Abstract
Chronic kidney disease (CKD) is a major healthcare burden that takes a toll on the quality of life of many patients. Emerging evidence indicates that a substantial proportion of these patients carry a genetic defect that contributes to their disease. Any effort to reduce the percentage of patients with a diagnosis of nephropathy heading towards kidney replacement therapies should therefore be encouraged. Besides early genetic screenings and registries, in vitro systems that mimic the complexity and pathophysiological aspects of the disease could advance the screening for targeted and personalized therapies. In this regard, the use of patient-derived cell lines, as well as the generation of disease-specific cell lines via gene editing and stem cell technologies, have significantly improved our understanding of the molecular mechanisms underlying inherited kidney diseases. Furthermore, organs-on-chip technology holds great potential as it can emulate tissue and organ functions that are not found in other, more simple, in vitro models. The personalized nature of the chips, together with physiologically relevant read-outs, provide new opportunities for patient-specific assessment, as well as personalized strategies for treatment. In this review, we summarize the major kidney-on-chip (KOC) configurations and present the most recent studies on the in vitro representation of genetic kidney diseases using KOC-driven strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic kidney diseases are a major cause of chronic kidney disease (CKD) development and account for the majority of pediatric cases and ~ 10% of adult cases in need of kidney replacement therapy [1]. With the latest technological advances, like next generation sequencing and genome-wide association studies, it is expected that even more genetic cases will be identified. This is good news as in a majority of the cases a genetic diagnosis helps patients and doctors to gain insight into disease prognosis, treatment, or transplant decisions [2]. However, in many cases, there is still no therapy available to halt the disease progression and patients rely on kidney replacement therapy for their survival.
The kidney is a complex organ, home to multiple cell types with specific roles and functionalities. Their coordinated activity enables the clearance of metabolic waste products and balancing fluid and blood electrolytes. The nephron, the functional unit of the kidney, is comprised of different structural segments, each with its own assigned function. Every segment is susceptible to genetic mutations, with the majority of the genetic kidney disorders manifesting as glomerulopathies and tubulopathies. These disorders result in an overall malfunction of the kidney with life-threatening consequences. Although next-generation sequencing technologies can promote the clinical understanding of these genetic kidney diseases by providing guidance for molecular diagnostics and personalized treatment, this alone will not suffice. The combination of next-generation sequencing data and appropriate disease modeling will most likely provide a detailed vision of the complex genetic etiology. This was previously shown in a study where researchers found a variant of the PKHD1 gene, of which mutations cause polycystic kidney disease (PKD), and through in vitro testing of patient-derived cells were able to find ciliary defects in these cells, which were not detected earlier [3].
The development of disease models to understand the mechanisms of human genetic kidney disorders is of great interest. However, meeting the specificities of each disorder when proposing a model is not a trivial task. A plethora of disease models, ranging from zebrafish and C. elegans to rodents and in vitro patient-derived cell culturing, have contributed to the identification of novel genetic causes, new therapeutic targets, and to the development of new treatments. For a better overview of these models, we guide the reader to the work of Molinari et al. [4].
Although the ease of manipulation of in vitro cellular models allows the straightforward dissection of disease molecular mechanisms, the complexity of in vivo (animal) models supports the study of multiple cell interactions and tissue homeostasis in a (patho)physiologic context. Nevertheless, animal models do not fully genocopy or phenocopy human disease due to interspecies differences. They are too physiologically complex for a minimal reconstruction approach. The heterogeneity and the low frequency of certain kidney disorders prompt for the quest of more accurate and human patient-specific disease models [5]. With the current developments in (bio)manufacturing and the robust isolation and generation of disease-specific cells and organoids, in vitro disease models based on organs-on-chip (OOC) technology have been pushed to the forefront of scientific discoveries. OOC technology refers to microfluidic cell culture systems with controlled, dynamic conditions that directly replicate the microenvironment of tissues in the human body. OOC-based approaches are beyond traditional, flat, 2D cultures, as they exhibit tissue- and organ- level functions that are not found in other, more simple, in vitro models. Due to their physiologically relevant read-outs, OOC are used for preclinical drug testing and, more recently, as controlled, miniature representation of specific patients and diseases [5, 6]. In this review, we will first consider the major traits of genetic kidney diseases and discuss the cell-based in vitro models for their study. In this context, we will provide an overview of the current advances in the development of OOC-based in vitro models towards studying genetic kidney disease and discuss their potential use as tools to facilitate further insights into disease pathomechanisms and development of new therapeutical interventions.
