronavirus associated with diarrhoea in swine in China: identification, prevalence and full-length genome sequence analysis. Transbound Emerg Dis 62(6):575–580. https://doi.org/10.1111/tbed.12399
Masters PS (2006) the molecular biology of coronaviruses. In: Advances in virus research, pp 193–292. https://doi.org/10.1016/s0065-3527(06)66005-3
Woo PC, Huang Y, Lau SK, Yuen KY (2010) Coronavirus genomics and bioinformatics analysis. Viruses 2(8):1804–1820. https://doi.org/10.3390/v2081803
Chen F, Zhu Y, Wu M, Ku X, Yao L, He Q (2015) Full-length genome characterization of chinese porcine deltacoronavirus strain CH/SXD1/2015. Genome Announc 3(5). https://doi.org/10.1128/genomeA.01284-15
Zhang J, Chen J, Shi D, Shi H, Zhang X, Liu J, Cao L, Zhu X, Liu Y, Wang X, Ji Z, Feng L (2019) Porcine deltacoronavirus enters cells via two pathways: a protease-mediated one at the cell surface and another facilitated by cathepsins in the endosome. J Biol Chem 294(25):9830–9843. https://doi.org/10.1074/jbc.RA119.007779
Zhu X, Liu S, Wang X, Luo Z, Shi Y, Wang D, Peng G, Chen H, Fang L, **ao S (2018) Contribution of porcine aminopeptidase N to porcine deltacoronavirus infection. Emerg Microbes Infect 7(1):65. https://doi.org/10.1038/s41426-018-0068-3
Chen R, Fu J, Hu J, Li C, Zhao Y, Qu H, Wen X, Cao S, Wen Y, Wu R, Zhao Q, Yan Q, Huang Y, Ma X, Han X, Huang X (2020) Identification of the immunodominant neutralizing regions in the spike glycoprotein of porcine deltacoronavirus. Virus Res 276:197834. https://doi.org/10.1016/j.virusres.2019.197834
Shang J, Zheng Y, Yang Y, Liu C, Geng Q, Tai W, Du L, Zhou Y, Zhang W, Li F (2018) Cryo-electron microscopy structure of porcine deltacoronavirus spike protein in the prefusion state. J Virol 92(4). https://doi.org/10.1128/JVI.01556-17
Li F (2016) Structure, function, and evolution of coronavirus spike proteins. Annu Rev Virol 3(1):237–261. https://doi.org/10.1146/annurev-virology-110615-042301
Schoeman D, Fielding BC (2019) Coronavirus envelope protein: current knowledge. Virol J 16(1):69. https://doi.org/10.1186/s12985-019-1182-0
McBride R, van Zyl M, Fielding BC (2014) The coronavirus nucleocapsid is a multifunctional protein. Viruses 6(8):2991–3018. https://doi.org/10.3390/v6082991
Lee S, Lee C (2015) Functional characterization and proteomic analysis of the nucleocapsid protein of porcine deltacoronavirus. Virus Res 208:136–145. https://doi.org/10.1016/j.virusres.2015.06.013
Likai J, Shasha L, Wenxian Z, **gjiao M, Jianhe S, Hengan W, Yaxian Y (2019) Porcine deltacoronavirus nucleocapsid protein suppressed IFN-beta production by interfering porcine RIG-I dsRNA-binding and K63-linked polyubiquitination. Front Immunol 10:1024. https://doi.org/10.3389/fimmu.2019.01024
Fang P, Fang L, Liu X, Hong Y, Wang Y, Dong N, Ma P, Bi J, Wang D, **ao S (2016) Identification and subcellular localization of porcine deltacoronavirus accessory protein NS6. Virology 499:170–177. https://doi.org/10.1016/j.virol.2016.09.015
Choi S, Lee C (2019) Functional characterization and proteomic analysis of porcine deltacoronavirus accessory protein NS7. J Microbiol Biotechnol 29(11):1817–1829. https://doi.org/10.4014/jmb.1908.08013
Fang P, Fang L, Hong Y, Liu X, Dong N, Ma P, Bi J, Wang D, **ao S (2017) Discovery of a novel accessory protein NS7a encoded by porcine deltacoronavirus. J Gen Virol 98(2):173–178. https://doi.org/10.1099/jgv.0.000690
