Abstract
Interspersed repetitions called transposable elements (TEs), commonly referred to as mobile elements, make up a significant portion of the genomes of higher animals. TEs contribute in controlling the expression of genes locally and even far away at the transcriptional and post-transcriptional levels, which is one of their significant functional effects on gene function and genome evolution. There are different mechanisms through which TEs control the expression of genes. First, TEs offer cis-regulatory regions in the genome with their inherent regulatory features for their own expression, making them potential factors for controlling the expression of the host genes. Promoter and enhancer elements contain cis-regulatory sites generated from TE, which function as binding sites for a variety of trans-acting factors. Second, a significant portion of miRNAs and long non-coding RNAs (lncRNAs) have been shown to have TEs that encode for regulatory RNAs, revealing the TE origin of these RNAs. Furthermore, it was shown that TE sequences are essential for these RNAs' regulatory actions, which include binding to the target mRNA. By being a member of cis-regulatory and regulatory RNA sequences, TEs therefore play essential regulatory roles. Additionally, it has been suggested that TE-derived regulatory RNAs and cis-regulatory regions both contribute to the evolutionary novelty of gene regulation. Additionally, these regulatory systems arising from TE frequently have tissue-specific functions. The objective of this review is to discuss TE-mediated gene regulation, with a particular emphasis on the processes, contributions of various TE types, differential roles of various tissue types, based mostly on recent studies on humans.
Similar content being viewed by others
Introduction
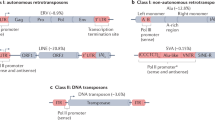
Transposable elements (TEs), also called mobile elements, are DNA fragments that may move about inside a host genome and typically make new copies of themselves while they do so. They are present across all forms of life, accounting for 50% of the mammalian genome [1,2,3]. TEs are present in the genomes of bacteria, plants and mammals, and are divided in two major classes known as Class I retrotransposons and Class II DNA transposons [4], and these two groups vary from one another in terms of the way they transpose. Class II TEs are less common (3.5%) in the human genome and are regarded as DNA fossils because no family of DNA transposons is still active today [5]. The development of genomics and large-scale functional tests has revealed new knowledge on the many functions of TEs [6].
While DNA transposons move commonly by a cut-and-paste mechanism, retrotransposons do so by a copy-and-paste fashion [7]. The transcription of class I retrotransposons results in an intermediate RNA molecule that may be reverse-transcribed into DNA using reverse transcriptase to create a new copy of the retrotransposon in the genome. On the other hand, Class II DNA transposons produce an enzyme called transposase that separates the parental sequence from the genome before mediating its reintegration into another region of the genome [4, 8].
Retrotransposons come in a variety of forms, such as non-LTR retrotransposons and endogenous retroviruses (ERVs), which are distinguished by the presence of long terminal repeats (LTRs). Long nuclear elements (LINEs), short-interspersed elements (SINEs), and SVAs are further classifications for non-LTR retrotransposons [4, 9]. LINEs make up the majority of non-LTR retrotransposons in the human genome, accounting for 20.4% of it, followed by SINEs (13.1%), LTRs (9.1%), and SVAs (0.1%) [10, 11].
The consequences of TE insertions on host gene expression might be beneficial or harmful, like any mutational process. Regardless of their transposition competency, TE regulatory sequences can be co-opted for host regulatory activities. Evidence gives new thoughts on TE mobility and regulatory potential and serves as a vital resource for population history and disease genetics research [12]. Mechanistically, TEs can influence gene expression either transcriptionally [13], post-transcriptionally [14], or at the step of translation [2, 15] through their encoded products which include both proteins and non-coding RNAs (ncRNAs). More complex than originally thought, the mechanisms by which TEs affect host gene-regulatory networks include: the addition of TFBSs, promoters, and enhancers, alteration of 3D chromatin organization, production of regulatory ncRNAs, co-option/exaptation/domestication of TE-derived coding sequences as new transcriptional effector proteins, and collateral consequences of TE silencing mechanisms [2]. The objective of this review is to discuss TE-mediated gene regulation, with a particular emphasis on the mechanisms, contributions of various TE types, and differential roles of various tissue types, based mostly on recent studies on humans.
The roles of transposable elements in the human genome and cell
The evolution of genetic information, as well as DNA duplication, stability, and gene expression, are just a few of the numerous facets of DNA function that TEs may affect. The discovery of TEs' involvement in genome evolution and gene function has altered the previously held belief that TEs are junk, parasitic, colonizing, or selfish DNA [16]. New genes with crucial host functions can be produced as a result of TEs [29]. A recent study has established how lineage-specific TEs can promote evolutionary turnover and divergence of innate immune regulatory networks and reveals a novel function for B2 SINEs as inducible enhancer elements that affect immunity in mice [30].
Transposons could be altered to incorporate a reporter gene that, when randomly inserted into the bacterial chromosome, can fuse to a gene on the chromosome [31]. This kind of transposon library screening for reporter expression under various situations enables the identification of fusions that are appropriate to stress conditions or a particular therapy. A genome-wide picture of the bacterial regulatory network organization may be obtained from the characterization of these fusions [31]. In addition, a study demonstrated that the expression of retrotransposon is clearly related to aging in Drosophila [32].
The negative roles of transposable elements in the human genome
Through processes dependent on and independent of transposition, TEs can lead to genomic/epigenomic instability, which may result in different disease conditions, cell death or the development of cancer [20, 26, 33, 73], histone alterations [74], and mRNA editing [72]. The majority of TEs in somatic cells are silenced by one of these processes, DNA methylation [75] (Fig. 1).
TEs are regulated in both healthy and cancerous cells. Epigenetic changes such as DNA methylation, histone modification, and non-coding RNA (eg cirRNA, miRNA, and lncRNA) inhibit the function of TEs in healthy cells (left panel). During cellular transformation, hypomethylation with increased S-Adenosyl methionine (SAM), various histone modifications (like methylation and acetylation), and oncogenic non-coding RNAs, which inhibit the expression of tumor suppressor genes (TSGs), all contribute to the loss of repressive signals and the uncontrolled production of TEs in cancer cells (right panel). DNA breakdown, mutations, and genomic instability result from all these (arrows indicate the increased activity, cross circle indicates inhibition; ( +) sign indicates increment, (–) sign indicates decrement, and cross sign indicates inhibition) [26]
The interactions between the TEs and a large number of non-coding RNAs are the basis of one well-known germline process [76]. The majority of research on regulatory RNAs has focused on the PIWI-interacting RNAs (piRNAs), which interact with TEs at various levels [77,94]. Numerous of these transcripts are co-opted as regulatory RNAs or chimera transcripts, and L1s produce hundreds of these developmentally regulated and cell-type-specific transcripts. One human-specific transcript expressed only during brain development is LINC01876, an L1-derived lncRNA. L1s are implicated in human-specific developmental processes as a result of decreased size of cerebral organoids and premature differentiation of neural progenitors caused by CRISPRi-silencing of LINC01876. Therefore, it has been demonstrated that L1-derived transcripts offer a previously unrecognized layer of transcriptome complexity that is unique to humans and primates and contributes to the functional diversity of the human brain [94,95,96]. Given that TEs can play a dual and contradictory function in the proper differentiation and development of neuronal mosaicism and in the start of neurological illness, the manifestation of TEs in the brain is symbolic of this "double-edged sword" phenomenon.
These are intriguing illustrations of how TE evolution has included both mechanisms to prevent these invasive sequences from having a negative impact on genome function and systems to allow them to play an active and beneficial part in it. It is becoming apparent that TEs are a crucial component of the genome's regulatory toolbox [97]. RNA translation, alternative splicing, and gene transcription are just a few biological processes for which repetitive sequences have shown promise as regulators [98]. More and more evidence is mounting that TEs can play a crucial role in regulating gene expression in a variety of mechanisms (Fig. 2).