Genetic kidney diseases
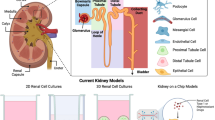
A large number of human genetic diseases (> 150) that affect kidney development or kidney tissue maintenance lead to functional and structural defects [1], as observed in most cases of CKD [7]. The range of phenotypes associated with genetic aberrations is remarkably wide. The renal cells that are most involved in the pathogenesis of various inherited kidney diseases include the following: (1) podocytes, the main performers of the renal glomerular filtration barrier, and (2) the (proximal) tubular epithelial cells of the nephron responsible of secretion, reabsorption and ion exchange [8]. A comprehensive overview of genetic kidney aberrations, their molecular and biological implications and associated pathological presentation has been prepared by others and is beyond the scope of the current review [9, 10]. As a summary, Fig. 1 includes the most common diseases and their altered phenotype throughout the segments of the nephron. The increased acceptance of early genetic testing and the establishment of national and international registries, allow for a real-time overview of genetic, medical, and family history. In this direction, the web-based registry, established by European Rare Kidney Disease Reference Network (ERKNet) (https://www.erknet.org), covers patients with rare kidney diseases, provides epidemiological and therapeutic management information, and includes patient cohorts for clinical research [11]. The in-depth documentation of new phenotypes and the isolation of patient-derived cells, made available to the scientific community, would support the development of disease-specific in vitro representations of the disease toward personalized therapeutic discoveries, with the ultimate goal to delay the onset of early-stage CKD [12].
Overview of the different segments of the nephron and their associated kidney genetic diseases. Glomerulus: congenital nephrotic syndrome (CNS) groups the genetic disorders that affect key glomerular filtration barrier (GFB) structural proteins. Slit diaphragms are affected by mutations in the genes NPHS1 (Finnish type CNS), NPHS2 (non-Finnish CNS) and WT1 (Denys-Drash syndrome) [7][7]. Within the GBM, the genetic disorders alter the encoding of collagen IV isoforms (Alport Syndrome, AS) [37], laminin (Pierson syndrome) [38], and LMX1B (less known, nail patella syndrome) [35]. Proximal tubule: formed by ciliated epithelial cells that are in close contact with capillaries. On the one hand, most of the disorders affecting the proximal tubule derive from aberrations in genes encoding for transporters of calcium, sodium, and potassium [50]. For instance, mutations in the genes CLCN5, OCRL1, causing Dent disease 1 and 2, respectively. Additionally, mutations in OCRL develop into Lowe syndrome [51]. On the other hand, mutations in genes encoding for key components in metabolic pathways are also associated with proximal tubulopathies. Defects in the CTNS gene lead to an abnormal intra-lysosomal accumulation of cystine that causes cystinosis and acidosis, which could eventually lead to Fanconi Syndrome and kidney failure. Distal tubule: the distal tubule epithelial cells express epithelial sodium channel (ENaC) and potassium channels in the apical membrane. Thus, genetic diseases often affect the regulation of Na–K reabsorption [60]. Mutations in the genes SCNN1B or SNCN1G have been linked to Liddle syndrome, characterized by excessive Na + reabsorption [67]. distal renal tubular acidosis (dRTA) is rooted in mutations in SLC4A1 [68]. The dominant pseudohypoaldosteronism type 1 (PHA1) is associated with NR3C3 mutations whilst recessive refers to SCNN1A, 1B and 1G [69]. Magnesium transporters are also subjected to genetic diseases, such as Bartter syndrome and its variant, Gitelman syndrome. While the former appears in the ascending limb of Henle’s loop, the latter is localized in the distal convoluted tube [70]. Gitelman has been associated with mutations in the SLC12A3 gene, which codes for the sodium-chloride cotransporter, and with CLCNKB gene, coding for renal chloride channel ClC-kb [70]. Collecting duct: the last segment of the nephron is in charge of water and Na + reabsorption. Aquaporin channels (AQP2) are also susceptible to mutations leading to dysfunction [72]
In vitro cell-based models of genetic kidney diseases
Several disease models (animal and cellular) for inherited kidney diseases have been developed to identify aberrant pathways and novel therapeutic targets, or as drug screening and testing platforms [4]. Deviating from animal models that are unable to fully recapitulate human pathophysiology, human models are becoming highly appealing [13]. In vitro cell cultures have been shown to be efficiently applied to dissect the molecular pathways of genetic diseases, while providing a fast and cheap platform for high-throughput drug screening. Special attention has been paid to identify new cell sources in such a way that patient-derived cells or cells with a predefined mutation can be tested. Hence, not only can we better understand the pathological manifestations of the disease, but also design targeted and personalized therapies towards the phenotype and/or repairing the defective gene [10]. The easy and non-invasive isolation of podocytes and proximal tubule epithelial cells (PTECs) from patient urine samples can generate primary kidney cell cultures that retain the genetic signature of the disease [10, 14]. However, these cells cannot be cultured indefinitely and/or lose their phenotypical signature and spontaneously dedifferentiate, mainly due to the lack of a kidney-like microenvironment. Via immortalization, their phenotype can be conserved. Similarly, cells can be obtained from patient’s kidney biopsies and subsequently immortalized; however, the invasive retrieval procedure and the lack of availability of kidney material make this approach challenging and less used in practice [10, 15]. Still, immortalized, patient-derived cell lines may lose their sensitivity to external stimuli, such as drugs, becoming a faux representation of an otherwise dynamic and highly responsive biological system. Moreover, since these cell lines cannot be generated from every single patient, they are solely a representation of one specific patient, and the extrapolation to a population of distinct and unique individuals is limited.
