Abstract
Given the existence of plenty of river valleys connecting Southeast and East Asia, it is possible that some inland route(s) might have been adopted by the initial settlers to migrate into the interior of East Asia. Here we analyzed mitochondrial DNA (mtDNA) HVS variants of 845 newly collected individuals from 14 Myanmar populations and 5,907 published individuals from 115 populations from Myanmar and its surroundings. Enrichment of basal lineages with the highest genetic diversity in Myanmar suggests that Myanmar was likely one of the differentiation centers of the early modern humans. Intriguingly, some haplogroups were shared merely between Myanmar and southwestern China, hinting certain genetic connection between both regions. Further analyses revealed that such connection was in fact attributed to both recent gene flow and certain ancient dispersals from Myanmar to southwestern China during 25–10 kya, suggesting that, besides the coastal route, the early modern humans also adopted an inland dispersal route to populate the interior of East Asia.
Similar content being viewed by others
Introduction
It is generally agreed that the peopling of East Asia resulted mainly from the Late Pleistocene south-to-north migrations, initiated from Southeast Asia by the earliest settlers after they migrated from Africa via a coastal route at ~60 kilo-years ago (kya)1,2,3,4,5,6,7. However, so far it remains elusive how these initial settlers migrated into the interior of East Asia. Although it is plausible that modern humans kept adopting the coastal route and moved along the coastline of the ancient Sundaland, reaching and finally leading to the settlement of East Asia, an alternative possibility that the settlers might have adopted an inland route into the interior of East Asia via river valleys could not be ruled out. Actually, as suggested by GIS-based analysis, river valleys had likely played an important role in populating the interior of South Asia by modern humans8. This opinion echoes with the suggestion that the major river systems in the northern mainland Southeast Asia, such as Ayeyarwady, Salween and Mekong, created diverse environments and paths for human dispersal and thus were of great help for early hominine adaptation9. The modern human cranium excavated from northern Laos, dated to 51–46 kya10, evinced the very early presence of modern humans in the interior of Southeast Asia and lent further support to this hypothesized inland dispersal scenario that likely occurred in the Late Pleistocene.
Unfortunately, so far no genetic trace of this inland dispersal(s) was observed, notwithstanding much progress has been achieved on dissecting the genetic landscapes in East Asians2,3,7,11,12,13,7,15,16,17,22. One possible reason is that only a few study took into account the genetic data from both East and Southeast Asian populations; more importantly, another reason may be attributable to the scarcity of genetic information from Myanmar, the largest country in mainland Southeast Asia which locates at the junction connecting South, Southeast and East Asia. Although Myanmar likely served as the corridor where the initial settlers had adopted to enter and colonize southeastern Asia during their migration along Asian coast4,6, previous studies focused either on the genetic structure of some ethnic populations18 or on the distribution of a single haplogroup (viz. M31) in Myanmar19.
Therefore, if this hypothesized ancient inland dispersal route did exist, Myanmar likely served as the corridor. In fact, the two major rivers (i.e. Ayeyarwady and Salween) in Myanmar can trace their upstream back to southwestern China. The existence of such river valleys would facilitate the potential population movement northwards into the interior zones. Coincidentally, our recent study has observed the enrichment of a number of new basal mtDNA lineages in southwestern China (especially Yunnan Province) and suggested this region likely to be the genetic reservoir of the modern humans after they entered East Asia13, further favoring the possibility of directly genetic contribution from Southeast Asia, say Myanmar, to southwestern China possibly occurred in the Late Pleistocene.