The role of TEs in epigenetic gene expression control
Only recently has the role of TEs in 3D genome architecture been studied. TEs have an impact on 3D chromatin architecture with a direct effect on the folding of chromosomes [2, 104]. It is yet unknown what chemical mechanism causes this correlation between the suppression of TEs and the rise in histone repressive marks [104]. KZFP/KAP1 (Krüppel associated box (KRAB) zinc finger protein/KRAB-associated protein 1) complex plays a crucial role in maintaining heterochromatin, with DNA methylation marks at TEs shielding the loci from TET-mediated demethylation, according to new research in naive murine embryonic stem cells prior to implantation [105]. Ecco et al. [106] discovered that two KRAB/ZFP (Kinc Finger Protein) family members control TE targets by histone-based processes in differentiated tissues, which are not necessarily associated with the DNA methylation state of the loci. Additionally, it has been demonstrated that ZFP92 controls the transcription of particular genes in different tissues by the repression of particular TEs [107].
The work demonstrates that the interactions between the TEs and their KRAB-ZFP controllers affect the expression of neighboring genes. It has been shown that primate-specific ERVs serve as docking sites for the co-repressor protein KAP1 (also known as TRIM28) to produce local heterochromatin in human brain progenitor cells, making this connection even more obvious there [108]. KAP1 binds to the ERVs and represses them, which controls the expression of nearby genes crucial for brain development [108]. The interactions of the transcriptional regulators human silencing hub (HUSH) and microrchidia family CW-type zinc finger 2 (MORC2) with evolutionarily young full-length L1s situated in the transcriptionally permissive euchromatic region, which promotes the deposition of histone H3K9me3, a specific mark for transcriptional silencing, are another example of the regulation of neighboring genes by TEs. A reduction in mRNA expression and potential effects on the RNA polymerase II (POL II) elongation rate might result from this MORC2/HUSH-bound L1 specific impact spreading to nearby genes [109].
Certain kinds of TEs, particularly younger LINEs, have been discovered to affect chromatin accessibility in the livers of several inbred mouse strains, serving as a source of chromatin diversity. This demonstrates the ability of TEs to control tissue-specific genes, which may lead to phenotypic variability among populations [110]. Transposable elements can actively reorganize the chromatin structure to regulate gene expression over a lengthy period of time. About 10% of TE families have been discovered to be enriched in active genomic areas generally and across various organs. While L1 LINEs and ERV LTRs are the most often enriched TE classes in the repressed areas targeted with the H3K9me3 epigenetic mark, SINEs and DNA transposons are the most frequently enriched classes in the active chromatin regions [111].
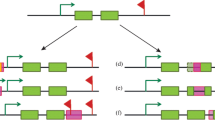
Intriguingly, open euchromatin areas show the strongest epigenetic impact of TEs. By comparing the epigenomes of two D. melanogaster strains, it has been shown, for example, that the enrichment of repressive epigenetic marks around euchromatic TEs is caused by the presence of TEs rather than by the preferential insertion of TEs into genomic regions already enriched with repressive epigenetic marks. This pattern explained why TE-flanking alleles had lower transcript levels and greater histone 3 dimethyl lysine 9 (H3K9me2) enrichment than similar alleles without neighboring TE insertions [112]. Similar to this, the analysis of epigenetic marks in flies with and without Bari-Jheh, a natural transposon that affects the expression of nearby genes, revealed significant differences in histone 3 trimethyl lysine 4 (H3K4me3), H3K9me3, and histone 3 trimethyl lysine 27 (H3K27me3) histone marking in relation to oxidative stress conditions, highlighting that this TE element influences gene expression by affecting the local chromatin state. These illustrations imply that the gene expression of neighboring genes is significantly influenced by the various TE distributions seen in the germlines of various organisms/strains and species [113] (Fig. 3A).
The consequence of TE distribution on the epigenetic control of gene expression due to changes in methylation of histones and DNA across species, tissues of the same organisms, and stimuli or circumstances. A The epigenetic control of a particular gene is altered by the varied distribution of TEs in evolution. TE element controls gene expression by influencing the local chromatin state due to changes in methylation of histones and DNA. B The expression of a particular gene is impacted by the differential redistribution of TEs in various cells and tissues of the same organism during development. C The expression of a certain gene is influenced by the relocalization of TEs sequence in the same cell following a particular stimulus or circumstance
The hypothesis that TEs have been co-opted and that their distributions have co-evolved with the control of gene expression is supported by the intriguing fact that varied TE distributions resulting from somatic transpositions impacting gene expression are also relevant in the same individual [114]. In actuality, TE enrichment differs between tissues, and TEs have binding sites for tissue-specific master transcription regulators [111]. The fact that integration is only permitted in open chromatin areas explains one aspect of TE targeting. The vicinity of neuronal genes is where somatic LINE insertions are abundant in mammalian brains.
The non-random and targeted tissue-specific distribution of TEs might be viewed as a way to genetically fix a landmark, which could result in an epigenetic regulation of neighboring gene expression, if we consider that TEs can be the target of epigenetic marks (Fig. 3B). The discovery that various environmental conditions cause L1 transposition through various basic helix-loop-helix PER-ARNT-SIM (bHLH/PAS) proteins raises the prospect of L1 insertions being targeted differently under various forms of stress [115]. Experimental data suggest that TE insertions are targeted in ways that go beyond the ostensible mechanistic need for accessible chromatin [116].
A temporal and functional hierarchy of transcriptional and epigenomic alterations in response to stress is established in Arabidopsis thaliana by the increased DNA methylation that silences TEs near environmental-induced genes [117]. It is feasible to suggest that, in response to particular stimuli, the mobilization and insertion of TEs may also be regulated in adult tissues and post-mitotic cells to drive the epigenetic regulation of particular genes (Fig. 3C).
Transposable elements in long-range regulation
Different families of TEs have developed many binding sites for transcription factors during the course of evolution, resulting in various transcriptome landscapes [14]. The ENCODE (Encyclopedia of DNA Elements) data comprises roughly 2 million transcription factors binding sites (TFBSs) that coincide with putatively regulation-competent human retrotransposons. For example, these retrotransposons (44% SINEs, 33% LINEs, and 23% LR/ERVs) are situated in a 5-kb gene promoter neighborhood [118]. According to the findings, SINEs are more common than LINE-derived transcription factor binding sites (TFBSs) outside of a 5-kb region close to the transcription start site, but the opposite is true within that region [118].
In addition, while it has long been hypothesized that the repeat sequences that TEs disperse across genomes serve as a source of TFBSs that encourage the emergence of new gene regulatory networks [2], it has only recently become clear that the proteins that TEs encode themselves offer complementary pathways to achieve this result. It was suggested that the process of transposase capture may be a recurring idea in the formation of transcription factors by the discovery that well-characterized transcription factors, such as the paired box (PAX) proteins, feature DNA-binding domains that appear to have arisen from transposases [119].
Additionally, the pathways most significantly influenced by the various retrotransposon distributions have been connected to crucial procedures such as cell stress and immunological responses, ribosome biogenesis, chromatin remodeling, DNA replication, mitotic spindle organization, and cell cycle advancement [118]. The discovery that an evolutionary conserved genomic region called AS3 9, made up of three TEs inserted side by side, serves as a distal enhancer for wnt5a expression during the morphogenesis of the mammalian secondary palate was made by Nishihara et al. [120].
Functional analyses have demonstrated that the AmnSINE1, X6b DNA, and MER117 retrotransposons were co-opted by a retroposition/transposition mechanism during the evolution of mammals. This co-option resulted in the acquisition of a specific Msx1 protein binding site within the X6b DNA sequence, which together with Wnt5a is involved in palatogenesis. According to this study, the great variety of numerous cis-regulatory elements (CREs) may have evolved as a result of the combination of several TEs that were all present in the same DNA segment [120].
