Abstract
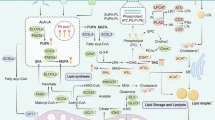
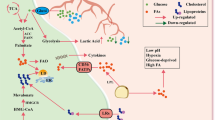
The tumor microenvironment represents the dynamic network consisting of tumor cells, stromal cells, and multiple lineages of immune subsets. It is well recognized that metabolic crosstalk within the tumor microenvironment (TME) greatly shapes both the composition and functionality of the infiltrated immune cells and therefore critically regulate the antitumor immunity. In general, most solid tumors are considered as lipid-enriched environment, which is beneficial for tumor cell growth and immune escape. Here we briefly summarize the effects of accumulated lipids on tumor cells and immune cells. We also discuss the possibility of targeting lipid metabolism within the TME and potential strategies for optimizing cancer treatment.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Egeblad M, et al. Tumors as organs: complex tissues that interface with the entire organism. Dev Cell. 2010;18(6):884–901.
Reina-Campos M, et al. Metabolism shapes the tumor microenvironment. Curr Opin Cell Biol. 2017;48:47–53.
Lyssiotis CA, Kimmelman AC. Metabolic interactions in the tumor microenvironment. Trends Cell Biol. 2017;27(11):863–75.
Binnewies M, et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med. 2018;24(5):541–50.
Ma X, et al. Cholesterol induces CD8(+) T cell exhaustion in the tumor microenvironment. Cell Metab. 2019;30(1):143–15.6.e5.
Anderson KG, et al. Obstacles posed by the tumor microenvironment to T cell activity: a case for synergistic therapies. Cancer Cell. 2017;31(3):311–25.
Cheng C, et al. Lipid metabolism reprogramming and its potential targets in cancer. Cancer Commun (Lond). 2018;38(1):27.
Cartier A, Hla T. Sphingosine 1-phosphate: lipid signaling in pathology and therapy. Science. 2019;366(6463):eaar5551.
Marra AN, et al. Prostaglandin signaling regulates renal multiciliated cell specification and maturation. Proc Natl Acad Sci U S A. 2019;116(17):8409–18.
Eichmann TO, Lass A. DAG tales: the multiple faces of diacylglycerol—stereochemistry, metabolism, and signaling. Cell Mol Life Sci. 2015;72(20):3931–52.
Marat AL, Haucke V. Phosphatidylinositol 3-phosphates-at the interface between cell signalling and membrane traffic. EMBO J. 2016;35(6):561–79.
Michalek RD, et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J Immunol. 2011;186(6):3299–303.
Huang SC, et al. Cell-intrinsic lysosomal lipolysis is essential for alternative activation of macrophages. Nat Immunol. 2014;15(9):846–55.
Silvente-Poirot S, Poirot M. Cancer. Cholesterol and cancer, in the balance. Science. 2014;343(6178):1445–6.
Qian H, et al. Structural basis for catalysis and substrate specificity of human ACAT1. Nature. 2020;581(7808):333–8.
de Gonzalo-Calvo D, et al. Intratumor cholesteryl ester accumulation is associated with human breast cancer proliferation and aggressive potential: a molecular and clinicopathological study. BMC Cancer. 2015;15:460.
Bemlih S, et al. Acyl-coenzyme A: cholesterol acyltransferase inhibitor Avasimibe affect survival and proliferation of glioma tumor cell lines. Cancer Biol Ther. 2010;9(12):1025–32.
Yue S, et al. Cholesteryl ester accumulation induced by PTEN loss and PI3K/AKT activation underlies human prostate cancer aggressiveness. Cell Metab. 2014;19(3):393–406.
**ang W, et al. Monoacylglycerol lipase regulates cannabinoid receptor 2-dependent macrophage activation and cancer progression. Nat Commun. 2018;9(1):2574.
Currie E, et al. Cellular fatty acid metabolism and cancer. Cell Metab. 2013;18(2):153–61.
Rohrig F, Schulze A. The multifaceted roles of fatty acid synthesis in cancer. Nat Rev Cancer. 2016;16(11):732–49.
Santos CR, Schulze A. Lipid metabolism in cancer. FEBS J. 2012;279(15):2610–23.
Shimano H, Sato R. SREBP-regulated lipid metabolism: convergent physiology—divergent pathophysiology. Nat Rev Endocrinol. 2017;13(12):710–30.
Pascual G, et al. Targeting metastasis-initiating cells through the fatty acid receptor CD36. Nature. 2017;541(7635):41–5.
Yang P, et al. Dietary oleic acid-induced CD36 promotes cervical cancer cell growth and metastasis via up-regulation Src/ERK pathway. Cancer Lett. 2018;438:76–85.
Nath A, et al. Elevated free fatty acid uptake via CD36 promotes epithelial-mesenchymal transition in hepatocellular carcinoma. Sci Rep. 2015;5:14752.
Goldstein JL, Brown MS. A century of cholesterol and coronaries: from plaques to genes to statins. Cell. 2015;161(1):161–72.
Lopes-Coelho F, et al. Breast cancer metabolic cross-talk: fibroblasts are hubs and breast cancer cells are gatherers of lipids. Mol Cell Endocrinol. 2018;462(Pt B):93–106.
Santi A, et al. Cancer associated fibroblasts transfer lipids and proteins to cancer cells through cargo vesicles supporting tumor growth. Biochim Biophys Acta. 2015;1853(12):3211–23.
Nieman KM, et al. Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nat Med. 2011;17(11):1498–503.
Iwamoto H, et al. Cancer lipid metabolism confers antiangiogenic drug resistance. Cell Metab. 2018;28(1):104–117.e5.