Results
Classification of mtDNA sequences in Myanmar populations
As shown in Supplementary Table S1 online, among the 845 Myanmar mtDNAs which were analyzed for their control-region and additional coding-region sites, the majority (532/845, 62.96%) could be allocated unambiguously into East Eurasian haplogroups, such as D, G, M7-M13, A, N9a, R9 and B4,11,13,16,20,21,22,23,24, whereas 4.26% (36/845) were assigned into haplogroups of South Asian ancestry6,25,26,27,28,29. Surprisingly, a high frequency of samples (269/845, 31.83%) could not be recognized based on control-region variation and partial coding-region information. Completely sequencing 64 representatives of these unrecognizable mtDNAs revealed that 225 of them belong in fact to certain sub-clades of the already defined haplogroups, e.g., M4, M5, M7, M20, M21, M24, M30, M33, M35, M45, M46, M49, M50, M51, M54, M55, M58, M60, M72, M76, M90, M91, R22, R31, N21 and HV (Supplementary Fig. S1 online); whereas the rest 44 mtDNAs were proven to represent 3 so far undefined basal lineages, for which could not find any sister clades after compared with over 20,666 mtDNA genomes worldwide (mtDNA tree Build 1630; http://www.phylotree.org/) and therefore were named as M82, M83 and M84 here (Supplementary Fig. S1 online).
Genetic relationship of Myanmar populations and their surrounding groups
After unambiguously determining all the 845 Myanmar mtDNAs under study, the proportion of haplogroups of East Eurasian (66.51%) and South Asian (17.40%) ancestries remains stable in the whole Myanmar population. The East Asian-prevalent haplogroups (i.e., M9a, A, D4, G and C2,11,Supplementary Table S1 online) which were chosen under the guideline of newly obtained mtDNA genome information (Supplementary Fig. S1 online). For any mtDNA whose phylogenetic status could not be identified yet, further complete sequencing work will be carried out. When naming the newly identified novel basal lineages, we followed the nomenclature listed in PhyloTree website (mtDNA tree Build 1630; http://www.phylotree.org/) and our recently suggested haplogroup scheme46. For haplogroups of interest, 28 additional representative samples were also chosen for complete sequencing. The experiments were carried out in accordance with the approved guideline of Chinese Academy of Sciences.
Data quality control
To ensure the quality of the complete genome data, our previously proposed quality-control measures, such as independent amplification, detecting errors by phylogenetic analysis and matching or near-matching method were followed6,11,28. Furthermore, to avoid the amplification of NUMT47 as well as the problem of artificial recombination that is easily introduced when dozens of primer sets are involved48, four pairs of PCR primers were designed to amplify the whole mtDNA genome. Each fragment was amplified independently. The amplified fragments, each of which contains more than 400bp overlap** regions with its neighbors49, were sequenced by use of 48 inner primers (12 for each fragment) reported in our previous studies6,20.
Data analyses
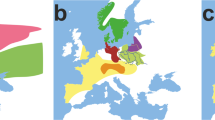
To facilitate the comparison with the reported data from the neighboring populations and thus to distill the genetic trace left by the Pleistocene inland immigrants, previously published mtDNA data, including 1,524 mtDNAs from mainland Southeast Asia and 3,120 mtDNAs from East Asia, 690 mtDNA sequences from northeast India and 302 from Bangladesh, were taken into account. In addition, the recently published Myanmar data (116 Barma and 155 Karen18) were also considered (Fig. 4 and Supplementary Table S6 online).
The PCA and CA were performed based on haplogroup frequencies by using Statistical Package for the Social Sciences (SPSS) 16.0 software. Reduced median network for each basal haplogroup was constructed manually and checked by the program NETWORK 4.510 (www.fluxus-engineering.com/sharenet.htm). Phylogenetic tree of haplogroup was reconstructed manually based on complete sequences and confirmed by mtphyl software. Contour maps of spatial frequencies of haplogroup were constructed using Kriging linear model of Surfer 8.0 (Golden Software Inc. Golden, Colorado, USA). Spatial analysis was performed using the PASSAGE software packet. Moran's I metrix was applied in correlogram analysis50. The time to the most recent common ancestor (TMRCA) of a haplogroup was estimated using ρ statistic method as described previously51. Nei's dA genetic distances52 and AMOVA were calculated by using the package Arlequin 3.11. Admixture estimation was performed by the Weighted Least Squares (WLS) Method53 using SPSS 16.0.