TEs’ role as cis-regulatory elements in the genome
It is believed that the majority of CREs newly evolved during primate evolution are directly derived from TEs [121, 122]. Transposable elements frequently contribute to cis-regulatory elements, tissue-specific expression, and alternative promoters in zebrafish, according to epigenomic analysis [123]. In mammalian genomes, transposable elements are a significant source of various cis-regulatory sequences (Fig. 2). According to some studies, 20% of the CREs found in the human genome may have been taken from TEs [124, 125]. By offering binding sites for trans-acting factors, TEs significantly contribute to all cis-regulatory regions (promoters, enhancers, silencers, and insulators) in the human genome [122]. TEs serve as a reservoir for a variety of regulatory functions and are crucial to the evolution of many regulatory components. They either offer substitute enhancers and promoters or change the activity of the current promoters [126, 127].
It has been well established that TEs may adapt to regulatory elements in the human genome and take on non-TE activities [128, 129]. The transcriptional activity of TE-derived sequences in regulatory elements has been empirically verified in several investigations [127, 130, 131]. According to one study, out of the 35,007 promoters, 75% were identified to have TE-derived sequences, with some promoters possessing as many as ten TEs [132]. However, only 6.8% of the TFBSs in promoters were found to be TE-derived, according to the study.
Studies have shown that TFs bind to TEs and that these proteins contain TF-binding sequence motifs [125, 132, 140]. Early in development, young TE families—often LTR elements with embryonic TFBSs in their ancestral sequence—display extremely particular transcriptional patterns [141, 142].
In somatic cells, TEs support cis-regulatory gene networks through the following mechanisms: overlap between the cis-regulatory programs of somatic cells and stem cells, retroviral hijacking of transcription factors expressed in different types of immune cells, or gain of somatic regulatory activity through TE sequence mutations that take place after genomic insertion [139, 143, 144].
Additionally, TE-derived regulatory sites frequently are species/lineage-specific and add innovation and variety to speciation. Future thorough analyses including all regulatory element types across a wide range of species ought to offer more information [127, 145].
However, it has been discovered that mobile element insertion polymorphisms are the most common structural variations in the human genome. Alu elements, L1s, and SVAs are the three groups of retrotransposons that are predominantly in charge of producing human TE polymorphisms [146,189].
Current clinical studies frequently target TEs or benefit from TE biology. Clinical studies using checkpoint inhibitor treatment for immune signaling against renal, ovarian, colorectal, and melanoma malignancies that include TE signaling pathways are currently being conducted [190].
In relation to TEs, both humoral and cell-mediated immunity have been investigated. Several malignancies, including ovarian and melanoma patients as well as teratocarcinoma cell lines, have been linked to anti-ERV-K antibodies [191]. Adaptive immune activity to target TEs as new therapeutic targets was found to be aided in cancer patients by T-cell-mediated and autologous humoral response.
Overexpression of transposon elements in different human diseases is due to demethylation of the TE loci [192]. The TE transcript mechanism, however, is occasionally independent of DNA methylation [193]. This raises the possibility of additional TE regulatory mechanism for non-coding RNAs and histone alterations. Human disorders are significantly influenced by RNA modification [194]. It has been revealed that transposon RNA M(6)A underwent one of its modifications [195]. Transposons may have a role in certain human disease mechanisms, by making use of attractive targets for treatments. In addition to RNA changes, one may look at the uncharacterized DNA modifications of TEs for additional study. One such is m6dA, which is found in the human genome at certain locations, is linked to enhanced transcription activity, and has been implicated in cancer [196, 197]. The link between TE loci and biomarkers raised in disease states would be an intriguing area for further research.
Limitation of the review
This review has a limitation in that it non-specifically addresses the role of transposable elements in the regulation of gene expression. It is more descriptive since it is a narrative review rather than a systematic review and/or meta-analysis, which are supported by statistical analyses and which can objectively answer a particular subject. Therefore, this review presents the authors' own perspectives on a more general topic.
Conclusion and perspective
There are a number of recent discoveries that support the increasingly clear active involvement of TEs in genome function, highlighting their impact on the control of gene expression. In addition to providing ready-to-use TFBSs or undergoing mutations to generate binding motifs for TFs, TEs have inherent regulatory mechanisms for controlling their own expression. Many genes' regulatory elements contain TE sequences, which are involved in both short- and long-range regulation of gene expression. By actively taking part in the production of regulatory RNAs, TEs also contribute to the control of genes. There is still much to learn about the function of transposable elements in gene regulation and their therapeutic potential.
Availability of data and materials
Not Applicable.
Abbreviations
- bHLH:
-
Basic helix-loop-helix
- CD8:
-
Cluster of differentiation 8
- CircRNA:
-
Circular RNAs
- cis-NATs:
-
Cis-natural antisense transcripts
- CREs:
-
Cis-regulatory elements
- CSD:
-
Cortical spreading depression
- H3K9:
-
Histone 3 lysine 9
- HP1a:
-
Heterochromatin protein 1a
- HuR:
-
Human antigen R
- HUSH:
-
Regulators human silencing hub
- KRAB:
-
Krüppel associated box
- LINEs:
-
Long interspersed nuclear elements
- LLERCPs:
-
LINE1-like elements and their reverse complementary pairs
- LTRs:
-
Long terminal repeats
- miRNAs:
-
MicroRNAs
- NSEs:
-
Nonsense-mediated RNA decay switch exons
- piRNAs:
-
P-element Induced WImpy testis (PIWI)-interacting RNAs
- POMC:
-
Proopiomelanocortin
- RBPs:
-
RNA binding proteins
- s/lncRNAs:
-
Short/Long non-coding Ribonucleic acids
- SINEs:
-
Short-interspersed nuclear elements
- SiRNAs:
-
Small-interfering RNAs
- SMD:
-
Staufen-mediated degradation
- SVAs:
-
SINEs Variable Number of Tandem Repeats Alu
- TEs:
-
Transposable elements
- TFBSs:
-
Transcription factor binding sites
- UORFs:
-
Upstream open reading frames
- UTR:
-
Untranslated region
References
Wells JN, Feschotte C. A field guide to eukaryotic transposable elements. Annu Rev Genet. 2020;54:539.
Fueyo R, Judd J, Feschotte C, Wysocka J. Roles of transposable elements in the regulation of mammalian transcription. Nat Rev Mol Cell Biol. 2022;23(7):481–97.
Hoyt SJ, Storer JM, Hartley GA, Grady PG, Gershman A, de Lima LG, Limouse C, Halabian R, Wojenski L, Rodriguez M. From telomere to telomere: the transcriptional and epigenetic state of human repeat elements. Science. 2022;376(6588):eabk3112.
Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell A, Leroy P, Morgante M, Panaud O. A unified classification system for eukaryotic transposable elements. Nat Rev Genet. 2007;8(12):973–82.
Pace JK, Feschotte C. The evolutionary history of human DNA transposons: evidence for intense activity in the primate lineage. Genome Res. 2007;17(4):422–32.
Bourque G, Burns KH, Gehring M, Gorbunova V, Seluanov A, Hammell M, Imbeault M, Izsvák Z, Levin HL, Macfarlan TS. Ten things you should know about transposable elements. Genome Biol. 2018;19(1):1–12.
Lodish H, Berk A, Kaiser CA, Kaiser C, Krieger M, Scott MP, Bretscher A, Ploegh H, Matsudaira P. Molecular cell biology. 8th ed. New York: W.H. Freeman; 2016.
Gifford WD, Pfaff SL, Macfarlan TS. Transposable elements as genetic regulatory substrates in early development. Trends Cell Biol. 2013;23(5):218–26.
Lee S-I, Kim N-S. Transposable elements and genome size variations in plants. Genomics Inf. 2014;12(3):87–97.
Tang W, Mun S, Joshi A, Han K, Liang P. Mobile elements contribute to the uniqueness of human genome with 15,000 human-specific insertions and 14 Mbp sequence increase. DNA Res. 2018;25(5):521–33.
Lancet D, Consortium HGS. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921.
Autio MI, Bin Amin T, Perrin A, Wong JY. Foo RS-Y, Prabhakar S: Transposable elements that have recently been mobile in the human genome. BMC Genomics. 2021;22(1):1–17.