Trudeau MP, Verma H, Sampedro F, Urriola PE, Shurson GC, Goyal SM (2017) Environmental persistence of porcine coronaviruses in feed and feed ingredients. PLoS One 12(5):e0178094. https://doi.org/10.1371/journal.pone.0178094
Cottingim KM, Verma H, Urriola PE, Sampedro F, Shurson GC, Goyal SM (2017) Feed additives decrease survival of delta coronavirus in nursery pig diets. Porcine Health Manage 3:5. https://doi.org/10.1186/s40813-016-0048-8
Trudeau MP, Verma H, Sampedro F, Urriola PE, Shurson GC, McKelvey J, Pillai SD, Goyal SM (2016) Comparison of thermal and non-thermal processing of swine feed and the use of selected feed additives on inactivation of porcine epidemic diarrhea virus (PEDV). PLoS One 11(6):e0158128. https://doi.org/10.1371/journal.pone.0158128
Environmental stability of PEDv (2014)
Quist-Rybachuk GV, Nauwynck HJ, Kalmar ID (2015) Sensitivity of porcine epidemic diarrhea virus (PEDV) to pH and heat treatment in the presence or absence of porcine plasma. Vet Microbiol 181(3–4):283–288. https://doi.org/10.1016/j.vetmic.2015.10.010
Zheng L, Li X, Yan M, Ren W, Zhang L, Lu C, Tian X, Han W (2017) Isolation, identification and biological characteristics analysis of porcine deltacoronavirus TJ1, China. Anim Husbandry Vet Med 45(1):219–224 (in Chinese)
Hulswit RJ, de Haan CA, Bosch BJ (2016) Coronavirus spike protein and tropism changes. Adv Virus Res 96:29–57. https://doi.org/10.1016/bs.aivir.2016.08.004
Graham RL, Baric RS (2010) Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission. J Virol 84(7):3134–3146. https://doi.org/10.1128/JVI.01394-09
Skowronski DM, Astell C, Brunham RC, Low DE, Petric M, Roper RL, Talbot PJ, Tam T, Babiuk L (2005) Severe acute respiratory syndrome (SARS): a year in review. Annu Rev Med 56:357–381. https://doi.org/10.1146/annurev.med.56.091103.134135
Jung K, Hu H, Eyerly B, Lu Z, Chepngeno J, Saif LJ (2015) Pathogenicity of 2 porcine deltacoronavirus strains in gnotobiotic pigs. Emerg Infect Dis 21(4):650–654. https://doi.org/10.3201/eid2104.141859
Torres CA, Hora AS, Tonietti PO, Taniwaki SA, Cecchinato M, Villarreal LY, Brandao PE (2016) Gammacoronavirus and deltacoronavirus in quail. Avian Dis 60(3):656–661. https://doi.org/10.1637/11412-032316-Reg.1
Banerjee A, Kulcsar K, Misra V, Frieman M, Mossman K (2019) bats and coronaviruses. Viruses 11(1). https://doi.org/10.3390/v11010041
Sowman HR, Cave NJ, Dunowska M (2018) A survey of canine respiratory pathogens in New Zealand dogs. N Z Vet J 66(5):236–242. https://doi.org/10.1080/00480169.2018.1490214
Liang Q, Zhang H, Li B, Ding Q, Wang Y, Gao W, Guo D, Wei Z, Hu H (2019) Susceptibility of chickens to porcine deltacoronavirus infection. Viruses 11(6). https://doi.org/10.3390/v11060573
Cruz-Pulido D, Boley PA, Ouma WZ, Alhamo MA, Saif LJ, Kenney SP (2021) Comparative transcriptome profiling of human and pig intestinal epithelial cells after porcine deltacoronavirus infection. Viruses 13(2). https://doi.org/10.3390/v13020292
Li W, Hulswit RJG, Kenney SP, Widjaja I, Jung K, Alhamo MA, van Dieren B, van Kuppeveld FJM, Saif LJ, Bosch BJ (2018) Broad receptor engagement of an emerging global coronavirus may potentiate its diverse cross-species transmissibility. Proc Natl Acad Sci USA 115(22):E5135–E5143. https://doi.org/10.1073/pnas.1802879115
Lee JH, Chung HC, Nguyen VG, Moon HJ, Kim HK, Park SJ, Lee CH, Lee GE, Park BK (2016) Detection and phylogenetic analysis of porcine deltacoronavirus in Korean swine farms, 2015. Transbound Emerg Dis 63(3):248–252. https://doi.org/10.1111/tbed.12490
