Abstract
Background
Metastasis is the major cause of morbidity and mortality in cancer that involves in multiple steps including epithelial–mesenchymal transition (EMT) process. Centrosome is an organelle that functions as the major microtubule organizing center (MTOC), and centrosome abnormalities are commonly correlated with tumor aggressiveness. However, the conclusive mechanisms indicating specific centrosomal proteins participated in tumor progression and metastasis remain largely unknown.
Methods
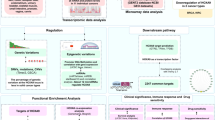
The expression levels of centriolar/centrosomal genes in various types of cancers were first examined by in silico analysis of the data derived from The Cancer Genome Atlas (TCGA), Gene Expression Omnibus (GEO), and European Bioinformatics Institute (EBI) datasets. The expression of STIL (SCL/TAL1-interrupting locus) protein in clinical specimens was further assessed by Immunohistochemistry (IHC) analysis and the oncogenic roles of STIL in tumorigenesis were analyzed using in vitro and in vivo assays, including cell migration, invasion, xenograft tumor formation, and metastasis assays. The transcriptome differences between low- and high-STIL expression cells were analyzed by RNA-seq to uncover candidate genes involved in oncogenic pathways. The quantitative polymerase chain reaction (qPCR) and reporter assays were performed to confirm the results. The chromatin immunoprecipitation (ChIP)-qPCR assay was applied to demonstrate the binding of transcriptional factors to the promoter.
Results
The expression of STIL shows the most significant increase in lung and various other types of cancers, and is highly associated with patients’ survival rate. Depletion of STIL inhibits tumor growth and metastasis. Interestingly, excess STIL activates the EMT pathway, and subsequently enhances cancer cell migration and invasion. Importantly, we reveal an unexpected role of STIL in tumor metastasis. A subset of STIL translocate into nucleus and associate with FOXM1 (Forkhead box protein M1) to promote tumor metastasis and stemness via FOXM1-mediated downstream target genes. Furthermore, we demonstrate that hypoxia-inducible factor 1α (HIF1α) directly binds to the STIL promoter and upregulates STIL expression under hypoxic condition.
Conclusions
Our findings indicate that STIL promotes tumor metastasis through the HIF1α-STIL-FOXM1 axis, and highlight the importance of STIL as a promising therapeutic target for lung cancer treatment.
Similar content being viewed by others
Background
Cancer metastasis is a complex process consisting of many critical steps, such as migration, invasion, adhesion, and metastatic colonization [1]. Gain of migrating and invading abilities are the most important steps during the development of metastasis. Cells experiencing these alterations undergo profound morphological changes, collectively referred to as the epithelial–mesenchymal transition (EMT) process. EMT is considered to be a critical mechanism in regulating cancer invasion and metastasis [1, 2]. It can be triggered by oncogenic activation or microenvironmental stimuli, such as hypoxia. Indeed, the hypoxia of the tumor microenvironment has been shown to be closely associated with metastasis [3]. Under hypoxic conditions, hypoxia-induced factor-1α (HIF-1α) becomes stabilized and up-regulates a number of EMT-related transcription factors (e.g., TWIST and SNAIL) that promote tumor metastasis [4, 5].
Cancer stem cells are a small population of cancer cells holding stemness properties known as cancer stemness (CS), which possess the ability to self-renew and contribute to unlimited cancer proliferation, tumor aggressiveness, drug treatment resistance, and metastasis [6, 7]. Cancer stem cells have been demonstrated to be regulated by several pluripotent transcription factors, such as OCT4, SOX2, and NANOG [8, 4g) in the SLUG promoter. We then conducted the quantitative ChIP-qPCR assay to test whether the FOXM1-STIL complex could bind to the SLUG promoter. Our result showed that both of STIL and FOXM1 bound to the SLUG promoter (Fig. 4g). Notably, the association of both STIL (Fig. 4h) and FOXM1 (Additional file 2: Fig. S6g) with the SLUG promoter was reduced in sh-FOXM1-treated cells. Reciprocally, the reduced SLUG promoter-binding activity of FOXM1 was observed in STIL-depleted CL1-5 cells, suggesting that STIL enhances the transcriptional activity of FOXM1 by increasing its promoter binding affinity (Additional file 2: Fig. S6h). Since FOXM1 directly interacts with the SLUG promoter [103] and STIL does not harbor any known DNA-binding domains, we hypothesize that STIL acts as a transcriptional coactivator of FOXM1 to promote SLUG expression.
Given that the CS core transcription factors (e.g., SOX2, OCT4/POU5F1, and NANOG) were reported to be directly regulated by FOXM1 [106], we tested whether STIL regulates the expression of these CS core genes in a FOXM1-dependent manner. As expected, overexpression of STIL significantly increased the mRNA expression levels of NANOG, SOX2, and POU5F1; however, these STIL-induced upregulations were suppressed upon FOXM1 depletion (Fig. 4i). Further studies demonstrated that STIL depletion affects the expression of these FOXM1-driven genes: overexpression of FOXM1 significantly increased these CS gene promoter-driven luciferase activities (e.g. SLUG, NANOG, SOX2 and POU5F1) in a dose-dependent manner, while knockdown of endogenous STIL reduced FOXM1-induced gene activation (Additional file 2: Fig. S6i). Together, our findings suggest that STIL associates with FOXM1 to enhance the FOXM1-modulated CS.
FOXM1 has also been reported to play an important role in cell cycle regulation that controls the expression of many genes required for G1/S and G2/M transition [107, 108]. To investigate whether STIL might modulate FOXM1-driven cell cycle related genes [109], the gene expression levels of the regulators for G1/S transition, such as CCND1, SKP2 (S-Phase Kinase Associated Protein 2) and CDC25A (Cell Division Cycle 25A), the components of G2/M phase progression including CCNB1 (Cyclin B1), CCNB2 (Cyclin B2), CDK1 (Cyclin Dependent Kinase 1) and PLK1 (Polo Like Kinase 1), and the activators of mitotic entry, such as AURKA (Aurora kinase A), AURKB (Aurora kinase B), CEBPB (CCAAT/enhancer-binding protein beta) and BUBR1/BUB1B (Budding uninhibited by benzimidazoles 1 homolog beta), were examined. Two CL1-5-based FOXM1 knockdown clones and their corresponding control cells overexpressing Flag vector or Flag-STIL were generated (Additional file 2: Fig. S7a). As expected, FOXM1 deletion resulted in the decreased mRNA expression including SKP2, CDC25A, CCNB1, CCNB2, CDK1, PLK1, AURKA, AURKB and BUBR1 (Additional file 2: Fig. S7b). Overexpression of STIL significantly increased the expression of the above genes; however, these STIL-induced upregulations were inhibited upon FOXM1 depletion. In contrast, FOXM1 knockdown or overexpression of STIL did not affect CCND1 and CEBPB mRNA (Additional file 2: Fig. S7c). The reason is not clear. Collectively, our findings suggest that the association of STIL with FOXM1 could upregulate some FOXM1-modulated genes involved in cell cycle regulation, such as SKP2, CDC25A, CCNB1, CCNB2, CDK1, PLK1, AURKA, AURKB, and BUBR1, but not CCND1 and CEBPB.