Chuong EB, Elde NC, Feschotte C. Regulatory activities of transposable elements: from conflicts to benefits. Nat Rev Genet. 2017;18(2):71–86.
Drongitis D, Aniello F, Fucci L, Donizetti A. Roles of transposable elements in the different layers of gene expression regulation. Int J Mol Sci. 2019;20(22):5755.
Fort V, Khelifi G, Hussein SM. Long non-coding RNAs and transposable elements: a functional relationship. Biochim Biophys Acta (BBA)-Mol Cell Rese. 2021;1868(1):118837.
Ayarpadikannan S, Kim H-S. The impact of transposable elements in genome evolution and genetic instability and their implications in various diseases. Genomics Inf. 2014;12(3):98–104.
Elisaphenko EA, Kolesnikov NN, Shevchenko AI, Rogozin IB, Nesterova TB, Brockdorff N, Zakian SM. A dual origin of the **st gene from a protein-coding gene and a set of transposable elements. PLoS ONE. 2008;3(6): e2521.
Mi S, Lee X. Li X-p, Veldman GM, Finnerty H, Racie L, LaVallie E, Tang X-Y, Edouard P, Howes S. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature. 2000;403(6771):785–9.
Schneider BK, Sun S, Lee M, Li W, Skvir N, Neretti N, Vijg J, Secombe J. Expression of retrotransposons contributes to aging in Drosophila. Genetics. 2023;224(2):iyad073.
Hsu P-S, Yu S-H, Tsai Y-T, Chang J-Y, Tsai L-K, Ye C-H, Song N-Y, Yau L-C, Lin S-P. More than causing (epi) genomic instability: emerging physiological implications of transposable element modulation. J Biomed Sci. 2021;28(1):1–14.
Villanueva-Cañas JL, Horvath V, Aguilera L, González J. Diverse families of transposable elements affect the transcriptional regulation of stress-response genes in Drosophila melanogaster. Nucleic Acids Res. 2019;47(13):6842–57.
Miao B, Fu S, Lyu C, Gontarz P, Wang T, Zhang B. Tissue-specific usage of transposable element-derived promoters in mouse development. Genome Biol. 2020;21(1):1–25.
Nikolaienko O, Patil S, Eriksen MS, Bramham CR. Arc protein: a flexible hub for synaptic plasticity and cognition. Seminars in cell & developmental biology. 2018;77:33–42.
Kong Y, Rose CM, Cass AA, Williams AG, Darwish M, Lianoglou S, Haverty PM, Tong A-J, Blanchette C, Albert ML. Transposable element expression in tumors is associated with immune infiltration and increased antigenicity. Nat Commun. 2019;10(1):5228.
Bourque G, Burns KH, Gehring M, Gorbunova V, Seluanov A, Hammell M, Imbeault M, Izsvák Z, Levin HL, Macfarlan TS. Ten things you should know about transposable elements. Genome Biol. 2018;19:1–12.
Bhat A, Ghatage T, Bhan S, Lahane GP, Dhar A, Kumar R, Pandita RK, Bhat KM, Ramos KS, Pandita TK. Role of transposable elements in genome stability: implications for health and disease. Int J Mol Sci. 2022;23(14):7802.
Zheng R, Dunlap M, Lyu J, Gonzalez-Figueroa C, Bobkov G, Harvey SE, Chan TW, Quinones-Valdez G, Choudhury M, Vuong A et al. LINE-associated cryptic splicing induces dsRNA-mediated interferon response and tumor immunity. bioRxiv. posted February 24, 2023.
Carrillo D, Reggiardo RE, Lim J, Mantalas G, Peddu V, Kim DH. Transposable element RNA dysregulation in mutant KRAS(G12C) 3D lung cancer spheroids. bioRxiv. posted February 28, 2023.
Grandi N, Tramontano E. Human endogenous retroviruses are ancient acquired elements still sha** innate immune responses. Front Immunol. 2018;9:2039.
Horton I, Kelly CJ, Dziulko A, Simpson DM, Chuong EB. Mouse B2 SINE elements function as IFN-inducible enhancers. Elife. 2023;12:e82617.
Figueroa-Bossi N, Balbontin R, Bossi L. Use of transposable reporters in the analysis of bacterial regulatory networks. Cold Spring Harb Protoc. 2023.
Schneider BK, Sun S, Lee M, Li W, Skvir N, Neretti N, Vijg J, Secombe J. Expression of retrotransposons contributes to aging in Drosophila. Genetics. 2023;224(2):iyad073.
Konkel MK, Batzer MA. A mobile threat to genome stability: The impact of non-LTR retrotransposons upon the human genome. Seminars in cancer biology. 2010;20(4):211–21.
Jang HS, Shah NM, Du AY, Dailey ZZ, Pehrsson EC, Godoy PM, Zhang D, Li D, **ng X, Kim S. Transposable elements drive widespread expression of oncogenes in human cancers. Nat Genet. 2019;51(4):611–7.
Cordaux R, Batzer MA. The impact of retrotransposons on human genome evolution. Nat Rev Genet. 2009;10(10):691–703.
Belancio VP, Hedges DJ, Deininger P. LINE-1 RNA splicing and influences on mammalian gene expression. Nucleic Acids Res. 2006;34(5):1512–21.
Lev-Maor G, Ram O, Kim E, Sela N, Goren A, Levanon EY, Ast G. Intronic Alu s influence alternative splicing. PLoS Genet. 2008;4(9): e1000204.
Shah N, Liang Y. Chimeric transcripts of transposable elements and genes are a source of tumor-specific antigens. Nat Genet. 2023;55(4):538–9.
Sultana T, Zamborlini A, Cristofari G, Lesage P. Integration site selection by retroviruses and transposable elements in eukaryotes. Nat Rev Genet. 2017;18(5):292–308.
Belancio VP, Deininger PL, Roy-Engel AM. LINE dancing in the human genome: transposable elements and disease. Genome Med. 2009;1(10):1–8.
Haig D. Transposable elements: self-seekers of the germline, team-players of the soma. BioEssays. 2016;38(11):1158–66.
Muotri AR, Chu VT, Marchetto MC, Deng W, Moran JV, Gage FH. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature. 2005;435(7044):903–910.
Kazazian HH, Wong C, Youssoufian H, Scott AF, Phillips DG, Antonarakis SE. Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature. 1988;332(6160):164–6.
Bingham PM, Kidwell MG, Rubin GM. The molecular basis of PM hybrid dysgenesis: the role of the P element, a P-strain-specific transposon family. Cell. 1982;29(3):995–1004.
Pereira CM, Stoffel TJ, Callegari-Jacques SM, Hua-Van A, Capy P, Loreto EL. The somatic mobilization of transposable element mariner-Mos1 during the Drosophila lifespan and its biological consequences. Gene. 2018;679:65–72.
Lee E, Iskow R, Yang L, Gokcumen O, Haseley P, Luquette LJ III, Lohr JG, Harris CC, Ding L, Wilson RK. Landscape of somatic retrotransposition in human cancers. Science. 2012;337(6097):967–71.
Tubio JM, Li Y, Ju YS, Martincorena I, Cooke SL, Tojo M, Gundem G, Pipinikas CP, Zamora J, Raine K. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science. 2014;345(6196):1251343.
Rodriguez-Martin B, Alvarez EG, Baez-Ortega A, Zamora J, Supek F, Demeulemeester J, Santamarina M, Ju YS, Temes J, Garcia-Souto D. Pan-cancer analysis of whole genomes identifies driver rearrangements promoted by LINE-1 retrotransposition. Nat Genet. 2020;52(3):306–19.
Tang Z, Steranka JP, Ma S, Grivainis M, Rodić N, Huang CRL, Shih I-M, Wang T-L, Boeke JD, Fenyö D. Human transposon insertion profiling: analysis, visualization and identification of somatic LINE-1 insertions in ovarian cancer. Proc Natl Acad Sci. 2017;114(5):E733–40.