Perez-Rivera C, Ramirez-Mendoza H, Mendoza-Elvira S, Segura-Velazquez R, Sanchez-Betancourt JI (2019) First report and phylogenetic analysis of porcine deltacoronavirus in Mexico. Transbound Emerg Dis 66(4):1436–1441. https://doi.org/10.1111/tbed.13193
Ajayi T, Dara R, Misener M, Pasma T, Moser L, Poljak Z (2018) Herd-level prevalence and incidence of porcine epidemic diarrhoea virus (PEDV) and porcine deltacoronavirus (PDCoV) in swine herds in Ontario, Canada. Transbound Emerg Dis 65(5):1197–1207. https://doi.org/10.1111/tbed.12858
Wang M, Wang Y, Baloch AR, Pan Y, Tian L, Xu F, Shivaramu S, Chen S, Zeng Q (2018) Detection and genetic characterization of porcine deltacoronavirus in Tibetan pigs surrounding the Qinghai-Tibet Plateau of China. Transbound Emerg Dis 65(2):363–369. https://doi.org/10.1111/tbed.12819
Zhang Y, Cheng Y, **ng G, Yu J, Liao A, Du L, Lei J, Lian X, Zhou J, Gu J (2019) Detection and spike gene characterization in porcine deltacoronavirus in China during 2016–2018. Infect Genet Evol 73:151–158. https://doi.org/10.1016/j.meegid.2019.04.023
Luo SX, Fan JH, Opriessnig T, Di JM, Liu BJ, Zuo YZ (2017) Development and application of a recombinant M protein-based indirect ELISA for the detection of porcine deltacoronavirus IgG antibodies. J Virol Methods 249:76–78. https://doi.org/10.1016/j.jviromet.2017.08.020
Su M, Li C, Guo D, Wei S, Wang X, Geng Y, Yao S, Gao J, Wang E, Zhao X, Wang Z, Wang J, Wu R, Feng L, Sun D (2016) A recombinant nucleocapsid protein-based indirect enzyme-linked immunosorbent assay to detect antibodies against porcine deltacoronavirus. J Vet Med Sci 78(4):601–606. https://doi.org/10.1292/jvms.15-0533
Ma L, Zeng F, Huang B, Cong F, Huang R, Ma J, Guo P (2018) Development of a conventional RT-PCR assay for rapid detection of porcine deltacoronavirus with the same detection limit as a SYBR green-based real-time RT-PCR assay. Biomed Res Int 2018:5035139. https://doi.org/10.1155/2018/5035139
Ding G, Fu Y, Li B, Chen J, Wang J, Yin B, Sha W, Liu G (2020) Development of a multiplex RT-PCR for the detection of major diarrhoeal viruses in pig herds in China. Transbound Emerg Dis 67(2):678–685. https://doi.org/10.1111/tbed.13385
Jia S, Feng B, Wang Z, Ma Y, Gao X, Jiang Y, Cui W, Qiao X, Tang L, Li Y, Wang L, Xu Y (2019) Dual priming oligonucleotide (DPO)-based real-time RT-PCR assay for accurate differentiation of four major viruses causing porcine viral diarrhea. Mol Cell Probes 47:101435. https://doi.org/10.1016/j.mcp.2019.101435
Li D, Feng H, Liu Y, Chen Y, Wei Q, Wang J, Liu D, Huang H, Su Y, Wang D, Cui Y, Zhang G (2018) Molecular evolution of porcine epidemic diarrhea virus and porcine deltacoronavirus strains in Central China. Res Vet Sci 120:63–69. https://doi.org/10.1016/j.rvsc.2018.06.001
Zhai SL, Wei WK, Li XP, Wen XH, Zhou X, Zhang H, Lv DH, Li F, Wang D (2016) Occurrence and sequence analysis of porcine deltacoronaviruses in southern China. Virol J 13:136. https://doi.org/10.1186/s12985-016-0591-6
Zhang F, Luo S, Gu J, Li Z, Li K, Yuan W, Ye Y, Li H, Ding Z, Song D, Tang Y (2019) Prevalence and phylogenetic analysis of porcine diarrhea associated viruses in southern China from 2012 to 2018. BMC Vet Res 15(1):470. https://doi.org/10.1186/s12917-019-2212-2
Zhang H, Liang Q, Li B, Cui X, Wei X, Ding Q, Wang Y, Hu H (2019) Prevalence, phylogenetic and evolutionary analysis of porcine deltacoronavirus in Henan province, China. Prev Vet Med 166:8–15. https://doi.org/10.1016/j.prevetmed.2019.02.017