Using gene editing tools, such as clustered regularly interspaced short palindromic repeat (CRISPR/Cas) technology, a candidate mutation can be precisely introduced in healthy cells, allowing a fast and easy generation of mutation-specific diseased cells, paving the way toward precision medicine [16]. An example includes the recent application of CRISPR/Cas9 on a human PTECs line to generate a model for cystinosis that revealed novel insights into the molecular mechanisms of the disease and the potential therapeutic effect of a new combination treatment to alleviate the symptomatology of the disease [17]. Using the same technology, we can target and modify virtually any cell type. By introducing the correct version of the gene into the genome of diseased cells, reversing the disease is in sight. Despite the versatility and the specificity of CRISPR/Cas technology that make it a powerful tool for the generation of disease models in vitro, this gene-editing tool still harbors some limitations. The editing efficiency of the CRISPR/Cas system differs vastly among various cell types, which hampers the generation of disease models due to high optimization costs and time consuming experiments. Additionally, CRISPR/Cas editing presents relatively high off-target effects, which could result in a misleading disease phenotype if not properly evaluated. Lastly, some cells may be sensitive to the DNA damage induced by CRISPR/Cas which can trigger apoptosis, preventing the generation and further study of these disease models [18].
The discovery of human-induced pluripotent stem cells (hiPSCs) has been instrumental for the growth of the in vitro disease modeling field [19,20,21]. These indefinitely growing cells are suitable alternatives that overcome ethical concerns related to animal studies and reflect the donor’s genetic background. In short, pluripotency is induced in adult somatic cells using four retrovirally transfected transcription factors (the Yamanaka factors) [22]. Then, hiPSCs can be directed towards the desired cell type by a finely tuned combination of growth factors [23]. Using hiPSCs, it is possible to, theoretically, generate all cell types existing in the kidney, which otherwise cannot be obtained by employing classical isolation strategies. When hiPSCs are isolated from a donor that carries a specific gene mutation, this characteristic will be maintained in the differentiated cells. Furthermore, using healthy hiPSCs and employing gene editing tools, such as CRISPR/Cas, insertions and deletions can be performed, which result in ‘diseased’ kidney cells after differentiation. Creating isogenic models also avoids misinterpretation of the results derived from differences in genetic background among donors. Nevertheless, the fact that these cells possess phenotypic heterogeneity, tumorigenicity derived from undifferentiated iPSCs in the cell population, and above all, inefficient recapitulation of late-onset diseases due to the lack of maturity in iPSC-derived cells limits their application in personalised medicine [24]. To date, glomerular and proximal tubule cells have been successfully generated from hiPSCs [25,26,27].
Kidney-derived cells from hiPSCs and adult stem cells (ASCs) can also be cultured as 3D self-organized multi-cellular structures, known as organoids, which offer a more in vivo-like representation of the cellular heterogeneity resulting in a superior disease model when compared to 2D cell culture. ASCs isolated from kidney tissue and/or urine were used to generate tubuloids, organoids recapitulating the adult kidney tubular epithelium. These tubuloids have then been thought to be included in a biobank in which different patient-derived tubuloids can be used for the study of inherited kidney tubulopathies [28, 29]. Increasing the repertoire of disease-specific organoids will allow us to unravel the role of genotype in the disease process and whether common or divergent disease mechanisms occur in patients with the same diagnosis but different genetic context. The use of high content analysis methodologies, such as RNA sequencing and proteomics, enable the identification of molecular networks that are altered [30, 31]. Nevertheless, organoid maturation is limited, comparable to embryonic kidney during the first trimester, even if maintained for long periods in culture [32]. Thus, their use to model fully developed kidney diseases is limited. Even more, organoid cultures lack physical directional cues that drive the appropriate self-organization for the cells within the organ. To improve the level of maturity of the organoid system, biomechanical stimulation (via flow) and the introduction of vascular networks (via flow or host implantation) have been proposed. These features are crucial for the complete study of phenotype and tissue- and organ- level manifestation of the disease [33]. The implementation of microfabrication and microfluidic devices, such as OOC, enables the introduction of flow in vitro, allowing a better representation of the in vivo situation [75]. The team studied the infection with pseudorabies virus on electrolyte transport, particularly the Na+–Cl– cotransporter and Na+K+–ATPase. A successful analysis of the Na+ reabsorption on the chip supported the evidence that the beforementioned genetic alterations could be investigated further in this setup. The results proved that cells exposed to FSS showed better Na+ reabsorption, tighter cell junctions, more acetylated tubulin expression, and larger cell heights with more microvilli on the apical side, when compared to the static controls [75]. Moreover, this report is the first one of its kind that included viral infections. This proof of concept opens the door to viral transductions on-chip, a field which remained unexplored and could potentially lead to the establishment of more precise in vitro models.