Pitkänen E, Cajuso T, Katainen R, Kaasinen E, Välimäki N, Palin K, Taipale J, Aaltonen LA, Kilpivaara O. Frequent L1 retrotranspositions originating from TTC28 in colorectal cancer. Oncotarget. 2014;5(3):853.
Brégnard C, Guerra J, Déjardin S, Passalacqua F, Benkirane M, Laguette N. Upregulated LINE-1 activity in the Fanconi anemia cancer susceptibility syndrome leads to spontaneous pro-inflammatory cytokine production. EBioMedicine. 2016;8:184–94.
Rodić N, Steranka JP, Makohon-Moore A, Moyer A, Shen P, Sharma R, Kohutek ZA, Huang CR, Ahn D, Mita P. Retrotransposon insertions in the clonal evolution of pancreatic ductal adenocarcinoma. Nat Med. 2015;21(9):1060–4.
Doucet-O’Hare TT, Sharma R, Rodić N, Anders RA, Burns KH, Kazazian HH Jr. Somatically acquired LINE-1 insertions in normal esophagus undergo clonal expansion in esophageal squamous cell carcinoma. Hum Mutat. 2016;37(9):942–54.
Abascal F, Tress ML, Valencia A. Alternative splicing and co-option of transposable elements: the case of TMPO/LAP2α and ZNF451 in mammals. Bioinformatics. 2015;31(14):2257–61.
Bernard A, Boidot R, Végran F. Alternative splicing in cancer and immune cells. Cancers. 2022;14(7):1726.
Kim WR, Park EG, Lee YJ, Bae WH, Lee DH, Kim H-S. Integration of TE induces cancer specific alternative splicing events. Int J Mol Sci. 2022;23(18):10918.
Krasileva KV. The role of transposable elements and DNA damage repair mechanisms in gene duplications and gene fusions in plant genomes. Curr Opin Plant Biol. 2019;48:18–25.
Doyle GA, Crist RC, Karatas ET, Hammond MJ, Ewing AD, Ferraro TN, Hahn C-G, Berrettini WH. Analysis of LINE-1 elements in DNA from postmortem brains of individuals with schizophrenia. Neuropsychopharmacology. 2017;42(13):2602–11.
Shpyleva S, Melnyk S, Pavliv O, Pogribny I, Jill James S. Overexpression of LINE-1 retrotransposons in autism brain. Mol Neurobiol. 2018;55(2):1740–9.
Alesi V, Genovese S, Lepri FR, Catino G, Loddo S, Orlando V, Di Tommaso S, Morgia A, Martucci L, Di Donato M et al. Deep Intronic LINE-1 insertions in NF1: expanding the spectrum of neurofibromatosis type 1-associated rearrangements. Biomolecules. 2023;13(5):725.
DeRosa H, Smith A, Geist L, Cheng A, Hunter RG, Kentner AC. Maternal immune activation alters placental histone-3 lysine-9 tri-methylation, offspring sensorimotor processing, and hypothalamic transposable element expression in a sex-specific manner. Neurobiol Stress. 2023;24: 100538.
Ahn HW, Worman ZF, Lechsinska A, Payer LM, Wang T, Malik N, Li W, Burns KH, Nath A, Levin HL. Retrotransposon insertions associated with risk of neurologic and psychiatric diseases. EMBO Rep. 2023;24(1): e55197.
Jahangir M, Li L, Zhou J-S, Lang B, Wang X-P. L1 Retrotransposons: a potential endogenous regulator for schizophrenia. Front Genet. 2022;13: 878508.
Bundo M, Toyoshima M, Okada Y, Akamatsu W, Ueda J, Nemoto-Miyauchi T, Sunaga F, Toritsuka M, Ikawa D, Kakita A. Increased l1 retrotransposition in the neuronal genome in schizophrenia. Neuron. 2014;81(2):306–13.
De Donno MD, Puricella A, D'Attis S, Specchia V, Bozzetti MP. Expression of transposable elements in the brain of the drosophila melanogaster model for fragile X syndrome. Genes (Basel). 2023;14(5):1060.
Guo C, Jeong H-H, Hsieh Y-C, Klein H-U, Bennett DA, De Jager PL, Liu Z, Shulman JM. Tau activates transposable elements in Alzheimer’s disease. Cell Rep. 2018;23(10):2874–80.
Stetler RA, Leak RK, Gan Y, Li P, Zhang F, Hu X, **g Z, Chen J, Zigmond MJ, Gao Y. Preconditioning provides neuroprotection in models of CNS disease: paradigms and clinical significance. Prog Neurobiol. 2014;114:58–83.
Rana G, Donizetti A, Virelli G, Piscopo M, Viggiano E, De Luca B, Fucci L. Cortical spreading depression differentially affects lysine methylation of H3 histone at neuroprotective genes and retrotransposon sequences. Brain Res. 2012;1467:113–9.
Drongitis D, Rainone S, Piscopo M, Viggiano E, Viggiano A, De Luca B, Fucci L, Donizetti A. Epigenetics and cortical spreading depression: changes of DNA methylation level at retrotransposon sequences. Mol Biol Rep. 2016;43(8):755–60.
Lipszyc A, Szuplewska M, Bartosik D. How do transposable elements activate expression of transcriptionally silent antibiotic resistance genes? Int J Mol Sci. 2022;23(15):8063.
Muñoz-Lopez M, Vilar-Astasio R, Tristan-Ramos P, Lopez-Ruiz C, Garcia-Pérez JL. Study of transposable elements and their genomic impact. Transposons Retrotransposons. 2016;1400:1–19.
Reik W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature. 2007;447(7143):425–32.
Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, De Klein A, Wessels L, De Laat W. Domain organization of human chromosomes revealed by map** of nuclear lamina interactions. Nature. 2008;453(7197):948–51.
Martens JH, O’Sullivan RJ, Braunschweig U, Opravil S, Radolf M, Steinlein P, Jenuwein T. The profile of repeat-associated histone lysine methylation states in the mouse epigenome. EMBO J. 2005;24(4):800–12.
Göke J, Lu X, Chan Y-S, Ng H-H, Ly L-H, Sachs F, Szczerbinska I. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell. 2015;16(2):135–41.
Hadjiargyrou M, Delihas N. The intertwining of transposable elements and non-coding RNAs. Int J Mol Sci. 2013;14(7):13307–28.
Ozata DM, Gainetdinov I, Zoch A, O’Carroll D, Zamore PD. PIWI-interacting RNAs: small RNAs with big functions. Nat Rev Genet. 2019;20(2):89–108.
Anzelon TA, Chowdhury S, Hughes SM, **ao Y, Lander GC, MacRae IJ. Structural basis for piRNA targeting. Nature. 2021;597(7875):285–9.
Yamashiro H, Siomi MC. PIWI-interacting RNA in Drosophila: biogenesis, transposon regulation, and beyond. Chem Rev. 2017;118(8):4404–21.
Le Thomas A, Rogers AK, Webster A, Marinov GK, Liao SE, Perkins EM, Hur JK, Aravin AA, Tóth KF. Piwi induces piRNA-guided transcriptional silencing and establishment of a repressive chromatin state. Genes Dev. 2013;27(4):390–9.
Clark JP, Lau NC. Piwi Proteins and piRNAs step onto the systems biology stage. Syst Biol RNA Binding Proteins. 2014;825:159–97.
Pezic D, Manakov SA, Sachidanandam R, Aravin AA. piRNA pathway targets active LINE1 elements to establish the repressive H3K9me3 mark in germ cells. Genes Dev. 2014;28(13):1410–28.
Lindehell H, Schwartz YB, Larsson J. Methylation of lysine 36 on histone H3 is required to control transposon activities in somatic cells. Life Sci Alliance. 2023;6(8):e202201832.
Iwasaki YW, Murano K, Ishizu H, Shibuya A, Iyoda Y, Siomi MC, Siomi H, Saito K. Piwi modulates chromatin accessibility by regulating multiple factors including histone H1 to repress transposons. Mol Cell. 2016;63(3):408–19.
Malone CD, Brennecke J, Dus M, Stark A, McCombie WR, Sachidanandam R, Hannon GJ. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell. 2009;137(3):522–35.