Mai K, Feng J, Chen G, Li D, Zhou L, Bai Y, Wu Q, Ma J (2018) The detection and phylogenetic analysis of porcine deltacoronavirus from Guangdong Province in Southern China. Transbound Emerg Dis 65(1):166–173. https://doi.org/10.1111/tbed.12644
Zhang F, Song D, Zhou X, Huang D, Li A, Peng Q, Chen Y, Wu Q, He H, Tang Y (2016) Establishment and application of a RT-PCR assay for detection of newly emerged porcine deltacoronavirus. Sci Agric Sin 49(7):1406–1416 (in Chinese)
Huang X, Chen J, Yao G, Guo Q, Wang J, Liu G (2019) A TaqMan-probe-based multiplex real-time RT-qPCR for simultaneous detection of porcine enteric coronaviruses. Appl Microbiol Biotechnol 103(12):4943–4952. https://doi.org/10.1007/s00253-019-09835-7
Otter JA, Donskey C, Yezli S, Douthwaite S, Goldenberg SD, Weber DJ (2016) Transmission of SARS and MERS coronaviruses and influenza virus in healthcare settings: the possible role of dry surface contamination. J Hosp Infect 92(3):235–250. https://doi.org/10.1016/j.jhin.2015.08.027
Ding Y, He L, Zhang Q, Huang Z, Che X, Hou J, Wang H, Shen H, Qiu L, Li Z, Geng J, Cai J, Han H, Li X, Kang W, Weng D, Liang P, Jiang S (2004) Organ distribution of severe acute respiratory syndrome (SARS) associated coronavirus (SARS-CoV) in SARS patients: implications for pathogenesis and virus transmission pathways. J Pathol 203(2):622–630. https://doi.org/10.1002/path.1560
Fan C, Lei D, Fang C, Li C, Wang M, Liu Y, Bao Y, Sun Y, Huang J, Guo Y, Yu Y, Wang S (2020) Perinatal transmission of COVID-19 associated SARS-CoV-2: should we worry? Clin Infect Dis. https://doi.org/10.1093/cid/ciaa226
Alonso C, Goede DP, Morrison RB, Davies PR, Rovira A, Marthaler DG, Torremorell M (2014) Evidence of infectivity of airborne porcine epidemic diarrhea virus and detection of airborne viral RNA at long distances from infected herds. Vet Res 45:73. https://doi.org/10.1186/s13567-014-0073-z
Lowe J, Gauger P, Harmon K, Zhang J, Connor J, Yeske P, Loula T, Levis I, Dufresne L, Main R (2014) Role of transportation in spread of porcine epidemic diarrhea virus infection, United States. Emerg Infect Dis 20(5):872–874. https://doi.org/10.3201/eid2005.131628
Marthaler D, Raymond L, Jiang Y, Collins J, Rossow K, Rovira A (2014) Rapid detection, complete genome sequencing, and phylogenetic analysis of porcine deltacoronavirus. Emerg Infect Dis 20(8):1347–1350. https://doi.org/10.3201/eid2008.140526
Hu H, Jung K, Vlasova AN, Saif LJ (2016) Experimental infection of gnotobiotic pigs with the cell-culture-adapted porcine deltacoronavirus strain OH-FD22. Arch Virol 161(12):3421–3434. https://doi.org/10.1007/s00705-016-3056-8
Greiner L (2016) Evaluation of the likelihood of detection of porcine epidemic diarrhea virus or porcine delta coronavirus ribonucleic acid in areas within feed mills. J Swine Health Prod 24(4):198–204
Lam TT, Shum MH, Zhu HC, Tong YG, Ni XB, Liao YS, Wei W, Cheung WY, Li WJ, Li LF, Leung GM, Holmes EC, Hu YL, Guan Y (2020) Identifying SARS-CoV-2 related coronaviruses in Malayan pangolins. Nature. https://doi.org/10.1038/s41586-020-2169-0
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, **ao GF, Shi ZL (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798):270–273. https://doi.org/10.1038/s41586-020-2012-7