The interaction between FOXM1 and STIL is required for the STIL-FOXM1 axis-mediated tumorigenic abilities
To validate the importance of the association of STIL with FOXM1 in STIL-mediated tumorigenic functions, we have mapped the FOXM1-interacting region of STIL. Our co-IP results showed that HA-FOXM1 forms complexes with full-length wild type STIL (a.a. 1–1288) and STIL-M (a.a. 420–780), but not STIL-N (a.a. 1–628) or STIL-C (a.a. 780–1288), implying that the region of STIL between a.a. 628 to 780 is responsible for FOXM1-binding (Fig. 5a). Because the coiled-coil domain (a.a 726–748) of STIL [110] was reported to be located within this region, we examined whether the deleted coiled-coil domain mutant of STIL (ΔCC) impairs its FOXM1-binding ability. As shown in Fig. 5b, the full-length STIL and two STIL truncated mutants (a.a. 1–1061 and a.a. 437–1288) containing the coiled-coil domain are coimmunoprecipitated with FOXM1, while the STIL mutant missing the coiled-coli domain (ΔCC) does not. We next examined the tumorigenic effects of the STIL mutant (ΔCC) on migration/invasion and SLUG gene expression. Our results showed that the STIL mutant (ΔCC) impaired its abilities to enhance cellular migration/invasion (Fig. 5c) and SLUG gene activation (Fig. 5d), suggesting that the interaction of FOXM1 with STIL is important for STIL-mediated tumorigenic abilities. Together, our data support a model wherein STIL acts as a coactivator that complexes with FOXM1 to upregulate the FOXM1-mediated downstream genes involved in metastasis, CS, and cell cycle.
The interaction of FOXM1 with STIL is required for STIL-induced tumorigenic abilities. a HEK293T cells were transiently co-transfected HA-FOXM1 with full-length or various truncated Flag-STIL mutants including STIL-N (a.a. 1–628), STIL-M (a.a. 420–780) and STIL-C (a.a. 780–1288) as indicated. Protein complexes were immunoprecipitated using anti-Flag antibody and analyzed by Western blotting using indicated antibodies. b HEK293T cells were transiently co-transfected HA-FOXM1 with the full length or various truncated GFP-STIL mutants (a.a. 1–1061, a.a. 437–1288, and the deleted coiled-coil domain (ΔCC)). Protein complexes were immunoprecipitated using anti-HA antibody and analyzed by Western blotting. c, d NCI-H1299 cells were transiently transfected with GFP-STIL-WT or GFP-STIL mutant (ΔCC), and the expressed proteins were analyzed by Western blotting (c, upper panel). Cell migration and invasion abilities were analyzed by Boyden chamber assay (c, lower panel), while the SLUG promoter activity was measured by reporter assay (d, upper panel) and the SLUG mRNA level (d, lower panel) was measured by qPCR method. Data information: Statistical data in c, d represent the mean ± SD (n = 3 independent experiments). Significance is determined by t-test (NS, not significant; **p < 0.01; ***p < 0.001)
STIL expression is induced by HIF1α under hypoxia
We next explored why STIL is up-regulated in lung cancer. We first examined the STIL DNA copy number using single-nucleotide polymorphism (SNP) array data (Additional file 2: Fig. S8a) and the DNA methylation status of STIL using 450K methylation array data (Additional file 2: Fig. S8b). No significant difference was observed among normal lungs, lung cancer cell lines, and lung cancer specimens in either datasets.
Hypoxia is an important micro-environmental characteristic that activates EMT-TFs and the HIF-mediated pathway during tumor metastasis [111]. The hypoxia pathway was also noted in our GSEA analysis of the STIL-regulated transcriptome (Fig. 3a), and four HIF1α DNA-binding sites ([A/G]CGTG) were identified in the STIL promoter (Fig. 6d). Interestingly, we found that an increase HIF1α level was accompanied by the elevated STIL expression (Fig. 2a) in CL1-0, CL1-3 and CL1-5 cells under the normoxic condition (Additional file 2: Fig. S8c). We thus investigated whether STIL is induced under the hypoxic condition. Figure 6a shows that the protein and mRNA levels of STIL were up-regulated in CL1-0, CL1-5, and NCI-H1229 cells under hypoxia, but that HIF1α depletion dramatically diminished the ability of hypoxia to induce STIL at the protein (Fig. 6a, upper panel) and mRNA (Fig. 6a, lower panel) levels in all three lung cancer lines. HIF1α, which is the major transcription factor in the cellular response to low oxygen, is easily degraded in normoxia. We thus examined whether overexpression of HIF1α (ΔODD), an HIF1α mutant that lacks the oxygen-degradation domain could induce STIL expression under normoxia. As shown in Fig. 6b, we observed increases of STIL at both the protein and mRNA levels in cells overexpressing HIF1α (ΔODD) under the normoxic condition. Consistent with this finding, HIF1α (ΔODD) overexpression could activate STIL promoter-driven luciferase activity in a dose-dependent manner under normoxia (Fig. 6c). Given that a small portion of STIL can be detected in nucleus (Fig. 3e) and STIL is upregulated by HIF1α (Fig. 6a), we further examined the effect of hypoxia on STIL nuclear localization. As shown in Additional file 2: Fig. S8d, the endogenous nuclear STIL was slightly increased under the hypoxic condition compared with that of normoxia (20% versus 11%). The ICC result further showed that HIF1α became stable and accumulated in the nucleus under hypoxia, and led to the increased FOXM1 in the nucleus (Additional file 2: Fig. S8e). Furthermore, hypoxia seems to partially promote the nuclear localization of GFP-STIL (Additional file 2: Fig. S8e), which may reflect with an increase of nuclear GFP-STIL protein detected by Western blotting under hypoxia (36% versus 29%) (Additional file 2: Fig. S8f). Together, our results indicate that HIF1α is an upstream factor that regulates STIL expression.