Collecting duct
The collecting duct (CD) is the final segment of the nephron and allows urine to be excreted as waste. The main process in this segment is the reabsorption of water and Na+ [76]. To date, the only available OOC-based model was proposed by Jang et al. by culturing rat inter-medullary CD cells on a single-channel chip [77]. They established that FSS, the addition of arginine-vasopressin and osmolar differences between channels regulate cytoskeletal organization and aquaporin migration towards the apical domain. Moreover, the migration process was reversed after terminating fluidic stimulation [77].
The step forward: a combined KOC model
In the last few years, a substantial number of attempts to develop KOC have been proposed, including conceptual approaches based on computational models [78]. Nevertheless, the tendency is to move towards a system of chips in which each segment can be separately replicated and then connected. The combination of glomerular filtration and tubular reabsorption processes was proposed by Sakolish and Mahler who combined a biotic filter, mimicking glomerular filtration, with a PT chip [79]. Even though the glomerular segment lacked podocytes, their data supports that the combination of segments is not only possible, but also suitable to sustain enhanced junctions, baso-apical polarization, cytoskeletal reorganization and up-regulation of transport proteins. KOC has been combined with remote organs, including the liver, to establish systemic models [80]. A “body-on-a-chip” including kidney, intestine, liver, heart, lung, skin, blood–brain barrier, and brain has been recently proposed [81]. The addition of a diseased KOC in an otherwise healthy line-up of organs would provide further insight into how those genetic diseases that are largely limited to the kidney affect the activity of remote organs. Many kidney diseases manifest in other organs as well and, thus, in the “patient-on-a-chip” approach in which cells carrying the specific mutation, but representing different organs, are loaded on tissue-specific chips, the systemic implications of the disease could be recapitulated and therapeutically targeted [54, 79, 82, 83].
Conclusion
Kidney-on-chip emerges as a powerful tool for the in vitro representation of kidney function as it provides a much-needed dynamic platform for disease modeling and drug testing. From glomerulus to distal tubule, researchers have successfully proposed models for the nephron’s segments focusing on the replication of exchange barriers. Despite promising results and a technology readily and commercially available, the current approaches fall short when mimicking genetic diseases. Novel protocols for harvesting primary cells and differentiation of hiPSCs into individual kidney cell types, combined with gene editing, would shape our understanding of the kidney (patho-)physiology and aid in the development of novel therapies. A step towards the personalized approach has been taken. Accelerating the process is a matter of unifying efforts and strengthening the dialogue between biologists, engineers, geneticists, clinicians, and patients, early in the development of the models. Taken together, the unique capabilities of OOC technology can be used to develop new in vitro models of human genetic disease models.
References
Devuyst O, Knoers NV, Remuzzi G, Schaefer F et al (2014) Rare inherited kidney diseases: challenges, opportunities, and perspectives. Lancet 383:1844–1859. https://doi.org/10.1016/S0140-6736(14)60659-0
Besse W (2020) Genetic analysis in kidney disease: advancing clinical diagnosis and research discovery. Kidney360 1:720–723. https://doi.org/10.34067/kid.0003632020
Molinari E, Srivastava S, Dewhurst RM, Sayer JA (2020) Use of patient derived urine renal epithelial cells to confirm pathogenicity of PKHD1 alleles. BMC Nephrol 21:435. https://doi.org/10.1186/s12882-020-02094-z
Molinari E, Sayer JA (2020) Disease modeling to understand the pathomechanisms of human genetic kidney disorders. Clin J Am Soc Nephrol 15:855–872. https://doi.org/10.2215/CJN.08890719
