Abstract
Multiple paternity in the olive ridley sea turtle (Lepidochelys olivacea) population nesting in Suriname was demonstrated using two microsatellite loci, viz., Ei8 and Cm84. The large number of offspring sampled per clutch (70 on average, ranging from 15 to 103) and the number of alleles found at the two loci (18 and eight alleles, respectively) enabled unambiguous assessment of the occurrence of multiple paternity. In two out of 10 clutches analysed, the offspring had been sired by at least two males, which was confirmed at both loci. In both clutches, unequal paternity occurred: 73% and 92% of the offspring had been sired by the primary male. The probability of detecting multiple paternity was 0.903, and therefore there is a small chance that multiple paternity occurred but remained undetected in some of the eight clutches that appeared to be singly sired. Analysis of 703 offspring revealed a high mutation rate for locus Ei8 (μ = 2.3 × 10−2) with all 33 mutations occurring in maternal alleles. In particular, one allele of 274 bp mutated at a high frequency in a clutch to which the mother contributed the allele, but in another clutch where the father contributed the same allele, no such mutations were observed. Inferred allele-specific mutation rates for Ei8 and expected numbers of mutations per clutch confirmed that maternal alleles for Ei8 are more likely to mutate in the olive ridley sea turtle than paternal alleles. Possible explanations are discussed.
Similar content being viewed by others
Introduction
Small or declining populations are vulnerable to inbreeding depression and random loss of genetic variation through genetic drift. Since genetic variation is important for adaptation to changing environmental conditions, the level of genetic diversity and the factors influencing the maintenance of genetic diversity have been a primary concern of conservation biologists. An important factor that affects the amount of genetic diversity in a population is the effective population size (Ne), which in turn is greatly influenced by the mating system of a species. In a polyandrous mating system, females mate with several males within a single reproductive cycle. This may result in multiple paternity of clutches, which occurs when several males contribute to the offspring in a single clutch. In such a mating system, Ne increases, and as a result, more genetic variation can be maintained than in a monogamous mating system (Sugg and Chesser, 1994; Moran and Garcia-Vazquez, 1998).
Populations of marine turtles (order Testudines) have decreased in many parts of the world due to human factors. Until recently, the Surinam olive ridley (Lepidochelys olivacea) population was considered to be the largest in the Atlantic, but the number of nesting females has declined rapidly (Reichart and Fretey, 1993). In the late 1960s, the population nesting in the Galibi Nature Reserve in East Suriname was large enough to produce arribadas (mass nesting events) (Schulz, 1975), but the estimated number of clutches laid has declined continuously from 3063 in 1968 (when counting started) to 335 in 1995 (Schulz, 1975; Reichart and Fretey, 1993; Hoekert et al, 1996).
In recent years, the occurrence of multiple paternity has been investigated using allozymes, minisatellites or microsatellites in several marine turtle species from the Cheloniidae and Dermochelyidae. Several studies indicated the occurrence of multiple paternity in loggerhead turtles (Caretta caretta) (Harry and Briscoe, 1988; Bollmer et al, 1999), the Kemp's ridley (Lepidochelys kempi) (Kichler et al, 1999), and green turtles (Chelonia mydas) (Parker et al, 1996). In most cases however, multiple paternity was either observed at only one of several loci, indicated by deviations from Mendelian expectations, or the number of offspring sampled per clutch was low. In other studies, some of which were extensive, multiple paternity was not observed, or only at a very low level in green turtles (FitzSimmons, 1998) or leatherback turtles (Dermochelys coriacea) (Rieder et al, 1998; Curtis et al, 2000; Dutton et al, 2000).
The highly polymorphic nature of microsatellite markers, caused by high mutation rates, makes them attractive tools in paternity studies. However, the occurrence of mutations may lead to the false conclusion of multiple paternity, if more than two paternal alleles are detected among the progeny. Therefore, in order to rule out this possibility, multiple loci should be analysed to test whether multiple paternity can be confirmed at different loci. The exact mechanism of microsatellite mutation is not yet known, but it is generally accepted that DNA polymerase slippage is the predominant mechanism creating new alleles (Tautz and Schlötterer, 1994), usually resulting in the gain or loss of a single repeat unit.
The aim of this study was to investigate whether multiple paternity occurs in the olive ridley sea turtle population nesting in Suriname using microsatellite markers. The extensive sampling also provided a good opportunity to collect data on microsatellite mutation.