STIL expression is induced by HIF1α under hypoxia. a STIL protein (upper panel) and mRNA levels (lower panel) were analyzed by Western blotting and qPCR method, respectively, in the indicated cells under normoxic (20% O2) or hypoxic (1% O2) conditions, or in HIF1α-knockdown cells under hypoxia. b STIL protein (upper panel) and mRNA levels (lower panel) were analyzed in cells transiently transfected with the indicated constructs under normoxia. c STIL promoter activity was measured by reporter assay in HEK293T cells transiently transfected with the pGL3-STIL promoter-luciferase plasmid and the indicated constructs under the normoxic condition. d STIL promoter activity was measured by reporter assay in CL1-0 cells transiently transfected with pGL3-STIL promoter-driven luciferase constructs encoding wild-type HIF1α, HIF1α DNA-binding site mutants, and/or HA-HIF1α (ΔODD) under hypoxia. e The binding of HIF1α to the STIL promoter was analyzed by the ChIP-qPCR assay in CL1-5 cells under hypoxia. VEGF promoter was served as the positive control for HIF1α binding, and the region of 6.0 kb upstream of STIL transcriptional start site within STIL promoter was used as the unrelated control. Methylation of histone H3 (H3-K9Me3) on SAT2 gene was used as a positive control and IgG as a negative control for the ChIP-qPCR assay. f Clinical correlations between STIL, HIF1α, and SLUG in 200 lung cancer specimens were analyzed by IHC and the data were assessed using Pearson’s correlation. Scale bar: 50 μm. g Overall survival of 1927 lung cancer patients stratified by STIL and SLUG expression was determined by Kaplan–Meier analysis. The median survival of the four groups is shown. Significance is determined by the log-rank test (p < 0.001). Data information: Statistical data in a–e represent the mean ± SD (n = 3 independent experiments). Significance is determined by t-test (NS, not significant; **p < 0.01; ***p < 0.001)
Since we identified four HIF1α consensus DNA-binding sites (Fig. 6d) in the STIL promoter (site1: nts − 195 to − 199; site2: − 894 to − 898; site3: − 1054 to − 1058; and site4: − 1235 to − 1239), we then generated mutations in these four sites ([A/G]CGTG mutated to [A/G]CTGT) and examined which site is responsible for HIF1α binding. Our results showed that the mutation within nts − 195 to − 199 (site1) dramatically inhibited HIF1α-induced luciferase activation in cells overexpressing HIF1α (ΔODD) under hypoxia (Fig. 6d). A similar effect was also observed in HIF1α (ΔODD)-overexpressing cells under normoxia (Additional file 2: Fig. S8g). To examine whether HIF1α directly binds to the STIL promoter, we performed ChIP-qPCR assay in CL1-5 cells that were pre-treated with hypoxia (to stabilize the HIF1α protein level). Our result showed that HIF1α binds to the STIL promoter under the hypoxic condition (Fig. 6e). Collectively, our findings suggest that HIF1α directly binds to the STIL promoter at the site1 region (nts − 195 to − 199) to drive STIL expression under hypoxia.
We next performed IHC analysis to assess the clinical relevance of STIL in relation to HIF1α and SLUG. Our results showed that the HIF1α intensity was positively correlated with the expression of STIL (Pearson’s co-efficient, R = 0.54) in lung cancer specimens (Fig. 6f). STIL expression was similarly associated with SLUG in the same patient cohort (R = 0.55, Fig. 6f). Intriguingly, the concordant expression of STIL with SLUG displayed an especially high correlation in patients with metastatic lymph nodes (R = 0.76, Additional file 2: Fig. S8h), suggesting that there is a strong association of the HIF1α-STIL-SLUG axis in lung cancer specimens. We thus evaluated the clinical application of the STIL-SLUG axis to predict survival among lung cancer patients. We found that patients with STILhigh and SLUGhigh were associated with poor patient survival (60.0 months) (Fig. 6g), whereas STILlow and SLUGlow patients exhibited prolonged survival (92.6 months). This suggests that the STIL-SLUG axis could be a useful prognostic marker for the survival rate of lung cancer patients.
Discussion
Tumorigenesis is a complex and dynamic process consisting of three major stages: initiation, progression, and metastasis. In the present studies, we identify a novel STIL-mediated mechanism that promotes tumor progression and metastasis. Our collective in vitro and in vivo results on cell proliferation, colony formation, and xenograft tumor assay (Additional file 2: Fig. S2) support a role of STIL in tumor progression. Furthermore, we demonstrated that STIL is associated with FOXM1 (Fig. 4f), and that this association promotes tumor metastasis by activating FOXM1-regulated downstream genes (e.g., SLUG, NANOG, SOX2, POU5F1, SKP2, CDC25A, CCNB1, CCNB2, CDK1, PLK1, AURKA, AURKB and BUBR1) that are involved in the EMT, CS, and cell cycle (Figs. 3b, 4i, and Additional file 2: Fig. S7b). Importantly, we demonstrate that hypoxia is a new factor contributing to STIL upregulation in cancers. HIF1-α directly binds the STIL promoter under hypoxia (Fig. 6e), consequently potentiating hypoxia-induced tumor metastasis. A model showing how STIL contributes to tumor development via the FOXM1-mediated transcriptional activation under hypoxia is shown in Fig. 7.
Schematic diagram illustrates the molecular mechanism how STIL contributes to tumor development via the FOXM1-mediated transcriptional activation of target genes under hypoxia. HIF1α becomes stable under hypoxia in cancer, and translocates to the nucleus to upregulate STIL expression. Furthermore, HIF1α was reported to drive FOXM1 expression [10, 11]. Thus, STIL can associate with FOXM1 to modulate the expression of downstream target genes involved in the EMT process and stemness, and consequently promotes tumor development, metastasis, and poor prognosis
Centrosome abnormalities are commonly observed in human cancers and are correlated with aneuploidy and poor patient prognosis. Previous studies used mouse models to focus on PLK4, which is a key regulator of centrosome duplication [112, 113]. However, the studies in mice with high expression of PLK4 provided contradictory results on the contribution of centrosome amplification to tumor progression. For example, centrosome amplification in neural progenitor cells resulted in microcephaly but did not promote tumorigenesis [114]. Furthermore, Kulukian et al. [115] and Vitre et al. [116] reported that overexpression of PLK4 in the skin epidermis induced an increase in centrosome number but failed to initiate or promote tumorigenesis in skin. In contrast, Sercin et al. [39] showed that PLK4 overexpression accelerates skin tumor formation in mice lacking P53 and Levine et al. [37] demonstrated that supernumerary centrosomes are sufficient to drive tumorigenesis in multiple tissues of mice. Thus, the issue of whether direct associations exist between centrosome abnormalities and cancers remains unclear.