Additional Information
Accession numbers: All of the sequences obtained in the present study have been deposited into GenBank, with accession numbers KP345975-KP346066 for whole mtDNA genomes and KP346067-KP346911 for control region sequences.
References
Su, B. et al. Y-Chromosome evidence for a northward migration of modern humans into Eastern Asia during the last Ice Age. Am. J. Hum. Genet. 65, 1718–1724 (1999).
Yao, Y. G., Kong, Q. P., Bandelt, H. J., Kivisild, T. & Zhang, Y. P. Phylogeographic differentiation of mitochondrial DNA in Han Chinese. Am. J. Hum. Genet. 70, 635–651 (2002).
Shi, H. et al. Y-chromosome evidence of southern origin of the East Asian–specific haplogroup O3-M122. Am. J. Hum. Genet. 77, 408–419 (2005).
Macaulay, V. et al. Single, rapid coastal settlement of Asia revealed by analysis of complete mitochondrial genomes. Science 308, 1034 (2005).
Stringer, C. Palaeoanthropology: coasting out of Africa. Nature 405, 24–27 (2000).
Palanichamy, M. et al. Phylogeny of mitochondrial DNA macrohaplogroup N in India, based on complete sequencing: implications for the peopling of South Asia. Am. J. Hum. Genet. 75, 966–978 (2004).
Abdulla, M. A. et al. Map** human genetic diversity in Asia. Science 326, 1541–1545 (2009).
Field, J. S., Petraglia, M. D. & Lahr, M. M. The southern dispersal hypothesis and the South Asian archaeological record: examination of dispersal routes through GIS analysis. J. Anthropol. Archaeol. 26, 88–108 (2007).
Schepartz, L., Miller-Antonio, S. & Bakken, D. Upland resources and the early Palaeolithic occupation of southern China, Vietnam, Laos, Thailand and Burma. World Archaeol. 32, 1–13 (2000).
Demeter, F. et al. Anatomically modern human in Southeast Asia (Laos) by 46 ka. Proc. Natl. Acad. Sci. USA 109, 14375–14380 (2012).
Kong, Q. P. et al. Updating the East Asian mtDNA phylogeny: a prerequisite for the identification of pathogenic mutations. Hum. Mol. Genet. 15, 2076 (2006).
Kong, Q. P. et al. Mitochondrial DNA sequence polymorphisms of five ethnic populations from northern China. Hum. Genet. 113, 391–405 (2003).
Kong, Q. P. et al. Large-scale mtDNA screening reveals a surprising matrilineal complexity in east asia and its implications to the peopling of the region. Mol. Biol. Evol. 28, 513–522 (2011).
Wen, B., **e, X., Gao, S., Li, H., Shi, H. & Song, X. Analyses of genetic structure of Tibeto-Burman populations reveals sex-biased admixture in southern Tibeto-Burmans. Am. J. Hum. Genet. 74, 856–865 (2004).
Hill, C. et al. A mitochondrial stratigraphy for island southeast Asia. Am. J. Hum. Genet. 80, 29–43 (2007).
Hill, C. et al. Phylogeography and ethnogenesis of aboriginal Southeast Asians. Mol. Biol. Evol. 23, 2480–2491 (2006).
Bodner, M. et al. Southeast Asian diversity: first insights into the complex mtDNA structure of Laos. BMC Evol. Biol. 11, 49 (2011).
Summerer, M. et al. Large-scale mitochondrial DNA analysis in Southeast Asia reveals evolutionary effects of cultural isolation in the multi-ethnic population of Myanmar. BMC Evol. Biol. 14, 17 (2014).
Wang, H. W. et al. Mitochondrial DNA evidence supports northeast Indian origin of the aboriginal Andamanese in the Late Paleolithic. J. Genet. Genomics. 38, 117–122 (2011).