Materials and methods
Fieldwork and sampling
Blood samples were collected from nesting olive ridley females in the Galibi Nature Reserve in East Suriname at the beginning of the 1995 nesting season. Ten females were sampled and immediately after egg deposition their clutches were transferred to a hatchery located on a nearby beach. In addition, blood samples from two other nesting females were taken. A sterile syringe (0.6 × 25 mm) was used to take 0.5 ml of blood from the dorsal cervical sinus (Bennet, 1986). Several days before the expected hatching date, screens were placed around the nests to collect the hatchlings. After hatching, blood samples (<0.2 ml) were taken from most of the emerged hatchlings, again from the dorsal cervical sinus (Bennet, 1986) before releasing the hatchlings. A total of 703 offspring from 10 clutches, ranging from 15 to 103 offspring per clutch, were sampled (Table 1). Blood samples were stored at ambient temperature in lysis buffer (100 mM Tris (pH 8), 100 mM EDTA, 10 mM NaCl, and 1% SDS). The nests were dug up after hatching and the hatching success (defined as the proportion of emerged offspring) was determined.
Microsatellite analysis
Nuclear DNA was extracted from the blood samples by proteinase K digestion (overnight) followed by ethanol precipitation (FitzSimmons et al, 1995). The occurrence of multiple paternity was investigated using Ei8 and Cm84, two dinucleotide microsatellite loci obtained from Eretmochelys imbricata (the hawksbill turtle) and C. mydas, respectively (FitzSimmons et al, 1995). For Cm84, a re-designed primer was used that resulted in smaller, better scorable alleles (K Kichler, personal communication). PCR was carried out in 10 μl reactions that contained 50–100 ng of nuclear DNA, 0.4 pmol of forward primer, end-labelled with γ32P-ATP, 1.6 pmol of unlabelled forward primer, 2 pmol of reverse primer, 0.1 mM dNTPs, 0.4 U Super Taq (Sphaero Q, Leiden, The Netherlands) and 1 μl Super Taq buffer. A 4 min denaturing step at 94°C was followed by 25 (Ei8) or 30 (Cm84) cycles of 30 sec denaturation at 94°C, 30 sec annealing at 56°C (Ei8) or 57°C (Cm84), and 25 sec extension at 72°C followed by a final extension of 5 min at 72°C. The PCR products were run on a 6% polyacrylamide gel (Sequagel) at 60 W for 4–5 h and visualised by autoradiography. The size of the alleles was determined by running them next to sequenced plasmid DNA with known sequence.
Data analysis
The genotypes of 25 unrelated adult individuals were determined in order to estimate the allele frequencies for Ei8 and Cm84 in the breeding population. These individuals consisted of the 12 nesting females sampled in 1995, 12 breeding males whose genotypes were inferred from the offspring data and one nesting female sampled in 1997. It was assumed that all the sampled females and the inferred males were unrelated. Allele frequencies at Ei8 and Cm84 were tested for deviation from Hardy-Weinberg equilibrium using a Markov chain method in GENEPOP 3.1 (Raymond and Rousset, 1995). The same method was used to test for linkage disequilibrium between the two loci. The probability of two unrelated individuals sharing the same genotype at both loci was calculated following Hanotte et al (1991). The probability of detecting multiple paternity (D) using Ei8 and Cm84 in the Surinam olive ridley population was calculated according to Westneat et al (1987). This value (D) depends on the allele frequencies in the breeding population and takes into account all genotypic combinations that will not allow the detection of multiple paternity (for instance, two males sharing the same genotype, or two males homozygous for different alleles). In order to detect with 95% confidence any male fathering at least five offspring, it is required to sample 50% of a clutch, regardless of the clutch size (Galbraith et al, 1989). Except for two clutches with a low hatching success, 50% to 81% of the eggs in each clutch was analysed. All offspring (n = 703) were screened at both loci, and paternal alleles were inferred for each clutch by comparing the progeny to their mother. Chi-square tests were used to test whether the distribution of genotypes in the progeny followed Mendelian expectations. To this end, mutant alleles were grouped with the allele they were presumed to have originated from, which is the allele that is closest in size to the mutant allele. The occurrence of an unexpected maternal allele in a descendant, ie, an allele that was not present in the mother, was designated a mutation, while the occurrence of more than two paternal alleles among the offspring in a clutch could either be explained by mutation or by multiple paternity. Multiple paternity was concluded when (i) more than two paternal alleles at a locus were found among the progeny, or if deviations from Mendelian inheritance were observed, and (ii) multiple paternity was supported at both loci in individual offspring, making the possibility of mutation highly unlikely.