In this study, we examined the T (tumor)/N (non-malignant) ratio of 14 centriolar/centrosomal genes in lung cancer patients and their paired adjacent non-malignant lung tissues. STIL showed the highest T/N ratio (3.5) among the studied genes (Table 1); this was found in patients with lung cancer, and was even higher than the T/N ratio (1.5) of PLK4 in these patients. Further analysis also showed that the STIL mRNA level was significantly increased in many other types of cancers (Additional file 1: Table S1a) and its high expression is associated with poor prognosis in patients with many cancer types (Additional file 1: Table S1b). STIL was previously reported to be present in the cytosol, specifically in the centriole [43], and act as a master regulator of PLK4 in initiating centriole duplication [45,46,47,48,49]. Here, we reveal an unexpected novel role of STIL in the nucleus. In addition to centriolar STIL, a subset of STIL translocate into the nucleus and function as the coactivator to enhance the downstream FOXM1-drievn genes via the association between STIL and FOXM1, and consequently contributes to the metastasis. This finding may explain the elevated STIL expression in lung cancer patients with metastatic lymph nodes or brain metastases.
A remaining open question is: Do the extra centrosomes induced by excess STIL promote tumor initiation and drive spontaneous tumorigenesis? The answer is not yet clear. Using si-SASS6 to block excess STIL-induced centriole amplification, we found that the migration and invasion abilities were significantly reduced in the si-SASS6-treated cells harboring excess STIL (Additional file 2: Fig. S4a). However, the si-SASS6 treatment does not completely block the migration/invasion abilities of STIL overexpressing cells (Additional file 2: Fig. S4a). These findings suggest that in addition to the excess STIL-mediated cancer cell migration and invasion, the possibility of supernumerary centrosome aberration-triggered tumorigenesis (e.g. aneuploidy and/or tissue architecture disruption) [20] can’t be ruled out. Future experiments are needed to clarify this discrepancy.
Finally, it has been proposed that the small increases in centrosome number induced by a low-to-moderate level of PLK4 are permissive for tumor development, whereas high levels of PLK4 trigger larger number of centrosomes and are likely to be harmful for long-term cell survival [37]. Since STIL is a master regulator of PLK4, we speculate that a low-to-moderate level of STIL could promote tumor initiation, as seen for PLK4, via a yet-unknown mechanism. Future experiments by generating a transgenic mouse line with low to moderate expression level of STIL could be a way to test the role of STIL in the initial stage of tumorigenesis.
In summary, we herein show that STIL is significantly up-regulated in lung and many other types of cancers, and that its expression level is highly correlated with patient survival, implicating its potential application in cancer detection and as a prognostic marker. Importantly, we demonstrate that STIL plays a versatile role in multistage tumorigenesis through the HIF1α-STIL-FOXM1 axis, and therefore may serve as a promising target for cancer therapy.
Conclusion
Our findings indicate that the centriolar protein STIL functions not only as a key regulator in centriole duplication but also as a transcriptional coactivator that regulates EMT and stemness to promote tumor metastasis. Our findings show that a subset of STIL enter the nucleus, which interact with FOXM1 to activate its downstream target genes in metastasis. Furthermore, we provide evidence to show that HIF1α directly binds to STIL promoter and drives STIL gene expression under hypoxia. Thus, STIL can serve as a potential diagnostic marker for early lung cancer detection, and a promising therapeutic target for lung cancer treatment.
Availability of data and materials
All data relevant to the study are included in the article and in additional files. The reagents used in this publication are available from the corresponding author on reasonable request.
Change history
09 April 2024
A Correction to this paper has been published: https://doi.org/10.1186/s12929-024-01021-w
Abbreviations
- ADC:
-
Adenocarcinoma
- ARG2:
-
Arginase 2
- AUC:
-
Area under the curve
- AURKA:
-
Aurora kinase A
- AURKB:
-
Aurora kinase B
- BUBR1/BUB1B:
-
Budding uninhibited by benzimidazoles 1 homolog beta
- CCK-8:
-
Cell counting kit-8
- CCNB1:
-
Cyclin B1
- CCNB2:
-
Cyclin B2
- CCND1:
-
Cyclin D1
- CDC25A:
-
Cell Division Cycle 25A
- CDK1:
-
Cyclin dependent kinase 1
- CEBPB:
-
CCAAT/enhancer-binding protein beta
- CEP63:
-
Centrosomal protein 63
- CEP120:
-
Centrosomal protein 120
- CEP152:
-
Centrosomal protein 152
- CEP135:
-
Centrosomal protein 135
- CEP295:
-
Centrosomal protein 295
- CETN1:
-
Centrin 1
- ChIP:
-
Chromatin immunoprecipitation
- CPAP:
-
Centrosomal P4.1-associated protein
- CS:
-
Cancer stemness
- DMEM:
-
Dulbecco’s Modified Eagle’s Medium
- Dox:
-
Doxycycline
- EMT:
-
Epithelial–mesenchymal transition
- FBS:
-
Fetal bovine serum
- FOXM1:
-
Forkhead box protein M1
- GEO:
-
Gene Expression Omnibus
- GSEA:
-
Gene set enrichment analysis
- HIF1α:
-
Hypoxia-inducible factor 1α
- ICC:
-
Immunocytochemistry
- IHC:
-
Immunohistochemistry
- IF:
-
Immunofluorescent
- IVIS:
-
In vivo imaging system
- MCPH:
-
Autosomal recessive primary microcephaly
- MT:
-
Microtubule
- MTOC:
-
Microtubule organizing center
- PCM:
-
Pericentriolar material
- PLK1:
-
Polo like kinase 1
- PLK4:
-
Polo-like kinase 4
- POC1B:
-
Proteome of centriole protein 1 beta
- POC5:
-
Proteome of centriole protein 5
- qPCR:
-
Quantitative polymerase chain reaction
- R:
-
Pearson correlation coefficient
- ROC:
-
Receptor operating characteristics
- RTTN:
-
Rotatin
- SAS6:
-
Spindle assembly abnormal protein 6
- SAT2:
-
Spermidine/Spermine N1-acetyltransferase family member 2
- SCC:
-
Squamous cell carcinoma
- SD:
-
Standard derivation
- SKP2:
-
S-phase kinase associated protein 2
- SNP:
-
Single-nucleotide polymorphism
- SPICE1:
-
Spindle and centriole-associated protein 1
- STIL:
-
SCL/TAL1-interrupting locus
- TCGA:
-
The Cancer Genome Atlas
- VEGF:
-
Vascular endothelial growth factor
References
Lambert AW, Pattabiraman DR, Weinberg RA. Emerging biological principles of metastasis. Cell. 2017;168(4):670–91.