Kong, Q. P. et al. Phylogeny of east Asian mitochondrial DNA lineages inferred from complete sequences. Am. J. Hum. Genet. 73, 671–676 (2003).
Zhao, M. et al. Mitochondrial genome evidence reveals successful Late Paleolithic settlement on the Tibetan Plateau. Proc. Natl. Acad. Sci. USA 106, 21230–21235 (2009).
Peng, M. S. et al. Tracing the Austronesian footprint in Mainland Southeast Asia: a perspective from mitochondrial DNA. Mol. Biol. Evol. 27, 2417–2430 (2010).
Tabbada, K. A. et al. Philippine Mitochondrial DNA Diversity: A Populated Viaduct between Taiwan and Indonesia? Mol. Biol. Evol. 27, 21–31 (2010).
Tanaka, M. et al. Mitochondrial Genome Variation in Eastern Asia and the Peopling of Japan. Genome Res. 14, 1832–1850 (2004).
Chandrasekar, A. et al. Updating phylogeny of mitochondrial DNA macrohaplogroup m in India: dispersal of modern human in South Asian corridor. PLoS ONE 4, e7447 (2009).
Chaubey, G. et al. Phylogeography of mtDNA haplogroup R7 in the Indian peninsula. BMC Evol. Biol. 8, 227 (2008).
Kumar, S. et al. The earliest settlers' antiquity and evolutionary history of Indian populations: evidence from M2 mtDNA lineage. BMC Evol. Biol. 8, 230 (2008).
Sun, C. et al. The dazzling array of basal branches in the mtDNA macrohaplogroup M from India as inferred from complete genomes. Mol. Biol. Evol. 23, 683–690 (2006).
Thangaraj, K. et al. Reconstructing the origin of Andaman Islanders. Science 308, 996–996 (2005).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–394 (2009).
Wen, B. et al. Genetic evidence supports demic diffusion of Han culture. Nature 431, 302–305 (2004).
Moore, E. Bronze and iron age sites in upper Myanmar: Chindwin, Samon and Pyu. SBBR 1, 24–39 (2003).
Kuroiwa, Y. & Verkuyten, M. Narratives and the constitution of a common identity: The Karen in Burma. Identities-Glob. Stud. 15, 391–412 (2008).
Brown, W., George, M. & Wilson, A. Rapid evolution of animal mitochondrial DNA. Proc. Natl. Acad. Sci. USA 76, 1967–1971 (1979).
Diran, R. K. The Vanishing Tribes Of Burma 226–231 (Amphoto Art, London, UN, 1997).
Atkinson, Q. D., Gray, R. D. & Drummond, A. J. mtDNA variation predicts population size in humans and reveals a major Southern Asian chapter in human prehistory. Mol. Biol. Evol. 25, 468–474 (2008).
Li, K. S. Economic and cultural exchange between Yunnan, China and Southeast Asia and South Asia: from prehistory to warring state and Qin-Han Period. J. Guangxi. Univ. Nationalities. Nat. Sci. Ed. 17, 37–41 (2011).
Peng, C. L. Early cultural relationships of Yunnan-Guizhou plateau and mainland Southeast Asia. Corpus of Guangxi Museum(4). (Guangxi People's Publishing House, Guangxi, China, 2012).
Aung-Thaw The “Neolithic” Culture of the Padah-lin Caves. AP 14, 123–133 (1971).
Aung-Thwin, M. A. Burma before Pagan: The status of archaeology today. AP 25, 1–21 (1982).
Aung-Thwin, M. A. Origins and Development of the Field of Prehistory in Burma. AP 40, 6–34 (2001).
Yao, Y. G., Kong, Q. P., Wang, C. Y., Zhu, C. L. & Zhang, Y. P. Different matrilineal contributions to genetic structure of ethnic groups in the silk road region in china. Mol. Biol. Evol. 21, 2265–2280 (2004).