In order to calculate allele-specific mutation rates at Ei8, the alleles that were found among the 703 offspring were grouped in size classes of 10 bp, and the number of mutations in each size class was divided by the number of observations of each size class. It was assumed that the alleles differed only in the number of repeat units, and that the mutation rate depended on the number of repeat units. The expected number of mutations per clutch was calculated as follows: for each allele in a clutch, the expected number of mutations was calculated by multiplying the number of offspring carrying the allele with the corresponding allele-specific mutation rate. The expected number of mutations per clutch was then calculated by summing the expected values for all the alleles in a clutch.
Results
Population variability
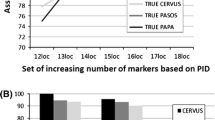
Among 25 unrelated individuals, 18 and eight alleles were found at microsatellites Ei8 and Cm84, respectively (Figure 1). Despite the fact that Cm84 is a dinucleotide repeat, most alleles for this locus differed by only one bp (Figure 1), which was also supported by two additional alleles (193 and 196 bp) that were found in unrelated olive ridley hatchlings sampled in 1997 (WEJ Hoekert et al, unpublished data). Allele frequencies at both loci were within expectations of Hardy-Weinberg equilibrium (P > 0.05), and no evidence for linkage disequilibrium between Ei8 and Cm84 was found (P > 0.05). The expected heterozygosity was 0.89 for Ei8 and 0.75 for Cm84. Observed heterozygosities were slightly higher: 0.92 and 0.80 for Ei8 and Cm84, respectively. The probability of two different males sharing the same genotype at Ei8 as well as Cm84 was 1.9 × 10−3, and the probability of detecting multiple paternity across both loci in the Surinam olive ridley population was 0.903.
Multiple paternity
The hatching success of the 10 clutches ranged from 14% to 83% (average 63%, Table 1). Maternal and inferred paternal alleles for each clutch are given in Table 1. Multiple paternity was found in two clutches, as shown by the number of paternal alleles and the distribution of paternal alleles (Table 1). Among the offspring in clutch Lo17, four paternal alleles were found at Ei8 and four paternal alleles were found at Cm84, strongly suggesting that this clutch had been fertilised by at least two males (Tables 1 and 2). Among the offspring in clutch Lo9, four paternal alleles were found at Ei8, and a strong deviation from Mendelian expectations was found at Cm84, both indicating multiple paternity. Of the two paternal alleles that were found at Cm84 in clutch Lo9, one (197 bp) was found at a very low frequency: only four out of 75 progeny contained this allele (Table 1). This considerable deviation from Mendelian expectations, under the assumption of a single father, strongly suggested the involvement of a second father that sired only a small proportion of this clutch. This was confirmed by the fact that the 197 bp allele was only present in offspring that had a 254 bp or 234 bp allele at Ei8 (Tables 1 and 2). Therefore, it was highly unlikely that the 197 bp alleles at Cm84 were the result of mutation, and this was also supported by the absence of mutations at Cm84 in the remaining progeny (see below).
All offspring in Lo9 and Lo17 that showed unexpected alleles at Ei8 also showed unexpected alleles at Cm84, meaning that multiple paternity was confirmed at both loci (Table 2). By combining data for both loci, unique genotypes could be assigned to each father, enabling the determination of the number of progeny sired by each male. Of the 71 offspring in clutch Lo17, 52 were sired by the primary male (defined as the male that sired most of the offspring) and 19 by the secondary male. Of the 75 offspring in Lo9, 69 were sired by the primary male and six by the secondary male.
In all 10 clutches, genotype frequencies in the offspring were in accordance with Mendelian expectations. When taking into account the two different fathers in clutches Lo9 and Lo17, Mendelian expectations for each father were met at both loci. Assuming null alleles, if the mother was heterozygous and the father was heterozygous for a null allele and an allele different from both maternal alleles, then three paternal alleles would be detected among the progeny, and a false conclusion of multiple paternity could be reached (Kichler et al, 1999). This scenario did not occur in the two clutches displaying multiple paternity (Tables 1 and 2).
Mutation rate
Analysis of 703 offspring provided information on the mutation rate at both microsatellite loci. At Ei8, 15 different mutant alleles were found in 33 out of 703 offspring (Table 3), giving a mutation rate of 2.3 × 10−2. At Cm84, no mutations were found. The unexpected alleles at Ei8 were identified as mutations by two observations: (i) none of the 33 progeny had unexpected alleles at Cm84, and (ii) each mutation occurred in the maternal allele, as shown by the absence of a maternal allele and the presence of a paternal allele in each of the 33 offspring. The second observation confirmed that the 33 unexpected alleles were caused by mutation and not by multiple paternity.