Tiwari N, Gheldof A, Tatari M, Christofori G. EMT as the ultimate survival mechanism of cancer cells. Semin Cancer Biol. 2012;22(3):194–207.
Wilson WR, Hay MP. Targeting hypoxia in cancer therapy. Nat Rev Cancer. 2011;11(6):393–410.
Casazza A, Di Conza G, Wenes M, Finisguerra V, Deschoemaeker S, Mazzone M. Tumor stroma: a complexity dictated by the hypoxic tumor microenvironment. Oncogene. 2014;33(14):1743–54.
Yang MH, Wu MZ, Chiou SH, Chen PM, Chang SY, Liu CJ, et al. Direct regulation of TWIST by HIF-1alpha promotes metastasis. Nat Cell Biol. 2008;10(3):295–305.
Mackillop WJ, Ciampi A, Till JE, Buick RN. A stem cell model of human tumor growth: implications for tumor cell clonogenic assays. J Natl Cancer Inst. 1983;70(1):9–16.
Clevers H. The cancer stem cell: premises, promises and challenges. Nat Med. 2011;17(3):313–9.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4):663–76.
Maherali N, Sridharan R, **e W, Utikal J, Eminli S, Arnold K, et al. Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell. 2007;1(1):55–70.
Tang C, Liu T, Wang K, Wang X, Xu S, He D, et al. Transcriptional regulation of FoxM1 by HIF1alpha mediates hypoxiainduced EMT in prostate cancer. Oncol Rep. 2019;42(4):1307–18.
**a LM, Huang WJ, Wang B, Liu M, Zhang Q, Yan W, et al. Transcriptional up-regulation of FoxM1 in response to hypoxia is mediated by HIF-1. J Cell Biochem. 2009;106(2):247–56.
Sun H, Teng M, Liu J, ** D, Wu J, Yan D, et al. FOXM1 expression predicts the prognosis in hepatocellular carcinoma patients after orthotopic liver transplantation combined with the Milan criteria. Cancer Lett. 2011;306(2):214–22.
Lu XF, Zeng D, Liang WQ, Chen CF, Sun SM, Lin HY. FoxM1 is a promising candidate target in the treatment of breast cancer. Oncotarget. 2018;9(1):842–52.
Kim IM, Ackerson T, Ramakrishna S, Tretiakova M, Wang IC, Kalin TV, et al. The Forkhead Box m1 transcription factor stimulates the proliferation of tumor cells during development of lung cancer. Cancer Res. 2006;66(4):2153–61.
Wierstra I, Alves J. FOXM1, a typical proliferation-associated transcription factor. Biol Chem. 2007;388(12):1257–74.
Halasi M, Gartel AL. FOX(M1) news—it is cancer. Mol Cancer Ther. 2013;12(3):245–54.
Alvarez-Fernandez M, Medema RH. Novel functions of FoxM1: from molecular mechanisms to cancer therapy. Front Oncol. 2013;3:30.
Raychaudhuri P, Park HJ. FoxM1: a master regulator of tumor metastasis. Cancer Res. 2011;71(13):4329–33.
Banterle N, Gonczy P. Centriole biogenesis: from identifying the characters to understanding the plot. Annu Rev Cell Dev Biol. 2017;33:23–49.
LoMastro GM, Holland AJ. The emerging link between centrosome aberrations and metastasis. Dev Cell. 2019;49(3):325–31.
Thornton GK, Woods CG. primary microcephaly: do all roads lead to Rome? Trends Genet. 2009;25(11):501–10.
Nigg EA, Holland AJ. Once and only once: mechanisms of centriole duplication and their deregulation in disease. Nat Rev Mol Cell Biol. 2018;19(5):297–312.
Brito DA, Gouveia SM, Bettencourt-Dias M. Deconstructing the centriole: structure and number control. Curr Opin Cell Biol. 2012;24(1):4–13.
Gonczy P. Towards a molecular architecture of centriole assembly. Nat Rev Mol Cell Biol. 2012;13(7):425–35.
Breslow DK, Holland AJ. Mechanism and regulation of centriole and cilium biogenesis. Annu Rev Biochem. 2019;88:691–724.
Cizmecioglu O, Arnold M, Bahtz R, Settele F, Ehret L, Haselmann-Weiss U, et al. Cep152 acts as a scaffold for recruitment of Plk4 and CPAP to the centrosome. J Cell Biol. 2010;191(4):731–9.
Hatch EM, Kulukian A, Holland AJ, Cleveland DW, Stearns T. Cep152 interacts with Plk4 and is required for centriole duplication. J Cell Biol. 2010;191(4):721–9.
Kim TS, Park JE, Shukla A, Choi S, Murugan RN, Lee JH, et al. Hierarchical recruitment of Plk4 and regulation of centriole biogenesis by two centrosomal scaffolds, Cep192 and Cep152. Proc Natl Acad Sci USA. 2013;110(50):E4849–57.
Sonnen KF, Gabryjonczyk AM, Anselm E, Stierhof YD, Nigg EA. Human Cep192 and Cep152 cooperate in Plk4 recruitment and centriole duplication. J Cell Sci. 2013;126(Pt 14):3223–33.
Lin YC, Chang CW, Hsu WB, Tang CJ, Lin YN, Chou EJ, et al. Human microcephaly protein CEP135 binds to hSAS-6 and CPAP, and is required for centriole assembly. EMBO J. 2013;32(8):1141–54.
Comartin D, Gupta GD, Fussner E, Coyaud E, Hasegan M, Archinti M, et al. CEP120 and SPICE1 cooperate with CPAP in centriole elongation. Curr Biol. 2013;23(14):1360–6.
Lin YN, Wu CT, Lin YC, Hsu WB, Tang CJ, Chang CW, et al. CEP120 interacts with CPAP and positively regulates centriole elongation. J Cell Biol. 2013;202(2):211–9.
Chen HY, Wu CT, Tang CC, Lin YN, Wang WJ, Tang TK. Human microcephaly protein RTTN interacts with STIL and is required to build full-length centrioles. Nat Commun. 2017;8(1):247.
Chang CW, Hsu WB, Tsai JJ, Tang CJ, Tang TK. CEP295 interacts with microtubules and is required for centriole elongation. J Cell Sci. 2016;129(13):2501–13.
Jayaraman D, Bae BI, Walsh CA. The genetics of primary microcephaly. Annu Rev Genomics Hum Genet. 2018;19:177–200.
Godinho SA, Pellman D. Causes and consequences of centrosome abnormalities in cancer. Philos Trans R Soc Lond B Biol Sci. 2014;369(1650):20130467.
