Abstract
X-chromosome inactivation (XCI) equalizes gene expression between the sexes by inactivating one of the two X chromosomes in female mammals. **st has been considered as a major cis-acting factor that inactivates the paternally derived X chromosome (Xp) in preimplantation mouse embryos (imprinted XCI). Ftx has been proposed as a positive regulator of **st. However, the physiological role of Ftx in female animals has never been studied. We recently reported that Ftx is located in the cis-acting regulatory region of the imprinted XCI and expressed from the inactive Xp, suggesting a role in the imprinted XCI mechanism. Here we examined the effects on imprinted XCI using targeted deletion of Ftx. Disruption of Ftx did not affect the survival of female embryos or expression of **st and other X-linked genes in the preimplantation female embryos. Our results indicate that Ftx is dispensable for imprinted XCI in preimplantation embryos.
Similar content being viewed by others
Introduction
Female mammals have a unique mechanism of gene dosage compensation called X-chromosome inactivation (XCI), which is an essential epigenetic process for development. In mice, inactivation of the paternally derived X chromosome (Xp), but not of the maternally derived X chromosome (Xm), is initiated in the preimplantation embryos (imprinted XCI)1,2,3. This imprinted inactivation of the Xp is maintained in post-implantation extra-embryonic tissues. At the blastocyst stage, reactivation of the Xp occurs in the inner cell mass that will later form the embryo. Thus, the imprinted XCI is erased once, subsequently, random XCI of either the Xp or Xm chromosomes is initiated in the post-implantation embryo by a cis-acting region on the X chromosome termed the X-inactivation centre (** are shown as red arrows. (b) Validation of recombination on the chromosomal long and short arm target sequences of ES cells (cell lines #64 and #77). (c) Genoty** results for wild-type females (+m/+p, where m is the maternally derived X chromosome and p is the paternally derived X chromosome), heterozygous females (+m/−p), homozygous females (−m/−p), wild-type males (+m/Y, where Y is the Y chromosome) and hemizygous males (−m/Y). The PCR primers (233, 234, and330) used for genoty** are depicted in Fig2-a. (d) Expression of Ftx in Ftx-deficient blastocysts. PCR primers used for RT–PCR are shown in parentheses. Uncropped images of the full-length gels are presented in Supplementary Figure S1. (e) The expression levels of miR-374-5p and miR-421-3p were quantified by q-PCR in wild-type female (+m/+p), male (+m/Y) and heterozygous female (+m/−p) blastocysts. The ratios of the number of cDNA copies for each miRNA to that of cDNA copies for miR-295 are indicated as expression levels. The bar graphs and lines show the mean ± standard deviation (SD) (n = 3).
Expression of miRNAs in Ftx-deficient blastocysts
Two miRNAs, miR-374-5p and miR-421-3p, are located within the Ftx intron in the same orientation as the Ftx transcript (Figure 1a). We confirmed that these miRNAs were not expressed in female blastocysts (+m/−p) when the Ftx-targeted allele was derived from the father, whereas these miRNAs were predominantly expressed in wild-type female blastocysts (+m/+p) (Figure 2e). This finding indicated that the Ftx-deficient mice failed to express miR-374-5p and miR-421-3p simultaneously and suggested that both miRNAs were transcribed from the same promoter as Ftx.
Mating of Ftx-deficient mice
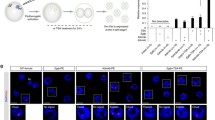
Paternal transmission of the ** gene, β-actin, are indicated as expression levels. The bar graphs and lines show the mean ± standard deviation (SD) (n = 3). (b) RNA FISH analysis for **st (red) expression in Ftx-deficient blastocysts. The nuclei of blastocysts were stained with DAPI (blue). Scale bar, 50 μm. (c) Summary of **st RNA FISH analysis. Three wild-type (+m/+p) and Three Ftx-deficient (+m/−p) blastocysts were examined. Cells with 2 **st signals were not observed. (d) Analysis of X-linked gene expression levels in Ftx-deficient blastocysts. No differences in the allelic expression patterns of X-linked genes between JF1 and C57BL/6N embryos were detected using RFLP. Uncropped images of the full-length gels are presented in Supplementary Figure S1.