Most mutations (n = 23) resulted in the gain of one repeat unit (two base pairs). In only one case, one repeat unit had been lost. In the other nine cases, two to six repeat units had been lost or gained (Table 3). Mutations were not evenly distributed among the clutches. Some clutches did not contain mutations at all, while in clutch Lo7r, 14% of the progeny (viz., 14 out of 103 individuals) inherited a mutation. Large alleles (more repeat units) were more sensitive to mutation than small ones (Tables 3 and 4).
All mutations occurred in maternal Ei8 alleles. In order to analyse this unequal distribution of mutations, allele-specific mutation rates were determined for all the Ei8 alleles observed in the 10 clutches (Table 4). Using the allele-specific mutation rates, the expected number of mutations could be calculated for each clutch. The expected number of mutations per clutch differed significantly from the observed number of mutations in clutch Lo7r (χ2 = 5.45, df = 1, P < 0.05) and clutch Lo17 (χ2 = 5.27, df = 1, P < 0.05) (Table 5). In clutch Lo7r, the difference was caused by an excess of mutations in the maternal 274 bp allele: only seven mutations were expected to occur in the 274 bp allele, yet 14 mutations occurred. In clutch Lo17, the difference was caused by an absence of mutations in the paternal 270 bp allele: at least four mutations were expected in the 270 bp allele, whereas none occurred (Table 5). Grou** of alleles in other size classes such as classes of 5 bp, 15 bp, or 20 bp, and/or shifting the size classes to different ranges yielded similar results: a significant difference between expected and observed mutations in clutches Lo7r and Lo17 (data not shown).
Discussion
Multiple paternity
The microsatellite data presented in this study unambiguously show that multiple paternity occurs in the olive ridley sea turtle. Clutches Lo9 and Lo17 were shown to be fertilised by at least two males, and in both cases, multiple paternity was confirmed at both loci (Tables 1 and 2). Furthermore, unequal paternal contribution occurred; 92% (Lo9) and 73% (Lo17) of the progeny had been sired by the primary male. The probability of detecting multiple paternity was 0.903, which means that for each clutch that was sired by one male, there is a chance of about 10% that multiple paternity did occur but remained undetected. Therefore, the level of multiple paternity in the Surinam olive ridley population might be slightly higher than appeared from this study.
We cannot tell whether the observed multiple paternity resulted from multiple matings within one breeding season, or whether sperm stored from a previous nesting season fertilised part of the eggs. During the nesting season, Pacific olive ridleys assemble into the so-called reproductive patch (RP), a large dynamic aggregation of mature reproductive males and females engaged in courtship and mating or waiting for the next arribada (Kalb et al, 1992). Under these conditions, multiple matings could easily occur, and thus promiscuity could be common in olive ridleys. However, in the Surinam olive ridley population, there is no evidence for the existence of a RP, and arribadas did not occur anymore since the late 1960s. Alternatively, sperm stored from a previous nesting season could have contributed to the progeny. Sperm storage tubules containing sperm have been identified within the oviduct of several turtle species including the olive ridley (Gist and Jones, 1989), and in box turtles (Terrapene carolina carolina), sperm storage for up to 4 years has been reported (Ewing, 1943). This is much longer than the average internesting period for Surinam olive ridleys which nest on average every 1 or 2 years (Schulz, 1975). In copperhead snakes, multiple paternity resulting from autumn- and subsequent spring copulations was demonstrated, and in two out of three cases, most or all offspring were sired by the first male (Schuett and Gillingham, 1986). However, no multiple paternity occurred when females mated only once a year, in spring, at the time of ovulation. Extending these results to the olive ridley, and considering the relatively high proportion of offspring (27%) sired by the secondary male in clutch Lo17 suggests that the matings in both multiply sired clutches might have occurred within the same nesting season. If true, the observed unequal paternity could be explained by sperm competition, cryptic female choice (Jennions and Petrie, 2000; and references therein), or differences in genetic compatibility between certain males and females (Zeh, 1997).
The level of multiple paternity found in this study is lower than the one found in Kemp's ridleys, for which it was suggested that probably all clutches might have been fertilised by multiple males (Kichler et al, 1999). It should be noted that the Surinam population is not representative of a stable olive ridley population. Numbers have been, and still are, declining rapidly (Van Tienen, personal communication) and arribadas have not been observed in the last 30 years. Therefore, levels of multiple paternity could differ considerably in other, larger populations.