van den Berg A, Mummery CL, Passier R, van der Meer AD (2019) Personalised organs-on-chips: functional testing for precision medicine. Lab Chip 19:198–205. https://doi.org/10.1039/c8lc00827b
Ashammakhi N, Wesseling-Perry K, Hasan A, Elkhammas E et al (2018) Kidney-on-a-chip: untapped opportunities. Kidney Int 94:1073–1086. https://doi.org/10.1016/j.kint.2018.06.034
Leeuwis JW, Nguyen TQ, Dendooven A, Kok RJ et al (2010) Targeting podocyte-associated diseases. Adv Drug Deliv Rev 62:1325–1336. https://doi.org/10.1016/j.addr.2010.08.012
Downie ML, Lopez Garcia SC, Kleta R, Bockenhauer D (2021) Inherited tubulopathies of the kidney: insights from genetics. Clin J Am Soc Nephrol 16:620–630. https://doi.org/10.2215/CJN.14481119
Hildebrandt F (2010) Genetic kidney diseases. Lancet 375:1287–1295. https://doi.org/10.1016/S0140-6736(10)60236-X
Bondue T, Arcolino FO, Veys KRP, Adebayo OC et al (2021) Urine-derived epithelial cells as models for genetic kidney diseases. Cells 10:1413. https://doi.org/10.3390/cells10061413
Bassanese G, Wlodkowski T, Servais A, Heidet L et al (2021) The European Rare Kidney Disease Registry (ERKReg): objectives, design and initial results. Orphanet J Rare Dis 16:251. https://doi.org/10.1186/s13023-021-01872-8
Stokman MF, Renkema KY, Giles RH, Schaefer F et al (2016) The expanding phenotypic spectra of kidney diseases: insights from genetic studies. Nat Rev Nephrol 12:472–483. https://doi.org/10.1038/nrneph.2016.87
Faria J, Ahmed S, Gerritsen KGF, Mihaila SM et al (2019) Kidney-based in vitro models for drug-induced toxicity testing. Arch Toxicol 93:3397–3418. https://doi.org/10.1007/s00204-019-02598-0
Ekulu PM, Adebayo OC, Decuypere JP, Bellucci L et al (2021) Novel human podocyte cell model carrying G2/G2 APOL1 high-risk genotype. Cells 10:1914. https://doi.org/10.3390/cells10081914
Little MH, Kumar SV, Forbes T (2019) Recapitulating kidney development: Progress and challenges. Semin Cell Dev Biol 91:153–168. https://doi.org/10.1016/j.semcdb.2018.08.015
WareJoncas Z, Campbell JM, Martinez-Galvez G, Gendron WAC et al (2018) Precision gene editing technology and applications in nephrology. Nat Rev Nephrol 14:663–677. https://doi.org/10.1038/s41581-018-0047-x
Jamalpoor A, van Gelder CA, Yousef Yengej FA, Zaal EA et al (2021) Cysteamine-bicalutamide combination therapy corrects proximal tubule phenotype in cystinosis. EMBO Mol Med 13:e13067. https://doi.org/10.15252/emmm.202013067
Yang Y, Xu J, Ge S, Lai L (2021) CRISPR/Cas: advances, limitations, and applications for precision cancer research. Front Med (Lausanne) 8:649896. https://doi.org/10.3389/fmed.2021.649896
Zhou T, Benda C, Duzinger S, Huang Y et al (2011) Generation of induced pluripotent stem cells from urine. J Am Soc Nephrol 22:1221–1228. https://doi.org/10.1681/ASN.2011010106
Zhou T, Benda C, Dunzinger S, Huang Y et al (2012) Generation of human induced pluripotent stem cells from urine samples. Nat Protoc 7:2080–2089. https://doi.org/10.1038/nprot.2012.115
Mulder J, Sharmin S, Chow T, Rodrigues DC et al (2020) Generation of infant- and pediatric-derived urinary induced pluripotent stem cells competent to form kidney organoids. Pediatr Res 87:647–655. https://doi.org/10.1038/s41390-019-0618-y
Takahashi K, Tanabe K, Ohnuki M, Narita M et al (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872. https://doi.org/10.1016/j.cell.2007.11.019
Low JH, Li P, Chew EGY, Zhou B et al (2019) Generation of human PSC-derived kidney organoids with patterned nephron segments and a de novo vascular network. Cell Stem Cell 25(373–387):e379. https://doi.org/10.1016/j.stem.2019.06.009
Doss MX, Sachinidis A (2019) Current challenges of iPSC-based disease modeling and therapeutic implications. Cells 8:403. https://doi.org/10.3390/cells8050403
Roye Y, Bhattacharya R, Mou X, Zhou Y et al (2021) A personalized glomerulus chip engineered from stem cell-derived epithelium and vascular endothelium. Micromachines (Basel) 12:967. https://doi.org/10.3390/mi12080967
Musah S, Mammoto A, Ferrante TC, Jeanty SSF et al (2017) Mature induced-pluripotent-stem-cell-derived human podocytes reconstitute kidney glomerular-capillary-wall function on a chip. Nat Biomed Eng 1:0069. https://doi.org/10.1038/s41551-017-0069