Lippman Z, Gendrel A-V, Black M, Vaughn MW, Dedhia N, Richard McCombie W, Lavine K, Mittal V, May B, Kasschau KD. Role of transposable elements in heterochromatin and epigenetic control. Nature. 2004;430(6998):471–6.
Yang G, Lee Y-H, Jiang Y, Shi X, Kertbundit S, Hall TC. A two-edged role for the transposable element Kiddo in the rice ubiquitin2 promoter. Plant Cell. 2005;17(5):1559–68.
Baba Y, Yagi T, Sawayama H, Hiyoshi Y, Ishimoto T, Iwatsuki M, Miyamoto Y, Yoshida N, Baba H. Long interspersed element-1 methylation level as a prognostic biomarker in gastrointestinal cancers. Digestion. 2018;97(1):26–30.
Guan Y, Gao H, Leu NA, Vourekas A, Alexiou P, Maragkakis M, Kang Z, Mourelatos Z, Liang G, Wang PJ. The MOV10 RNA helicase is a dosage-dependent host restriction factor for LINE1 retrotransposition in mice. PLoS Genet. 2023;19(5): e1010566.
Gasparotto E, Burattin FV, Di Gioia V, Panepuccia M, Ranzani V, Marasca F, Bodega B. Transposable elements co-option in genome evolution and gene regulation. Int J Mol Sci. 2023;24(3):2610.
Baillie JK, Barnett MW, Upton KR, Gerhardt DJ, Richmond TA, De Sapio F, Brennan PM, Rizzu P, Smith S, Fell M. Somatic retrotransposition alters the genetic landscape of the human brain. Nature. 2011;479(7374):534–7.
Upton KR, Gerhardt DJ, Jesuadian JS, Richardson SR, Sánchez-Luque FJ, Bodea GO, Ewing AD, Salvador-Palomeque C, Van Der Knaap MS, Brennan PM. Ubiquitous L1 mosaicism in hippocampal neurons. Cell. 2015;161(2):228–39.
Coufal NG, Garcia-Perez JL, Peng GE, Yeo GW, Mu Y, Lovci MT, Morell M, O’Shea KS, Moran JV, Gage FH. L1 retrotransposition in human neural progenitor cells. Nature. 2009;460(7259):1127–31.
Garza R, Atacho D, Adami A, Gerdes P, Vinod M, Hsieh P, Karlsson O, Horvath V, Johansson PA, Pandiloski N. L1 retrotransposons drive human neuronal transcriptome complexity and functional diversification. bioRxiv. Posted March 06, 2023..531072.
Muotri AR, Marchetto MC, Coufal NG, Oefner R, Yeo G, Nakashima K, Gage FH. L1 retrotransposition in neurons is modulated by MeCP2. Nature. 2010;468(7322):443–6.
Thomas CA, Paquola AC, Muotri AR. LINE-1 retrotransposition in the nervous system. Annu Rev Cell Dev Biol. 2012;28:555–73.
Slotkin RK, Martienssen R. Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet. 2007;8(4):272–85.
de Souza FS, Franchini LF, Rubinstein M. Exaptation of transposable elements into novel cis-regulatory elements: is the evidence always strong? Mol Biol Evol. 2013;30(6):1239–51.
Choudhary MNK, Quaid K, **ng X, Schmidt H, Wang T. Widespread contribution of transposable elements to the rewiring of mammalian 3D genomes. Nat Commun. 2023;14(1):634.
Kirkland JG, Raab JR, Kamakaka RT. TFIIIC bound DNA elements in nuclear organization and insulation. Biochim Biophys Acta (BBA)-Gene Regul Mech. 2013;1829(3–4):418–424.
Ichiyanagi T, Katoh H, Mori Y, Hirafuku K, Boyboy BA, Kawase M, Ichiyanagi K. B2 SINE copies serve as a transposable boundary of DNA methylation and histone modifications in the mouse. Mol Biol Evol. 2021;38(6):2380–95.
Lunyak VV, Prefontaine GG, Núñez E, Cramer T, Ju B-G, Ohgi KA, Hutt K, Roy R, García-Díaz A, Zhu X. Developmentally regulated activation of a SINE B2 repeat as a domain boundary in organogenesis. Science. 2007;317(5835):248–51.
Hollister JD, Gaut BS. Epigenetic silencing of transposable elements: a trade-off between reduced transposition and deleterious effects on neighboring gene expression. Genome Res. 2009;19(8):1419–28.
Berrens RV, Andrews S, Spensberger D, Santos F, Dean W, Gould P, Sharif J, Olova N, Chandra T, Koseki H. An endosiRNA-based repression mechanism counteracts transposon activation during global DNA demethylation in embryonic stem cells. Cell Stem Cell. 2017;21(5):694–703. e697.
Coluccio A, Ecco G, Duc J, Offner S, Turelli P, Trono D. Individual retrotransposon integrants are differentially controlled by KZFP/KAP1-dependent histone methylation, DNA methylation and TET-mediated hydroxymethylation in naïve embryonic stem cells. Epigenet Chromatin. 2018;11(1):1–18.
Ecco G, Cassano M, Kauzlaric A, Duc J, Coluccio A, Offner S, Imbeault M, Rowe HM, Turelli P, Trono D. Transposable elements and their KRAB-ZFP controllers regulate gene expression in adult tissues. Dev Cell. 2016;36(6):611–23.
Osipovich AB, Dudek KD, Trinh LT, Kim LH, Shrestha S, Cartailler JP, Magnuson MA. ZFP92, a KRAB domain zinc finger protein enriched in pancreatic islets, binds to B1/Alu SINE transposable elements and regulates retroelements and genes. PLoS Genet. 2023;19(5): e1010729.
Brattås PL, Jönsson ME, Fasching L, Wahlestedt JN, Shahsavani M, Falk R, Falk A, Jern P, Parmar M, Jakobsson J. TRIM28 controls a gene regulatory network based on endogenous retroviruses in human neural progenitor cells. Cell Rep. 2017;18(1):1–11.
Liu N, Lee CH, Swigut T, Grow E, Gu B, Bassik MC, Wysocka J. Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators. Nature. 2018;553(7687):228–32.
Du J, Leung A, Trac C, Lee M, Parks BW, Lusis AJ, Natarajan R, Schones DE. Chromatin variation associated with liver metabolism is mediated by transposable elements. Epigenet Chromatin. 2016;9(1):1–16.
Trizzino M, Kapusta A, Brown CD. Transposable elements generate regulatory novelty in a tissue-specific fashion. BMC Genomics. 2018;19(1):1–12.
Lee YCG, Karpen GH. Pervasive epigenetic effects of Drosophila euchromatic transposable elements impact their evolution. Elife. 2017;6: e25762.
Guio L, Vieira C, González J. Stress affects the epigenetic marks added by natural transposable element insertions in Drosophila melanogaster. Sci Rep. 2018;8(1):1–10.
Rebollo R, Romanish MT, Mager DL. Transposable elements: an abundant and natural source of regulatory sequences for host genes. Annu Rev Genet. 2012;46(1):21–42.
Ishizaka Y, Okudaira N, Tamura M, Iijima K, Shimura M, Goto M, Okamura T. Modes of retrotransposition of long interspersed element-1 by environmental factors. Front Microbiol. 2012;3:191.
Mourier T, Nielsen LP, Hansen AJ, Willerslev E. Transposable elements in cancer as a by-product of stress-induced evolvability. Front Genet. 2014;5:156.
Secco D, Wang C, Shou H, Schultz MD, Chiarenza S, Nussaume L, Ecker JR, Whelan J, Lister R. Stress induced gene expression drives transient DNA methylation changes at adjacent repetitive elements. elife. 2015;4:e09343.
Nikitin D, Penzar D, Garazha A, Sorokin M, Tkachev V, Borisov N, Poltorak A, Prassolov V, Buzdin AA. Profiling of human molecular pathways affected by retrotransposons at the level of regulation by transcription factor proteins. Front Immunol. 2018;9:30.