Levine MS, Bakker B, Boeckx B, Moyett J, Lu J, Vitre B, et al. Centrosome amplification is sufficient to promote spontaneous tumorigenesis in mammals. Dev Cell. 2017;40(3):313-22.e5.
Coelho PA, Bury L, Shahbazi MN, Liakath-Ali K, Tate PH, Wormald S, et al. Over-expression of Plk4 induces centrosome amplification, loss of primary cilia and associated tissue hyperplasia in the mouse. Open Biol. 2015;5(12):150209.
Sercin O, Larsimont JC, Karambelas AE, Marthiens V, Moers V, Boeckx B, et al. Transient PLK4 overexpression accelerates tumorigenesis in p53-deficient epidermis. Nat Cell Biol. 2016;18(1):100–10.
Aplan PD, Lombardi DP, Kirsch IR. Structural characterization of SIL, a gene frequently disrupted in T-cell acute lymphoblastic leukemia. Mol Cell Biol. 1991;11(11):5462–9.
Castiel A, Danieli MM, David A, Moshkovitz S, Aplan PD, Kirsch IR, et al. The Stil protein regulates centrosome integrity and mitosis through suppression of Chfr. J Cell Sci. 2011;124(Pt 4):532–9.
Arquint C, Sonnen KF, Stierhof YD, Nigg EA. Cell-cycle-regulated expression of STIL controls centriole number in human cells. J Cell Sci. 2012;125(Pt 5):1342–52.
Tang CJ, Lin SY, Hsu WB, Lin YN, Wu CT, Lin YC, et al. The human microcephaly protein STIL interacts with CPAP and is required for procentriole formation. EMBO J. 2011;30(23):4790–804.
Vulprecht J, David A, Tibelius A, Castiel A, Konotop G, Liu F, et al. STIL is required for centriole duplication in human cells. J Cell Sci. 2012;125(Pt 5):1353–62.
Arquint C, Gabryjonczyk AM, Imseng S, Bohm R, Sauer E, Hiller S, et al. STIL binding to Polo-box 3 of PLK4 regulates centriole duplication. Elife. 2015;4:e07888.
Dzhindzhev NS, Tzolovsky G, Lipinszki Z, Schneider S, Lattao R, Fu J, et al. Plk4 phosphorylates Ana2 to trigger Sas6 recruitment and procentriole formation. Curr Biol. 2014;24(21):2526–32.
Klebba JE, Buster DW, McLamarrah TA, Rusan NM, Rogers GC. Autoinhibition and relief mechanism for Polo-like kinase 4. Proc Natl Acad Sci USA. 2015;112(7):E657–66.
Moyer TC, Clutario KM, Lambrus BG, Daggubati V, Holland AJ. Binding of STIL to Plk4 activates kinase activity to promote centriole assembly. J Cell Biol. 2015;209(6):863–78.
Ohta M, Ashikawa T, Nozaki Y, Kozuka-Hata H, Goto H, Inagaki M, et al. Direct interaction of Plk4 with STIL ensures formation of a single procentriole per parental centriole. Nat Commun. 2014;5:5267.
Erez A, Perelman M, Hewitt SM, Cojacaru G, Goldberg I, Shahar I, et al. Sil overexpression in lung cancer characterizes tumors with increased mitotic activity. Oncogene. 2004;23(31):5371–7.
Wang J, Zhang Y, Dou Z, Jiang H, Wang Y, Gao X, et al. Knockdown of STIL suppresses the progression of gastric cancer by down-regulating the IGF-1/PI3K/AKT pathway. J Cell Mol Med. 2019;23(8):5566–75.
Lu TP, Lai LC, Tsai MH, Chen PC, Hsu CP, Lee JM, et al. Integrated analyses of copy number variations and gene expression in lung adenocarcinoma. PLoS ONE. 2011;6(9):e24829.
Lai LC, Tsai MH, Chen PC, Chen LH, Hsiao JH, Chen SK, et al. SNP rs10248565 in HDAC9 as a novel genomic aberration biomarker of lung adenocarcinoma in non-smoking women. J Biomed Sci. 2014;21:24.
Lu TP, Hsiao CK, Lai LC, Tsai MH, Hsu CP, Lee JM, et al. Identification of regulatory SNPs associated with genetic modifications in lung adenocarcinoma. BMC Res Notes. 2015;8:92.
Wei TY, Juan CC, Hisa JY, Su LJ, Lee YC, Chou HY, et al. Protein arginine methyltransferase 5 is a potential oncoprotein that upregulates G1 cyclins/cyclin-dependent kinases and the phosphoinositide 3-kinase/AKT signaling cascade. Cancer Sci. 2012;103(9):1640–50.
Wei TY, Hsia JY, Chiu SC, Su LJ, Juan CC, Lee YC, et al. Methylosome protein 50 promotes androgen- and estrogen-independent tumorigenesis. Cell Signal. 2014;26(12):2940–50.
Sanchez-Palencia A, Gomez-Morales M, Gomez-Capilla JA, Pedraza V, Boyero L, Rosell R, et al. Gene expression profiling reveals novel biomarkers in nonsmall cell lung cancer. Int J Cancer. 2011;129(2):355–64.
Suraokar MB, Nunez MI, Diao L, Chow CW, Kim D, Behrens C, et al. Expression profiling stratifies mesothelioma tumors and signifies deregulation of spindle checkpoint pathway and microtubule network with therapeutic implications. Ann Oncol. 2014;25(6):1184–92.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483(7391):603–7.
Tarca AL, Lauria M, Unger M, Bilal E, Boue S, Kumar Dey K, et al. Strengths and limitations of microarray-based phenotype prediction: lessons learned from the IMPROVER Diagnostic Signature Challenge. Bioinformatics. 2013;29(22):2892–9.
Rousseaux S, Debernardi A, Jacquiau B, Vitte AL, Vesin A, Nagy-Mignotte H, et al. Ectopic activation of germline and placental genes identifies aggressive metastasis-prone lung cancers. Sci Transl Med. 2013;5(186):186ra66.
Director’s Challenge Consortium for the Molecular Classification of Lung A, Shedden K, Taylor JM, Enkemann SA, Tsao MS, Yeatman TJ, et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med. 2008;14(8):822–7.
Botling J, Edlund K, Lohr M, Hellwig B, Holmberg L, Lambe M, et al. Biomarker discovery in non-small cell lung cancer: integrating gene expression profiling, meta-analysis, and tissue microarray validation. Clin Cancer Res. 2013;19(1):194–204.
Jabs V, Edlund K, Konig H, Grinberg M, Madjar K, Rahnenfuhrer J, et al. Integrative analysis of genome-wide gene copy number changes and gene expression in non-small cell lung cancer. PLoS ONE. 2017;12(11):e0187246.