Musah S, Dimitrakakis N, Camacho DM, Church GM et al (2018) Directed differentiation of human induced pluripotent stem cells into mature kidney podocytes and establishment of a Glomerulus Chip. Nat Protoc 13:1662–1685. https://doi.org/10.1038/s41596-018-0007-8
Schutgens F, Rookmaaker M, Verhaar M (2021) A perspective on a urine-derived kidney tubuloid biobank from patients with hereditary tubulopathies. Tissue Eng Part C Methods 27:177–182. https://doi.org/10.1089/ten.TEC.2020.0366
Schutgens F, Rookmaaker MB, Margaritis T, Rios A et al (2019) Tubuloids derived from human adult kidney and urine for personalized disease modeling. Nat Biotechnol 37:303–313. https://doi.org/10.1038/s41587-019-0048-8
Taguchi A, Nishinakamura R (2017) Higher-order kidney organogenesis from pluripotent stem cells. Cell Stem Cell 21(730–746):e736. https://doi.org/10.1016/j.stem.2017.10.011
Little MH, Howden SE, Lawlor KT, Vanslambrouck JM (2022) Determining lineage relationships in kidney development and disease. Nat Rev Nephrol 18:8–21. https://doi.org/10.1038/s41581-021-00485-5
Wu H, Uchimura K, Donnelly EL, Kirita Y et al (2018) Comparative analysis and refinement of human PSC-derived kidney organoid differentiation with single-cell transcriptomics. Cell Stem Cell 23(869–881):e868. https://doi.org/10.1016/j.stem.2018.10.010
Nishinakamura R (2019) Human kidney organoids: progress and remaining challenges. Nat Rev Nephrol 15:613–624. https://doi.org/10.1038/s41581-019-0176-x
Romero-Guevara R, Ioannides A, **naris C (2020) Kidney organoids as disease models: strengths, weaknesses and perspectives. Front Physiol 11:563981. https://doi.org/10.3389/fphys.2020.563981
Gijzen L, Yousef Yengej FA, Schutgens F, Vormann MK et al (2021) Culture and analysis of kidney tubuloids and perfused tubuloid cells-on-a-chip. Nat Protoc 16:2023–2050. https://doi.org/10.1038/s41596-020-00479-w
Wiraja C, Mori Y, Ichimura T, Hwang J et al (2021) Nephrotoxicity assessment with human kidney tubuloids using spherical nucleic acid-based mRNA nanoflares. Nano Lett 21:5850–5858. https://doi.org/10.1021/acs.nanolett.1c01840
Quiros-Solano WF, Gaio N, Stassen O, Arik YB et al (2018) Microfabricated tuneable and transferable porous PDMS membranes for Organs-on-Chips. Sci Rep 8:13524. https://doi.org/10.1038/s41598-018-31912-6
Clarke GA, Hartse BX, Niaraki Asli AE, Taghavimehr M et al (2021) Advancement of sensor integrated organ-on-chip devices. Sensors (Basel) 21:1367. https://doi.org/10.3390/s21041367
van der Helm MW, Odijk M, Frimat JP, van der Meer AD et al (2016) Direct quantification of transendothelial electrical resistance in organs-on-chips. Biosens Bioelectron 85:924–929. https://doi.org/10.1016/j.bios.2016.06.014
Henry OYF, Villenave R, Cronce MJ, Leineweber WD et al (2017) Organs-on-chips with integrated electrodes for trans-epithelial electrical resistance (TEER) measurements of human epithelial barrier function. Lab Chip 17:2264–2271. https://doi.org/10.1039/c7lc00155j
Booth R, Noh S, Kim H (2014) A multiple-channel, multiple-assay platform for characterization of full-range shear stress effects on vascular endothelial cells. Lab Chip 14:1880–1890. https://doi.org/10.1039/c3lc51304a
Rothbauer M, Ertl P (2020) Emerging biosensor trends in organ-on-a-chip. Adv Biochem Eng Biotechnol. https://doi.org/10.1007/10_2020_129
Toepke MW, Beebe DJ (2006) PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 6:1484–1486. https://doi.org/10.1039/b612140c
Probst C, Schneider S, Loskill P (2018) High-throughput organ-on-a-chip systems: Current status and remaining challenges. Current Opinion in Biomedical Engineering 6:33–41. https://doi.org/10.1016/j.cobme.2018.02.004
Low LA, Mummery C, Berridge BR, Austin CP et al (2021) Organs-on-chips: into the next decade. Nat Rev Drug Discov 20:345–361. https://doi.org/10.1038/s41573-020-0079-3
Tajeddin A, Mustafaoglu N (2021) Design and fabrication of organ-on-chips: promises and challenges. Micromachines (Basel) 12:1443. https://doi.org/10.3390/mi12121443