Hu B, Zeng LP, Yang XL, Ge XY, Zhang W, Li B, **e JZ, Shen XR, Zhang YZ, Wang N, Luo DS, Zheng XS, Wang MN, Daszak P, Wang LF, Cui J, Shi ZL (2017) Discovery of a rich gene pool of bat SARS-related coronaviruses provides new insights into the origin of SARS coronavirus. PLoS Pathog 13(11):e1006698. https://doi.org/10.1371/journal.ppat.1006698
Simas PV, Barnabe AC, Duraes-Carvalho R, Neto DF, Caserta LC, Artacho L, Jacomassa FA, Martini MC, Bianchi Dos Santos MM, Felippe PA, Ferreira HL, Arns CW (2015) Bat coronavirus in Brazil related to appalachian ridge and porcine epidemic diarrhea viruses. Emerg Infect Dis 21(4):729–731. https://doi.org/10.3201/eid2104.141783
Zhou P, Fan H, Lan T, Yang XL, Shi WF, Zhang W, Zhu Y, Zhang YW, **e QM, Mani S, Zheng XS, Li B, Li JM, Guo H, Pei GQ, An XP, Chen JW, Zhou L, Mai KJ, Wu ZX, Li D, Anderson DE, Zhang LB, Li SY, Mi ZQ, He TT, Cong F, Guo PJ, Huang R, Luo Y, Liu XL, Chen J, Huang Y, Sun Q, Zhang XL, Wang YY, **ng SZ, Chen YS, Sun Y, Li J, Daszak P, Wang LF, Shi ZL, Tong YG, Ma JY (2018) Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 556(7700):255–258. https://doi.org/10.1038/s41586-018-0010-9
Woo PC, Lau SK, Lam CS, Lai KK, Huang Y, Lee P, Luk GS, Dyrting KC, Chan KH, Yuen KY (2009) Comparative analysis of complete genome sequences of three avian coronaviruses reveals a novel group 3c coronavirus. J Virol 83(2):908–917. https://doi.org/10.1128/JVI.01977-08
Wang L, Su S, Bi Y, Wong G, Gao GF (2018) Bat-origin coronaviruses expand their host range to pigs. Trends Microbiol 26(6):466–470. https://doi.org/10.1016/j.tim.2018.03.001
Zhao S, Li W, Schuurman N, van Kuppeveld F, Bosch BJ, Egberink H (2019) Serological screening for coronavirus infections in cats. Viruses 11(8). https://doi.org/10.3390/v11080743
Wang D, Fang L, **ao S (2016) Porcine epidemic diarrhea in China. Virus Res 226:7–13. https://doi.org/10.1016/j.virusres.2016.05.026
Guo R, Fan B, Chang X, Zhou J, Zhao Y, Shi D, Yu Z, He K, Li B (2020) Characterization and evaluation of the pathogenicity of a natural recombinant transmissible gastroenteritis virus in China. Virology 545:24–32. https://doi.org/10.1016/j.virol.2020.03.001
Cavanagh D (2007) Coronavirus avian infectious bronchitis virus. Vet Res 38(2):281–297. https://doi.org/10.1051/vetres:2006055
Gomez DE, Arroyo LG, Poljak Z, Viel L, Weese JS (2017) Detection of bovine coronavirus in healthy and diarrheic dairy calves. J Vet Intern Med 31(6):1884–1891. https://doi.org/10.1111/jvim.14811
Haick AK, Rzepka JP, Brandon E, Balemba OB, Miura TA (2014) Neutrophils are needed for an effective immune response against pulmonary rat coronavirus infection, but also contribute to pathology. J Gen Virol 95(Pt 3):578–590. https://doi.org/10.1099/vir.0.061986-0
Tekes G, Thiel HJ (2016) Feline coronaviruses: pathogenesis of feline infectious peritonitis. Adv Virus Res 96:193–218. https://doi.org/10.1016/bs.aivir.2016.08.002
Curry SM, Gibson KA, Burrough ER, Schwartz KJ, Yoon KJ, Gabler NK (2017) Nursery pig growth performance and tissue accretion modulation due to porcine epidemic diarrhea virus or porcine deltacoronavirus challenge. J Anim Sci 95(1):173–181. https://doi.org/10.2527/jas.2016.1000
Zhang M, Li W, Zhou P, Liu D, Luo R, Jongkaewwattana A, He Q (2020) Genetic manipulation of porcine deltacoronavirus reveals insights into NS6 and NS7 functions: a novel strategy for vaccine design. Emerg Microbes Infect 9(1):20–31. https://doi.org/10.1080/22221751.2019.1701391