Lohr M, Hellwig B, Edlund K, Mattsson JS, Botling J, Schmidt M, et al. Identification of sample annotation errors in gene expression datasets. Arch Toxicol. 2015;89(12):2265–72.
Goldmann T, Marwitz S, Nitschkowski D, Krupar R, Backman M, Elfving H, et al. PD-L1 amplification is associated with an immune cell rich phenotype in squamous cell cancer of the lung. Cancer Immunol Immunother. 2021;70(9):2577–87.
Der SD, Sykes J, Pintilie M, Zhu CQ, Strumpf D, Liu N, et al. Validation of a histology-independent prognostic gene signature for early-stage, non-small-cell lung cancer including stage IA patients. J Thorac Oncol. 2014;9(1):59–64.
Borczuk AC, Sole M, Lu P, Chen J, Wilgus ML, Friedman RA, et al. Progression of human bronchioloalveolar carcinoma to invasive adenocarcinoma is modeled in a transgenic mouse model of K-ras-induced lung cancer by loss of the TGF-beta type II receptor. Cancer Res. 2011;71(21):6665–75.
Zhu CQ, Ding K, Strumpf D, Weir BA, Meyerson M, Pennell N, et al. Prognostic and predictive gene signature for adjuvant chemotherapy in resected non-small-cell lung cancer. J Clin Oncol. 2010;28(29):4417–24.
Luke F, Blazquez R, Yamaci RF, Lu X, Pregler B, Hannus S, et al. Isolated metastasis of an EGFR-L858R-mutated NSCLC of the meninges: the potential impact of CXCL12/CXCR4 axis in EGFRmut NSCLC in diagnosis, follow-up and treatment. Oncotarget. 2018;9(27):18844–57.
Lanczky A, Gyorffy B. Web-based survival analysis tool tailored for medical research (KMplot): development and implementation. J Med Internet Res. 2021;23(7):e27633.
Li B, Shin H, Gulbekyan G, Pustovalova O, Nikolsky Y, Hope A, et al. Development of a drug-response modeling framework to identify cell line derived translational biomarkers that can predict treatment outcome to erlotinib or sorafenib. PLoS ONE. 2015;10(6):e0130700.
Avila-Moreno F, Armas-Lopez L, Alvarez-Moran AM, Lopez-Bujanda Z, Ortiz-Quintero B, Hidalgo-Miranda A, et al. Overexpression of MEOX2 and TWIST1 is associated with H3K27me3 levels and determines lung cancer chemoresistance and prognosis. PLoS ONE. 2014;9(12):e114104.
Lee W, Jiang Z, Liu J, Haverty PM, Guan Y, Stinson J, et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature. 2010;465(7297):473–7.
Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM, et al. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci Transl Med. 2010;2(62):62ra93.
Broet P, Dalmasso C, Tan EH, Alifano M, Zhang S, Wu J, et al. Genomic profiles specific to patient ethnicity in lung adenocarcinoma. Clin Cancer Res. 2011;17(11):3542–50.
Wilkerson MD, Yin X, Walter V, Zhao N, Cabanski CR, Hayward MC, et al. Differential pathogenesis of lung adenocarcinoma subtypes involving sequence mutations, copy number, chromosomal instability, and methylation. PLoS ONE. 2012;7(5):e36530.
Aramburu A, Zudaire I, Pajares MJ, Agorreta J, Orta A, Lozano MD, et al. Combined clinical and genomic signatures for the prognosis of early stage non-small cell lung cancer based on gene copy number alterations. BMC Genomics. 2015;16:752.
Walter K, Holcomb T, Januario T, Du P, Evangelista M, Kartha N, et al. DNA methylation profiling defines clinically relevant biological subsets of non-small cell lung cancer. Clin Cancer Res. 2012;18(8):2360–73.
Shi J, Marconett CN, Duan J, Hyland PL, Li P, Wang Z, et al. Characterizing the genetic basis of methylome diversity in histologically normal human lung tissue. Nat Commun. 2014;5:3365.
Karlsson A, Jonsson M, Lauss M, Brunnstrom H, Jonsson P, Borg A, et al. Genome-wide DNA methylation analysis of lung carcinoma reveals one neuroendocrine and four adenocarcinoma epitypes associated with patient outcome. Clin Cancer Res. 2014;20(23):6127–40.
Dennis JL, Hvidsten TR, Wit EC, Komorowski J, Bell AK, Downie I, et al. Markers of adenocarcinoma characteristic of the site of origin: development of a diagnostic algorithm. Clin Cancer Res. 2005;11(10):3766–72.
Wang YW, Tu PH, Lin KT, Lin SC, Ko JY, Jou YS. Identification of oncogenic point mutations and hyperphosphorylation of anaplastic lymphoma kinase in lung cancer. Neoplasia. 2011;13(8):704–15.
Chu YW, Yang PC, Yang SC, Shyu YC, Hendrix MJ, Wu R, et al. Selection of invasive and metastatic subpopulations from a human lung adenocarcinoma cell line. Am J Respir Cell Mol Biol. 1997;17(3):353–60.
Pan CM, Wang ML, Chiou SH, Chen HY, Wu CW. Oncostatin M suppresses metastasis of lung adenocarcinoma by inhibiting SLUG expression through coordination of STATs and PIASs signalings. Oncotarget. 2016;7(37):60395–406.
Yeh HW, Hsu EC, Lee SS, Lang YD, Lin YC, Chang CY, et al. PSPC1 mediates TGF-beta1 autocrine signalling and Smad2/3 target switching to promote EMT, stemness and metastasis. Nat Cell Biol. 2018;20(4):479–91.
Chen HY, Hu JY, Chen TH, Lin YC, Liu X, Lin MY, et al. KLHL39 suppresses colon cancer metastasis by blocking KLHL20-mediated PML and DAPK ubiquitination. Oncogene. 2015;34(40):5141–51.
Schneider CA, Rasband WS, Eliceiri KW. NIH image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9(7):671–5.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102(43):15545–50.
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34(3):267–73.
Milewski D, Balli D, Ustiyan V, Le T, Dienemann H, Warth A, et al. FOXM1 activates AGR2 and causes progression of lung adenomas into invasive mucinous adenocarcinomas. PLoS Genet. 2017;13(12):e1007097.
Carbajo-Pescador S, Ordonez R, Benet M, Jover R, Garcia-Palomo A, Mauriz JL, et al. Inhibition of VEGF expression through blockade of Hif1alpha and STAT3 signalling mediates the anti-angiogenic effect of melatonin in HepG2 liver cancer cells. Br J Cancer. 2013;109(1):83–91.