Mastrangeli M, Millet S, Mummery C, Loskill P et al (2019) Building blocks for a European Organ-on-Chip roadmap. ALTEX 36:481–492. https://doi.org/10.14573/altex.1905221
Valverde MG, Mille LS, Figler KP, Cervantes E et al (2022) Biomimetic models of the glomerulus. Nat Rev Nephrol. https://doi.org/10.1038/s41581-021-00528-x
Petrosyan A, Cravedi P, Villani V, Angeletti A et al (2019) A glomerulus-on-a-chip to recapitulate the human glomerular filtration barrier. Nat Commun 10:3656. https://doi.org/10.1038/s41467-019-11577-z
Iampietro C, Bellucci L, Arcolino FO, Arigoni M et al (2020) Molecular and functional characterization of urine-derived podocytes from patients with Alport syndrome. J Pathol 252:88–100. https://doi.org/10.1002/path.5496
Wang L, Tao T, Su W, Yu H et al (2017) A disease model of diabetic nephropathy in a glomerulus-on-a-chip microdevice. Lab Chip 17:1749–1760. https://doi.org/10.1039/c7lc00134g
Zhou M, Zhang X, Wen X, Wu T et al (2016) Development of a functional glomerulus at the organ level on a chip to mimic hypertensive nephropathy. Sci Rep 6:31771. https://doi.org/10.1038/srep31771
Yang SH, Choi JW, Huh D, Jo HA et al (2017) Roles of fluid shear stress and retinoic acid in the differentiation of primary cultured human podocytes. Exp Cell Res 354:48–56. https://doi.org/10.1016/j.yexcr.2017.03.026
Homan KA, Gupta N, Kroll KT, Kolesky DB et al (2019) Flow-enhanced vascularization and maturation of kidney organoids in vitro. Nat Methods 16:255–262. https://doi.org/10.1038/s41592-019-0325-y
**e R, Korolj A, Liu C, Song X et al (2020) h-FIBER: microfluidic topographical hollow fiber for studies of glomerular filtration barrier. ACS Cent Sci 6:903–912. https://doi.org/10.1021/acscentsci.9b01097
Tuffin J, Burke M, Richardson T, Johnson T et al (2019) A composite hydrogel scaffold permits self-organization and matrix deposition by cocultured human glomerular cells. Adv Healthc Mater 8:e1900698. https://doi.org/10.1002/adhm.201900698
Slater SC, Beachley V, Hayes T, Zhang D et al (2011) An in vitro model of the glomerular capillary wall using electrospun collagen nanofibres in a bioartificial composite basement membrane. PLoS One 6:e20802. https://doi.org/10.1371/journal.pone.0020802
Li Z, Tuffin J, Lei IM, Ruggeri FS et al (2018) Solution fibre spinning technique for the fabrication of tuneable decellularised matrix-laden fibres and fibrous micromembranes. Acta Biomater 78:111–122. https://doi.org/10.1016/j.actbio.2018.08.010
Waters JP, Richards YC, Skepper JN, Southwood M et al (2017) A 3D tri-culture system reveals that activin receptor-like kinase 5 and connective tissue growth factor drive human glomerulosclerosis. J Pathol 243:390–400. https://doi.org/10.1002/path.4960
Wang PC, Takezawa T (2005) Reconstruction of renal glomerular tissue using collagen vitrigel scaffold. J Biosci Bioeng 99:529–540. https://doi.org/10.1263/jbb.99.529
Soo JY, Jansen J, Masereeuw R, Little MH (2018) Advances in predictive in vitro models of drug-induced nephrotoxicity. Nat Rev Nephrol 14:378–393. https://doi.org/10.1038/s41581-018-0003-9
Vormann MK, Gijzen L, Hutter S, Boot L et al (2018) Nephrotoxicity and kidney transport assessment on 3D perfused proximal tubules. AAPS J 20:90. https://doi.org/10.1208/s12248-018-0248-z
Maass C, Sorensen NB, Himmelfarb J, Kelly EJ et al (2019) Translational assessment of drug-induced proximal tubule injury using a kidney microphysiological system. CPT Pharmacometrics Syst Pharmacol 8:316–325. https://doi.org/10.1002/psp4.12400
Jang KJ, Mehr AP, Hamilton GA, McPartlin LA et al (2013) Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr Biol (Camb) 5:1119–1129. https://doi.org/10.1039/c3ib40049b
Frohlich EM, Zhang X, Charest JL (2012) The use of controlled surface topography and flow-induced shear stress to influence renal epithelial cell function. Integr Biol (Camb) 4:75–83. https://doi.org/10.1039/c1ib00096a
Vedula EM, Alonso JL, Arnaout MA, Charest JL (2017) A microfluidic renal proximal tubule with active reabsorptive function. PLoS One 12:e0184330. https://doi.org/10.1371/journal.pone.0184330