Do VT, Jang J, Park J, Dao HT, Kim K, Hahn TW (2020) Recombinant adenovirus carrying a core neutralizing epitope of porcine epidemic diarrhea virus and heat-labile enterotoxin B of Escherichia coli as a mucosal vaccine. Arch Virol 165(3):609–618. https://doi.org/10.1007/s00705-019-04492-7
Yuan X, Lin H, Li B, He K, Fan H (2017) Efficacy and immunogenicity of recombinant swinepox virus expressing the truncated S protein of a novel isolate of porcine epidemic diarrhea virus. Arch Virol 162(12):3779–3789. https://doi.org/10.1007/s00705-017-3548-1
Yuan X, Lin H, Fan H (2015) Efficacy and immunogenicity of recombinant swinepox virus expressing the A epitope of the TGEV S protein. Vaccine 33(32):3900–3906. https://doi.org/10.1016/j.vaccine.2015.06.057
Liniger M, Zuniga A, Tamin A, Azzouz-Morin TN, Knuchel M, Marty RR, Wiegand M, Weibel S, Kelvin D, Rota PA, Naim HY (2008) Induction of neutralising antibodies and cellular immune responses against SARS coronavirus by recombinant measles viruses. Vaccine 26(17):2164–2174. https://doi.org/10.1016/j.vaccine.2008.01.057
Ke Y, Yu D, Zhang F, Gao J, Wang X, Fang X, Wang H, Sun T (2019) Recombinant vesicular stomatitis virus expressing the spike protein of genotype 2b porcine epidemic diarrhea virus: a platform for vaccine development against emerging epidemic isolates. Virology 533:77–85. https://doi.org/10.1016/j.virol.2019.05.009
Bukreyev A, Lamirande EW, Buchholz UJ, Vogel LN, Elkins WR, St Claire M, Murphy BR, Subbarao K, Collins PL (2004) Mucosal immunisation of African green monkeys (Cercopithecus aethiops) with an attenuated parainfluenza virus expressing the SARS coronavirus spike protein for the prevention of SARS. Lancet 363(9427):2122–2127. https://doi.org/10.1016/S0140-6736(04)16501-X
Kato H, Takayama-Ito M, Iizuka-Shiota I, Fukushi S, Posadas-Herrera G, Horiya M, Satoh M, Yoshikawa T, Yamada S, Harada S, Fujii H, Shibamura M, Inagaki T, Morimoto K, Saijo M, Lim CK (2019) Development of a recombinant replication-deficient rabies virus-based bivalent-vaccine against MERS-CoV and rabies virus and its humoral immunogenicity in mice. PLoS One 14(10):e0223684. https://doi.org/10.1371/journal.pone.0223684
Wang C, Yan F, Zheng X, Wang H, ** H, Wang C, Zhao Y, Feng N, Wang T, Gao Y, Yang S, **a X (2017) Porcine epidemic diarrhea virus virus-like particles produced in insect cells induce specific immune responses in mice. Virus Genes 53(4):548–554. https://doi.org/10.1007/s11262-017-1450-2
Xu PW, Wu X, Wang HN, Ma BC, Ding MD, Yang X (2016) Assembly and immunogenicity of baculovirus-derived infectious bronchitis virus-like particles carrying membrane, envelope and the recombinant spike proteins. Biotechnol Lett 38(2):299–304. https://doi.org/10.1007/s10529-015-1973-3
Chen WH, Chag SM, Poongavanam MV, Biter AB, Ewere EA, Rezende W, Seid CA, Hudspeth EM, Pollet J, McAtee CP, Strych U, Bottazzi ME, Hotez PJ (2017) Optimization of the production process and characterization of the yeast-expressed SARS-CoV recombinant receptor-binding domain (RBD219-N1), a SARS vaccine candidate. J Pharm Sci 106(8):1961–1970. https://doi.org/10.1016/j.xphs.2017.04.037
Jiang X, Hou X, Tang L, Jiang Y, Ma G, Li Y (2016) A phase trial of the oral Lactobacillus casei vaccine polarizes Th2 cell immunity against transmissible gastroenteritis coronavirus infection. Appl Microbiol Biotechnol 100(17):7457–7469. https://doi.org/10.1007/s00253-016-7424-9
LeCureux JS, Dean GA (2018) Lactobacillus mucosal vaccine vectors: immune responses against bacterial and viral antigens. mSphere 3(3). https://doi.org/10.1128/mSphere.00061-18