Samudio I, et al. Pharmacologic inhibition of fatty acid oxidation sensitizes human leukemia cells to apoptosis induction. J Clin Invest. 2010;120(1):142–56.
Liu PP, et al. Elimination of chronic lymphocytic leukemia cells in stromal microenvironment by targeting CPT with an antiangina drug perhexiline. Oncogene. 2016;35(43):5663–73.
Jones SF, Infante JR. Molecular pathways: fatty acid synthase. Clin Cancer Res. 2015;21(24):5434–8.
Ginestier C, et al. Mevalonate metabolism regulates basal breast cancer stem cells and is a potential therapeutic target. Stem Cells. 2012;30(7):1327–37.
Kuzu OF, et al. The role of cholesterol in cancer. Cancer Res. 2016;76(8):2063–70.
Li J, et al. Abrogating cholesterol esterification suppresses growth and metastasis of pancreatic cancer. Oncogene. 2016;35(50):6378–88.
Potente M, Carmeliet P. The link between angiogenesis and endothelial metabolism. Annu Rev Physiol. 2017;79:43–66.
De Bock K, et al. Role of PFKFB3-driven glycolysis in vessel sprouting. Cell. 2013;154(3):651–63.
Cantelmo AR, et al. Inhibition of the glycolytic activator PFKFB3 in endothelium induces tumor vessel normalization, impairs metastasis, and improves chemotherapy. Cancer Cell. 2016;30(6):968–85.
Kalucka J, et al. Quiescent endothelial cells upregulate fatty acid beta-oxidation for Vasculoprotection via redox homeostasis. Cell Metab. 2018;28(6):881–894.e13.
Schoors S, et al. Fatty acid carbon is essential for dNTP synthesis in endothelial cells. Nature. 2015;520(7546):192–7.
Gajewski TF, et al. Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol. 2013;14(10):1014–22.
Vitale I, et al. Macrophages and metabolism in the tumor microenvironment. Cell Metab. 2019;30(1):36–50.
Ngambenjawong C, et al. Progress in tumor-associated macrophage (TAM)-targeted therapeutics. Adv Drug Deliv Rev. 2017;114:206–21.
Su P, et al. Enhanced lipid accumulation and metabolism are required for the differentiation and activation of tumor-associated macrophages. Cancer Res. 2020;80(7):1438–50.
Wu H, et al. Lipid droplet-dependent fatty acid metabolism controls the immune suppressive phenotype of tumor-associated macrophages. EMBO Mol Med. 2019;11(11):e10698.
Zhang D, et al. The Agpat4/LPA axis in colorectal cancer cells regulates antitumor responses via p38/p65 signaling in macrophages. Signal Transduct Target Ther. 2020;5:24.
Wu L, et al. RIPK3 orchestrates fatty acid metabolism in tumor-associated macrophages and hepatocarcinogenesis. Cancer Immunol Res. 2020;8(5):710–21.
Goossens P, et al. Membrane cholesterol efflux drives tumor-associated macrophage reprogramming and tumor progression. Cell Metab. 2019;29(6):1376–1389.e4.
Hossain F, et al. Inhibition of fatty acid oxidation modulates immunosuppressive functions of myeloid-derived suppressor cells and enhances cancer therapies. Cancer Immunol Res. 2015;3(11):1236–47.
Wang H, et al. CD36-mediated metabolic adaptation supports regulatory T cell survival and function in tumors. Nat Immunol. 2020;21(3):298–308.
Joyce JA, Fearon DT. T cell exclusion, immune privilege, and the tumor microenvironment. Science. 2015;348(6230):74–80.
Raud B, et al. Fatty acid metabolism in CD8(+) T cell memory: challenging current concepts. Immunol Rev. 2018;283(1):213–31.
Zhang Y, et al. Enhancing CD8(+) T cell fatty acid catabolism within a metabolically challenging tumor microenvironment increases the efficacy of melanoma immunotherapy. Cancer Cell. 2017;32(3):377–391.e9.
Herber DL, et al. Lipid accumulation and dendritic cell dysfunction in cancer. Nat Med. 2010;16(8):880–6.
Michelet X, et al. Metabolic reprogramming of natural killer cells in obesity limits antitumor responses. Nat Immunol. 2018;19(12):1330–40.
Qin WH, et al. High serum levels of cholesterol increase antitumor functions of nature killer cells and reduce growth of liver tumors in mice. Gastroenterology. 2020;158(6):1713–27.
Yang W, et al. Potentiating the antitumour response of CD8(+) T cells by modulating cholesterol metabolism. Nature. 2016;531(7596):651–5.
Ma X, et al. Cholesterol negatively regulates IL-9-producing CD8(+) T cell differentiation and antitumor activity. J Exp Med. 2018;215(6):1555–69.
Acknowledgments
The authors would like to thank members of Zhang Lab for helpful discussions. L.Z. is supported by Natural Science Foundation of China (NSFC 81971466) and Innovation Fund from the Chinese Academy of Medical Sciences (2016-I2M-1-005).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Ma, K., Zhang, L. (2021). Overview: Lipid Metabolism in the Tumor Microenvironment. In: Li, Y. (eds) Lipid Metabolism in Tumor Immunity. Advances in Experimental Medicine and Biology, vol 1316. Springer, Singapore. https://doi.org/10.1007/978-981-33-6785-2_3
Download citation
DOI: https://doi.org/10.1007/978-981-33-6785-2_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-33-6784-5
Online ISBN: 978-981-33-6785-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)