Vriend J, Peters JGP, Nieskens TTG, Skovronova R et al (2020) Flow stimulates drug transport in a human kidney proximal tubule-on-a-chip independent of primary cilia. Biochim Biophys Acta Gen Subj 1864:129433. https://doi.org/10.1016/j.bbagen.2019.129433
Naik S, Wood AR, Ongenaert M, Saidiyan P et al (2021) A 3D renal proximal tubule on chip model phenocopies Lowe syndrome and Dent II disease tubulopathy. Int J Mol Sci 22:5361. https://doi.org/10.3390/ijms22105361
Oliveira B, Unwin R, Walsh SB (2019) Inherited proximal tubular disorders and nephrolithiasis. Urolithiasis 47:35–42. https://doi.org/10.1007/s00240-018-01103-z
Weber EJ, Chapron A, Chapron BD, Voellinger JL et al (2016) Development of a microphysiological model of human kidney proximal tubule function. Kidney Int 90:627–637. https://doi.org/10.1016/j.kint.2016.06.011
Lin NYC, Homan KA, Robinson SS, Kolesky DB et al (2019) Renal reabsorption in 3D vascularized proximal tubule models. Proc Natl Acad Sci U S A 116:5399–5404. https://doi.org/10.1073/pnas.1815208116
Trepiccione F, Prosperi F, de la Motte LR, Hubner CA et al (2017) New findings on the pathogenesis of distal renal tubular acidosis. Kidney Dis (Basel) 3:98–105. https://doi.org/10.1159/000478781
Furgeson SB, Linas S (2010) Mechanisms of type I and type II pseudohypoaldosteronism. J Am Soc Nephrol 21:1842–1845. https://doi.org/10.1681/ASN.2010050457
Knoers NV (2006) Gitelman syndrome. Adv Chronic Kidney Dis 13:148–154. https://doi.org/10.1053/j.ackd.2006.01.014
Wang J, Wang C, Xu N, Liu ZF et al (2019) A virus-induced kidney disease model based on organ-on-a-chip: pathogenesis exploration of virus-related renal dysfunctions. Biomaterials 219:119367. https://doi.org/10.1016/j.biomaterials.2019.119367
Jung HJ, Kwon TH (2016) Molecular mechanisms regulating aquaporin-2 in kidney collecting duct. Am J Physiol Renal Physiol 311:F1318–F1328. https://doi.org/10.1152/ajprenal.00485.2016
Jang KJ, Cho HS, Kang DH, Bae WG et al (2011) Fluid-shear-stress-induced translocation of aquaporin-2 and reorganization of actin cytoskeleton in renal tubular epithelial cells. Integr Biol (Camb) 3:134–141. https://doi.org/10.1039/c0ib00018c
Weinberg E, Kaazempur-Mofrad M, Borenstein J (2008) Concept and computational design for a bioartificial nephron-on-a-chip. Int J Artif Organs 31:508–514. https://doi.org/10.1177/039139880803100606
Sakolish Courtney M, Mahler GJ (2017) A novel microfluidic device to model the human proximal tubule and glomerulus. RSC Adv 7:4216–4225. https://doi.org/10.1039/C6RA25641D
Theobald J, Abu El Maaty MA, Kusterer N, Wetterauer B et al (2019) In vitro metabolic activation of vitamin D3 by using a multi-compartment microfluidic liver-kidney organ on chip platform. Sci Rep 9:4616. https://doi.org/10.1038/s41598-019-40851-9
Novak R, Ingram M, Marquez S, Das D et al (2020) Robotic fluidic coupling and interrogation of multiple vascularized organ chips. Nat Biomed Eng 4:407–420. https://doi.org/10.1038/s41551-019-0497-x
Borestrom C, Jonebring A, Guo J, Palmgren H et al (2018) A CRISP(e)R view on kidney organoids allows generation of an induced pluripotent stem cell-derived kidney model for drug discovery. Kidney Int 94:1099–1110. https://doi.org/10.1016/j.kint.2018.05.003
Sakolish CM, Philip B, Mahler GJ (2019) A human proximal tubule-on-a-chip to study renal disease and toxicity. Biomicrofluidics 13:014107. https://doi.org/10.1063/1.5083138
Funding
This study has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska–Curie Grant agreement no. 813839 (JF), H2020 WIDESPREAD-05–2018-TWINNING Remodel (SMM; RM), the IMAGEN project which is co-funded by the PPP Allowance made available by Health ~ Holland, Top Sector Life Sciences & Health, to stimulate public–private partnerships (IMplementation of Advancements in GENetic Kidney Disease, LSHM20009; ESG, MJJ, RM) and Utrecht Institute for Pharmaceutical Sciences (MGV).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
G. Valverde, M., Faria, J., Sendino Garví, E. et al. Organs-on-chip technology: a tool to tackle genetic kidney diseases. Pediatr Nephrol 37, 2985–2996 (2022). https://doi.org/10.1007/s00467-022-05508-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00467-022-05508-2