Feschotte C. Transposable elements and the evolution of regulatory networks. Nat Rev Genet. 2008;9(5):397–405.
Nishihara H, Kobayashi N, Kimura-Yoshida C, Yan K, Bormuth O, Ding Q, Nakanishi A, Sasaki T, Hirakawa M, Sumiyama K. Coordinately co-opted multiple transposable elements constitute an enhancer for wnt5a expression in the mammalian secondary palate. PLoS Genet. 2016;12(10): e1006380.
Trizzino M, Park Y, Holsbach-Beltrame M, Aracena K, Mika K, Caliskan M, Perry GH, Lynch VJ, Brown CD. Transposable elements are the primary source of novelty in primate gene regulation. Genome Res. 2017;27(10):1623–33.
Jacques P-É, Jeyakani J, Bourque G. The majority of primate-specific regulatory sequences are derived from transposable elements. PLoS Genet. 2013;9(5): e1003504.
Lee HJ, Hou Y, Maeng JH, Shah NM, Chen Y, Lawson HA, Yang H, Yue F, Wang T. Epigenomic analysis reveals prevalent contribution of transposable elements to cis-regulatory elements, tissue-specific expression, and alternative promoters in zebrafish. Genome Res. 2022; 32(7):1424–36.
Lowe CB, Haussler D. 29 mammalian genomes reveal novel exaptations of mobile elements for likely regulatory functions in the human genome. 2012;7(8):e43128.
Sundaram V, Wysocka J. Transposable elements as a potent source of diverse cis-regulatory sequences in mammalian genomes. Philos Trans R Soc B. 2020;375(1795):20190347.
Conley AB, Piriyapongsa J, Jordan IK. Retroviral promoters in the human genome. Bioinformatics. 2008;24(14):1563–7.
Franchini LF, López-Leal R, Nasif S, Beati P, Gelman DM, Low MJ, de Souza FJ, Rubinstein M. Convergent evolution of two mammalian neuronal enhancers by sequential exaptation of unrelated retroposons. Proc Natl Acad Sci. 2011;108(37):15270–5.
Brosius J, Gould SJ. On “genomenclature”: a comprehensive (and respectful) taxonomy for pseudogenes and other" junk DNA". Proc Natl Acad Sci. 1992;89(22):10706–10.
Gould SJ, Vrba ES. Exaptation—a missing term in the science of form. Paleobiology. 1982;8(1):4–15.
Brini AT, Lee GM, Kinet J-P. Involvement of Alu sequences in the cell-specific regulation of transcription of the gamma chain of Fc and T cell receptors. J Biol Chem. 1993;268(2):1355–61.
Bejerano G, Lowe CB, Ahituv N, King B, Siepel A, Salama SR, Rubin EM, James Kent W, Haussler D. A distal enhancer and an ultraconserved exon are derived from a novel retroposon. Nature. 2006;441(7089):87–90.
Kellner M, Makałowski W. Transposable elements significantly contributed to the core promoters in the human genome. Sci China Life Sci. 2019;62(4):489–97.
Sundaram V, Cheng Y, Ma Z, Li D, **ng X, Edge P, Snyder MP, Wang T. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. 2014;24(12):1963–76.
Zeng L, Pederson SM, Cao D, Qu Z, Hu Z, Adelson DL, Wei C. Genome-wide analysis of the association of transposable elements with gene regulation suggests that alu elements have the largest overall regulatory impact. J Comput Biol. 2018;25(6):551–62.
Samuelson LC, Wiebauer K, Snow C, Meisler MH. Retroviral and pseudogene insertion sites reveal the lineage of human salivary and pancreatic amylase genes from a single gene during primate evolution. Mol Cell Biol. 1990;10(6):2513–20.
Hambor J, Mennone J, Coon M, Hanke J, Kavathas P. Identification and characterization of an Alu-containing, T-cell-specific enhancer located in the last intron of the human CD8 alpha gene. Mol Cell Biol. 1993;13(11):7056–70.
Lamprecht B, Walter K, Kreher S, Kumar R, Hummel M, Lenze D, Köchert K, Bouhlel MA, Richter J, Soler E. Derepression of an endogenous long terminal repeat activates the CSF1R proto-oncogene in human lymphoma. Nat Med. 2010;16(5):571–9.
Pehrsson EC, Choudhary MN, Sundaram V, Wang T. The epigenomic landscape of transposable elements across normal human development and anatomy. Nat Commun. 2019;10(1):1–16.
Chuong EB, Elde NC, Feschotte C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science. 2016;351(6277):1083–7.
Rodriguez-Terrones D, Torres-Padilla M-E. Nimble and ready to mingle: transposon outbursts of early development. Trends Genet. 2018;34(10):806–20.
Flemr M, Malik R, Franke V, Nejepinska J, Sedlacek R, Vlahovicek K, Svoboda P. A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell. 2013;155(4):807–16.
Sakashita A, Maezawa S, Takahashi K, Alavattam KG, Yukawa M, Hu Y-C, Kojima S, Parrish NF, Barski A, Pavlicev M. Endogenous retroviruses drive species-specific germline transcriptomes in mammals. Nat Struct Mol Biol. 2020;27(10):967–77.
Uhlén M, Fagerberg L, Hallström BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson Å, Kampf C, Sjöstedt E, Asplund A. Proteomics. Tissue-based map of the human proteome. Science (New York, NY). 2015;347(6220):1260419–1260419.
Simonti CN, Pavličev M, Capra JA. Transposable element exaptation into regulatory regions is rare, influenced by evolutionary age, and subject to pleiotropic constraints. Mol Biol Evol. 2017;34(11):2856–69.
Rayan NA, Del Rosario RC, Prabhakar S. Massive contribution of transposable elements to mammalian regulatory sequences. Seminars in cell & developmental biology. 2016;57:51–6.
Genomes Project Consortium. A global reference for human genetic variation. Nature. 2015;526(7571):68.
Wang H, **ng J, Grover D, Hedges DJ, Han K, Walker JA, Batzer MA. SVA elements: a hominid-specific retroposon family. J Mol Biol. 2005;354(4):994–1007.
Batzer MA, Deininger PL. A human-specific subfamily of Alu sequences. Genomics. 1991;9(3):481–7.
Brouha B, Schustak J, Badge RM, Lutz-Prigge S, Farley AH, Moran JV, Kazazian HH Jr. Hot L1s account for the bulk of retrotransposition in the human population. Proc Natl Acad Sci. 2003;100(9):5280–5.
Daniel C, Lagergren J, Öhman M. RNA editing of non-coding RNA and its role in gene regulation. Biochimie. 2015;117:22–7.
Gong C, Maquat LE. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature. 2011;470(7333):284–8.
Lucas BA, Lavi E, Shiue L, Cho H, Katzman S, Miyoshi K, Siomi MC, Carmel L, Ares M Jr, Maquat LE. Evidence for convergent evolution of SINE-directed Staufen-mediated mRNA decay. Proc Natl Acad Sci. 2018;115(5):968–73.
Kelley DR, Hendrickson DG, Tenen D, Rinn JL. Transposable elements modulate human RNA abundance and splicing via specific RNA-protein interactions. Genome Biol. 2014;15(12):1–16.
Kim EZ, Wespiser AR, Caffrey DR. The domain structure and distribution of Alu elements in long noncoding RNAs and mRNAs. RNA. 2016;22(2):254–64.
Kralovicova J, Moreno PM, Cross NC, Pêgo AP, Vorechovsky I. Antisense oligonucleotides modulating activation of a nonsense-mediated RNA decay switch exon in the ATM gene. Nucleic Acid Therapeut. 2016;26(6):392–400.
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–9.
Ma L, Bajic VB, Zhang Z. On the classification of long non-coding RNAs. RNA Biol. 2013;10(6):924–33.
Fernandes JC, Acuña SM, Aoki JI, Floeter-Winter LM, Muxel SM. Long non-coding RNAs in the regulation of gene expression: physiology and disease. Non-coding RNA. 2019;5(1):17.
Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol. 2018;19(3):143–57.
Azlan A, Dzaki N, Azzam G. Argonaute: the executor of small RNA function. J Genet Genomics. 2016;43(8):481–94.
Piriyapongsa J, Jordan IK. A family of human microRNA genes from miniature inverted-repeat transposable elements. PLoS ONE. 2007;2(2): e203.
Mustafin RN, Khusnutdinova E. Perspective for studying the relationship of miRNAs with transposable elements. Curr Issues Mol Biol. 2023;45(4):3122–45.
Yuan Z, Sun X, Jiang D, Ding Y, Lu Z, Gong L, Liu H, **e J. Origin and evolution of a placental-specific microRNA family in the human genome. BMC Evol Biol. 2010;10(1):1–12.
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011;25(18):1915–27.
Kapusta A, Kronenberg Z, Lynch VJ, Zhuo X, Ramsay L, Bourque G, Yandell M, Feschotte C. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet. 2013;9(4): e1003470.
Kelley D, Rinn J. Transposable elements reveal a stem cell-specific class of long noncoding RNAs. Genome Biol. 2012;13(11):1–14.
Johnson R, Guigó R. The RIDL hypothesis: transposable elements as functional domains of long noncoding RNAs. RNA. 2014;20(7):959–76.
Kapusta A, Feschotte C. Volatile evolution of long noncoding RNA repertoires: mechanisms and biological implications. Trends Genet. 2014;30(10):439–52.
Fort A, Hashimoto K, Yamada D, Salimullah M, Keya CA, Saxena A, Bonetti A, Voineagu I, Bertin N, Kratz A. Deep transcriptome profiling of mammalian stem cells supports a regulatory role for retrotransposons in pluripotency maintenance. Nat Genet. 2014;46(6):558–66.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141–57.
Zhang X-O, Wang H-B, Zhang Y, Lu X, Chen L-L, Yang L. Complementary sequence-mediated exon circularization. Cell. 2014;159(1):134–47.
Chen L-L, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12(4):381–8.
Chen L, Zhang P, Fan Y, Lu Q, Li Q, Yan J, Muehlbauer GJ, Schnable PS, Dai M, Li L. Circular RNAs mediated by transposons are associated with transcriptomic and phenotypic variation in maize. New Phytol. 2018;217(3):1292–306.
Jung J, Lee S, Cho H-S, Park K, Ryu J-W, Jung M, Kim J, Kim H, Kim D-S. Bioinformatic analysis of regulation of natural antisense transcripts by transposable elements in human mRNA. Genomics. 2019;111(2):159–66.
Shin C, Nam J-W, Farh KK-H, Chiang HR, Shkumatava A, Bartel DP. Expanding the microRNA targeting code: functional sites with centered pairing. Mol Cell. 2010;38(6):789–802.
Spengler RM, Oakley CK, Davidson BL. Functional microRNAs and target sites are created by lineage-specific transposition. Hum Mol Genet. 2014;23(7):1783–93.
Gong C, Maquat LE. “Alu” strious long ncRNAs and their roles in shortening mRNA half-lives. Cell Cycle. 2011;10(12):1882–3.
Piriyapongsa J, Mariño-Ramírez L, Jordan IK. Origin and evolution of human microRNAs from transposable elements. Genetics. 2007;176(2):1323–37.
Qin S, ** P, Zhou X, Chen L, Ma F. The role of transposable elements in the origin and evolution of microRNAs in human. PLoS ONE. 2015;10(6): e0131365.
Mattick JS. Deconstructing the dogma: a new view of the evolution and genetic programming of complex organisms. Ann N Y Acad Sci. 2009;1178(1):29–46.
Kang D, Kim Y-J, Hong K, Han K. TE composition of human long noncoding RNAs and their expression patterns in human tissues. Genes Genomics. 2015;37(1):87–95.
Chishima T, Iwakiri J, Hamada M. Identification of transposable elements contributing to tissue-specific expression of long non-coding RNAs. Genes. 2018;9(1):23.
Kannan S, Chernikova D, Rogozin IB, Poliakov E, Managadze D, Koonin EV, Milanesi L. Transposable element insertions in long intergenic non-coding RNA genes. Front Bioeng Biotech. 2015;3:71.
Kitano S, Kurasawa H, Aizawa Y. Transposable elements shape the human proteome landscape via formation of cis-acting upstream open reading frames. Genes Cells. 2018;23(4):274–84.
Chen G, Wang R, Jiang Y, Dong X, Xu J, Xu Q, Kan Q, Luo Z, Springer NM, Li Q. A novel active transposon creates allelic variation through altered translation rate to influence protein abundance. Nucleic Acids Res. 2023;51(2):595–609.
Shen J, Liu J, **e K, **ng F, **ong F, **ao J, Li X, **ong L. Translational repression by a miniature inverted-repeat transposable element in the 3′ untranslated region. Nat Commun. 2017;8(1):1–10.
Volff JN. Cellular genes derived from Gypsy/Ty3 retrotransposons in mammalian genomes. Ann N Y Acad Sci. 2009;1178(1):233–43.
Pastuzyn ED, Day CE, Kearns RB, Kyrke-Smith M, Taibi AV, McCormick J, Yoder N, Belnap DM, Erlendsson S, Morado DR. The neuronal gene arc encodes a repurposed retrotransposon gag protein that mediates intercellular RNA transfer. Cell. 2018;172(1–2):275–288. e218.
Hudecek M, Izsvák Z, Johnen S, Renner M, Thumann G, Ivics Z. Going non-viral: the slee** beauty transposon system breaks on through to the clinical side. Crit Rev Biochem Mol Biol. 2017;52(4):355–80.
Taylor K, Yau HL, Chakravarthy A, Wang B, Shen SY, Ettayebi I, Ishak CA, Bedard PL, Razak AA, Hansen AR. An open-label, phase II multicohort study of an oral hypomethylating agent CC-486 and durvalumab in advanced solid tumors. J Immunother Cancer. 2020;8(2):e000883.
Hahn S, Ugurel S, Hanschmann K-M, Strobel H, Tondera C, Schadendorf D, Löwer J, Löwer R. Serological response to human endogenous retrovirus K in melanoma patients correlates with survival probability. AIDS Res Hum Retroviruses. 2008;24(5):717–23.
Anwar SL, Wulaningsih W, Lehmann U. Transposable elements in human cancer: causes and consequences of deregulation. Int J Mol Sci. 2017;18(5):974.
Zamudio N, Barau J, Teissandier A, Walter M, Borsos M, Servant N, Bourc’his D. DNA methylation restrains transposons from adopting a chromatin signature permissive for meiotic recombination. Genes Dev. 2015;29(12):1256–70.
Jonkhout N, Tran J, Smith MA, Schonrock N, Mattick JS, Novoa EM. The RNA modification landscape in human disease. RNA. 2017;23(12):1754–69.
Hwang S-Y, Jung H, Mun S, Lee S, Park K, Baek SC, Moon HC, Kim H, Kim B, Choi Y. L1 retrotransposons exploit RNA m6A modification as an evolutionary driving force. Nat Commun. 2021;12(1):1–14.
Pacini CE, Bradshaw CR, Garrett NJ, Koziol MJ. Characteristics and homogeneity of N6-methylation in human genomes. Sci Rep. 2019;9(1):1–12.
Koziol MJ, Bradshaw CR, Allen GE, Costa AS, Frezza C. Identification of methylated deoxyadenosines in genomic DNA by dA6m DNA immunoprecipitation. Bio-Protoc. 2016;6(21):e1990–e1990.
Acknowledgements
Not Applicable.
Funding
No fund has been obtained for this study.
Author information
Authors and Affiliations
Contributions
AG: Conception of research review, literature review, interpretation and write up of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Gebrie, A. Transposable elements as essential elements in the control of gene expression. Mobile DNA 14, 9 (2023). https://doi.org/10.1186/s13100-023-00297-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13100-023-00297-3