Ma F, Zhang E, Li Q, Xu Q, Ou J, Yin H, Li K, Wang L, Zhao X, Niu X, Li X, Zhang S, Wang Y, Deng R, Zhou E, Zhang G (2020) A plant-produced recombinant fusion protein-based newcastle disease subunit vaccine and rapid differential diagnosis platform. Vaccines (Basel) 8(1). https://doi.org/10.3390/vaccines8010122
Michon C, Langella P, Eijsink VG, Mathiesen G, Chatel JM (2016) Display of recombinant proteins at the surface of lactic acid bacteria: strategies and applications. Microb Cell Fact 15:70. https://doi.org/10.1186/s12934-016-0468-9
Sirichokchatchawan W, Temeeyasen G, Nilubol D, Prapasarakul N (2018) Protective effects of cell-free supernatant and live lactic acid bacteria isolated from Thai pigs against a pandemic strain of porcine epidemic diarrhea virus. Probiotics Antimicrob Proteins 10(2):383–390. https://doi.org/10.1007/s12602-017-9281-y
Szatraj K, Szczepankowska AK, Chmielewska-Jeznach M (2017) Lactic acid bacteria—promising vaccine vectors: possibilities, limitations, doubts. J Appl Microbiol 123(2):325–339. https://doi.org/10.1111/jam.13446
Wang X, Wang Z, Xu H, **ang B, Dang R, Yang Z (2016) Orally administrated whole yeast vaccine against porcine epidemic diarrhea virus induced high levels of IgA response in mice and piglets. Viral Immunol 29(9):526–531. https://doi.org/10.1089/vim.2016.0067
Chattha KS, Roth JA, Saif LJ (2015) Strategies for design and application of enteric viral vaccines. Annu Rev Anim Biosci 3:375–395. https://doi.org/10.1146/annurev-animal-022114-111038
Gerdts V, Zakhartchouk A (2017) Vaccines for porcine epidemic diarrhea virus and other swine coronaviruses. Vet Microbiol 206:45–51. https://doi.org/10.1016/j.vetmic.2016.11.029
Park S, Sestak K, Hodgins DC, Shoup DI, Ward LA, Jackwood DJ, Saif LJ (1998) Immune response of sows vaccinated with attenuated transmissible gastroenteritis virus (TGEV) and recombinant TGEV spike protein vaccines and protection of their suckling pigs against virulent TGEV challenge exposure. Am J Vet Res 59(8):1002–1008
Acknowledgments
This work was supported by the State National Key Laboratory of Animal Genetic Engineering Vaccine (grant no. AGVSKL-ZD-202007) and Leading Talents in Science and Technology Innovation of Zhongyuan Thousand Talents Project in Henan Province (grant no. 204200510012).
Funding
This work was supported by the State National Key Laboratory of Animal Genetic Engineering Vaccine (grant no. AGVSKL-ZD-202007), and Leading Talents in Science and Technology Innovation of Zhongyuan Thousand Talents Project in Henan Province (grant no. 204200510012).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. The idea and original draft preparation came from Pan Tang and Enhui Cui. Shujuan Wang and Congcong Lei contributed to the literature search and data analysis. Ruoqian Yan and **gyu Wang provided critical review and substantially revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
No conflict of interest exits in the submission of this manuscript, and the manuscript was approved by all authors for publication.
Additional information
Handling Editor Tim Skern.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tang, P., Cui, E., Song, Y. et al. Porcine deltacoronavirus and its prevalence in China: a review of epidemiology, evolution, and vaccine development. Arch Virol 166, 2975–2988 (2021). https://doi.org/10.1007/s00705-021-05226-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05226-4