Chang CH, Zanini M, Shirvani H, Cheng JS, Yu H, Feng CH, et al. Atoh1 controls primary ccilia formation to allow for SHH-triggered granule neuron progenitor proliferation. Dev Cell. 2019;48(2):184-99.e5.
Kim J, Kang J, Kang YL, Woo J, Kim Y, Huh J, et al. Ketohexokinase-A acts as a nuclear protein kinase that mediates fructose-induced metastasis in breast cancer. Nat Commun. 2020;11(1):5436.
Tang TK. Centriole biogenesis in multiciliated cells. Nat Cell Biol. 2013;15(12):1400–2.
Tang CJ, Fu RH, Wu KS, Hsu WB, Tang TK. CPAP is a cell-cycle regulated protein that controls centriole length. Nat Cell Biol. 2009;11(7):825–31.
Godinho SA, Picone R, Burute M, Dagher R, Su Y, Leung CT, et al. Oncogene-like induction of cellular invasion from centrosome amplification. Nature. 2014;510(7503):167–71.
Leidel S, Delattre M, Cerutti L, Baumer K, Gonczy P. SAS-6 defines a protein family required for centrosome duplication in C. elegans and in human cells. Nat Cell Biol. 2005;7(2):115–25.
Hajra KM, Chen DY, Fearon ER. The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res. 2002;62(6):1613–8.
Micalizzi DS, Farabaugh SM, Ford HL. Epithelial-mesenchymal transition in cancer: parallels between normal development and tumor progression. J Mammary Gland Biol Neoplasia. 2010;15(2):117–34.
Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. The human transcription factors. Cell. 2018;172(4):650–65.
Kalinichenko VV, Major ML, Wang X, Petrovic V, Kuechle J, Yoder HM, et al. Foxm1b transcription factor is essential for development of hepatocellular carcinomas and is negatively regulated by the p19ARF tumor suppressor. Genes Dev. 2004;18(7):830–50.
Yang C, Chen H, Tan G, Gao W, Cheng L, Jiang X, et al. FOXM1 promotes the epithelial to mesenchymal transition by stimulating the transcription of Slug in human breast cancer. Cancer Lett. 2013;340(1):104–12.
Yusuf D, Butland SL, Swanson MI, Bolotin E, Ticoll A, Cheung WA, et al. The transcription factor encyclopedia. Genome Biol. 2012;13(3):R24.
Sadasivam S, Duan S, DeCaprio JA. The MuvB complex sequentially recruits B-Myb and FoxM1 to promote mitotic gene expression. Genes Dev. 2012;26(5):474–89.
**e Z, Tan G, Ding M, Dong D, Chen T, Meng X, et al. Foxm1 transcription factor is required for maintenance of pluripotency of P19 embryonal carcinoma cells. Nucleic Acids Res. 2010;38(22):8027–38.
Laoukili J, Kooistra MR, Bras A, Kauw J, Kerkhoven RM, Morrison A, et al. FoxM1 is required for execution of the mitotic programme and chromosome stability. Nat Cell Biol. 2005;7(2):126–36.
Wang IC, Chen YJ, Hughes D, Petrovic V, Major ML, Park HJ, et al. Forkhead box M1 regulates the transcriptional network of genes essential for mitotic progression and genes encoding the SCF (Skp2-Cks1) ubiquitin ligase. Mol Cell Biol. 2005;25(24):10875–94.
Wierstra I. The transcription factor FOXM1 (Forkhead box M1): proliferation-specific expression, transcription factor function, target genes, mouse models, and normal biological roles. Adv Cancer Res. 2013;118:97–398.
Stevens NR, Dobbelaere J, Brunk K, Franz A, Raff JW. Drosophila Ana2 is a conserved centriole duplication factor. J Cell Biol. 2010;188(3):313–23.
Gilkes DM, Semenza GL, Wirtz D. Hypoxia and the extracellular matrix: drivers of tumour metastasis. Nat Rev Cancer. 2014;14(6):430–9.
Bettencourt-Dias M, Rodrigues-Martins A, Carpenter L, Riparbelli M, Lehmann L, Gatt MK, et al. SAK/PLK4 is required for centriole duplication and flagella development. Curr Biol. 2005;15(24):2199–207.
Habedanck R, Stierhof YD, Wilkinson CJ, Nigg EA. The Polo kinase Plk4 functions in centriole duplication. Nat Cell Biol. 2005;7(11):1140–6.
Marthiens V, Rujano MA, Pennetier C, Tessier S, Paul-Gilloteaux P, Basto R. Centrosome amplification causes microcephaly. Nat Cell Biol. 2013;15(7):731–40.
Kulukian A, Holland AJ, Vitre B, Naik S, Cleveland DW, Fuchs E. Epidermal development, growth control, and homeostasis in the face of centrosome amplification. Proc Natl Acad Sci USA. 2015;112(46):E6311–20.
Vitre B, Holland AJ, Kulukian A, Shoshani O, Hirai M, Wang Y, et al. Chronic centrosome amplification without tumorigenesis. Proc Natl Acad Sci USA. 2015;112(46):E6321–30.
Acknowledgements
We thank the sequencing core facility (IBMS, AS-CFII-111-211), flow cytometry core facility (AS-CFII-111-212), and the IBMS confocal imaging core facility of Academia Sinica. We thank Dr. Hsing-Jien Kung for his critical reading and comments on this manuscript.
Funding
This work was supported by grants from the Ministry of Science and Technology, Taiwan (MOST 109-2326-B001-010), IBMS-CRC, and Academia Sinica (AS-IA-109-L04; AS-TP-108-L08).
Author information
Authors and Affiliations
Contributions
Conceptualization-YWW, SCC, and TKT; Methodology-YWW, DLG, SCC, and TKT; Investigation-YWW, DLG, SCC, KTY, JJT, and YCY; Resources-YSJ and TYC; Original draft-YWW and TKT; Writing-YWW and TKT; Review and editing-YWW, SCC, and TKT. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All animal protocols were performed according to the guidelines and approved by the Institutional Animal Care and Use Committee of Academia Sinica. Human tissue arrays were purchased from Pantomics and US Biomax, whose companies provided a certificate statement to confirm the legitimacy of tissue resources.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1.
Additional Tables S1–S4.
Additional file 2.
Additional Figures S1–S8.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Wang, YW., Chen, SC., Gu, DL. et al. A novel HIF1α-STIL-FOXM1 axis regulates tumor metastasis. J Biomed Sci 29, 24 (2022). https://doi.org/10.1186/s12929-022-00807-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12929-022-00807-0