Coble, M. D. et al. Single nucleotide polymorphisms over the entire mtDNA genome that increase the power of forensic testing in Caucasians. Int. J. Legal. Med. 118, 137–146 (2004).
Finnila, S., Lehtonen, M. S. & Majamaa, K. Phylogenetic network for European mtDNA. Am. J. Hum. Genet. 68, 1475–1484 (2001).
Herrnstadt, C. et al. Reduced-median-network analysis of complete mitochondrial DNA coding-region sequences for the major African, Asian and European haplogroups. Am. J. Hum. Genet. 70, 1152–1171 (2002).
Kong, Q. P., Bandelt, H. J., Zhao, M., Zhang, Y. P. & Yao, Y. G. Reply to van Oven: Suggestions and caveats for naming mtDNA haplogroup. Proc. Natl. Acad. Sci. USA 107, E40–E41 (2010).
Yao, Y. G., Kong, Q. P., Salas, A. & Bandelt, H. J. Pseudomitochondrial genome haunts disease studies. J. Med. Genet. 45, 769–772 (2008).
Kong, Q. P. et al. Distilling artificial recombinants from large sets of complete mtDNA genomes. PLoS ONE 3, e3016 (2008).
Wang, H. W. et al. Strikingly different penetrance of LHON in two Chinese families with primary mutation G11778A is independent of mtDNA haplogroup background and secondary mutation G13708A. Mutat. Res. 643, 48–53 (2008).
Moran, P. A. Notes on continuous stochastic phenomena. BIOMETRIKA 37, 17–23 (1950).
Soares, P. et al. Correcting for purifying selection: an improved human mitochondrial molecular clock. Am. J. Hum. Genet. 84, 740–759 (2009).
Nei, M. Molecular evolutionary genetics. (Columbia University Press, New York & Guildford, England, 1987).
Long, J. C. et al. Genetic variation in Arizona Mexican Americans: estimation and interpretation of admixture proportions. Am. J. of Phys. Anthropol. 84, 141–157 (1991).
Andrews, R. M. et al. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat. Genet. 23, 147 (1999).
Acknowledgements
We thank the volunteers for participating in this project. We also thank Dr. Dong Wang for providing DNA of samples belonging to haplogroup M54. The work was supported by grants from the National Natural Science Foundation of China (31123005, 31322029).
Author information
Authors and Affiliations
Contributions
Q.-P.K. and Y.-P.Z. designed the research; L.-N.L., L.-Q.Y., C.-L.Z. and S.-F.W. colleted the samples; Y.-C.L., H.-W.W. and J.-Y.T. collected the data; Y.-C.L., H.-W.W., J.-Y.T. and C.-L.Z. performed the experiments; Y.-C.L., H.-W.W., J.-Y.T., Q.-P.K. and Y.-P.Z. analyzed data; Y.-C.L., Q.-P.K. and Y.-P.Z. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Supplementary Information
Dataset 1
Supplementary Information
Dataset 2
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, YC., Wang, HW., Tian, JY. et al. Ancient inland human dispersals from Myanmar into interior East Asia since the Late Pleistocene. Sci Rep 5, 9473 (2015). https://doi.org/10.1038/srep09473
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep09473
- Springer Nature Limited
This article is cited by
-
The history of prehistoric archaeology in Myanmar: a brief review
Asian Archaeology (2023)
-
The matrilineal ancestry of Nepali populations
Human Genetics (2023)
-
Investigating the palaeoenvironmental context of Late Pleistocene human dispersals into Southeast Asia: a review of stable isotope applications
Archaeological and Anthropological Sciences (2022)
-
Myanmar Burkholderia pseudomallei strains are genetically diverse and originate from Asia with phylogenetic evidence of reintroductions from neighbouring countries
Scientific Reports (2020)
-
Carriers of mitochondrial DNA macrohaplogroup L3 basal lineages migrated back to Africa from Asia around 70,000 years ago
BMC Evolutionary Biology (2018)