Mutation rate
The mutation rate at locus Ei8 (μ = 2.3 × 10−2) contrasted greatly with the one at Cm84, for which no mutations were observed. The high mutation rate for Ei8 differs from values reported in a study on the closely related Kemp's ridley (Kichler et al, 1999). In that study, maximum likelihood analyses were performed on data for three microsatellite loci (among which Ei8 and Cm84) in 161 offspring from 36 Kemp's ridley clutches. It was concluded that the maximum likelihood estimate of the overall mutation rate was 0, although rates as high as 0.01 could not be rejected (Kichler et al, 1999). The mutation rate at Ei8 is relatively high compared to values found in green turtles, which ranged from <5.7 × 10−4 to 9.6 × 10−3 (FitzSimmons, 1998), and comparable to the highest mutation rate observed so far for a microsatellite locus (3.6 × 10−2, Primmer and Ellegren, 1996). The fact that large alleles mutated more often than small ones (Tables 3 and 4) is in agreement with the hypothesis that microsatellite alleles with a larger number of repeats tend to have higher mutation rates (Weber, 1990), and with results from other studies (eg, Primmer and Ellegren, 1996; Schlötterer et al, 1998). Most of the mutations (85%) involved an increase in allele size (Table 3), which is also in agreement with other studies (Weber and Wong, 1993; Primmer and Ellegren, 1996; Cooper et al, 1999).
Surprisingly, all 33 mutations occurred in maternal Ei8 alleles. It is generally believed that mutation rates are much higher in males than in females because of the many more cell divisions in spermatogenesis than in oogenesis (Crow, 1993). Indeed, Weber and Wong (1993) observed a bias towards paternal mutation events in their study on human STRPs, and a strong tendency for mutations to occur in male germlines was found in a number of VNTRs (eg, Vergnaud et al, 1991). In birds, a male-biased mutation rate, manifested as faster evolution of the Z chromosome than of the W chromosome has been demonstrated (Carmichael et al, 2000, and references therein). On the other hand, no sexual bias was observed in mutation rates at microsatellite loci in green turtles (FitzSimmons, 1998) and leatherback turtles (Dutton et al, 2000), nor at particular human VNTR loci (eg, Jeffreys et al, 1991). One example, however, of female biased mutation is found at the fragile X mutation site, an exon in the human FMR-1 gene containing a highly polymorphic trinucleotide repeat (Fu et al, 1991, and references therein).
Most of the mutations (n = 21) occurred in two of the largest clutches, viz., Lo7r (n = 103) and Lo19 (n = 80). In these clutches, the mothers contributed large alleles and the fathers contributed small alleles (Table 1). Since larger alleles mutated at a higher rate, this may have contributed to the observed female-biased mutation. However, the fathers in other clutches had large alleles as well, and mutations in these alleles should have occurred if we assume no sexual bias. A conspicuous example of female-biased mutation at locus Ei8 is the absence of mutations in the paternal 274 bp allele among 14 offspring in clutch Lo3, while the maternal 274 bp allele in clutch Lo7r mutated in 14 out of 52 offspring (Tables 3 and 5). The observed number of mutations in clutch Lo7r differed significantly from the expected number of mutations (χ2 = 5.45, df = 1, P < 0.05), which was caused by an excess of mutations in the maternal 274 bp allele (Table 5). This observation suggests that maternal alleles for Ei8 are more prone to mutation than paternal alleles of the same size. The discrepancy might also be explained by the occurrence of allelic homoplasy due to variable flanking sequences. In a study on a human microsatellite locus, sequence data revealed that alleles of the same size could differ considerably in internal structure, and that a high degree of variability was due to variation in the flanking sequences (Grimaldi and Crouau-Roy, 1997). It is possible that the number of repeats in the maternal 274 bp allele in clutch Lo7r is higher than that of the paternal 274 bp allele in clutch Lo3 and that the difference in repeat number is masked by size differences in the flanking sequences. Indeed, sequence data from a Pacific olive ridley individual indicated that a 198 bp allele of locus Ei8 probably contains a CA16 repeat (FitzSimmons et al, 1995; N FitzSimmons, personal communication), leaving more than 160 bp of flanking sequences. Although allelic homoplasy could explain the observed results for a given allele, we regard it as unlikely that the overall absence of paternal mutations was caused by allelic homoplasy, because that would require the studied females and males to come from two distinct populations. Another explanation for the 274 bp alleles might be that the paternal 274 bp allele has a point mutation that interrupts the repeat, thus reducing the chance of DNA slippage.
Large size differences between the two microsatellite alleles in an individual might result in an increased mutation rate (Amos et al, 1996; Schlötterer et al, 1998). However, although the mother of clutch Lo7r did indeed have one of the largest size differences (206 and 274 bp), the size difference was even more pronounced in the fathers of clutch Lo3 (198 and 274 bp) and Lo17 (198 and 270 bp), two clutches in which no mutations were found. Moreover, the size difference in the mother of clutch Lo19 (254 and 266 bp) was very low, yet seven mutations occurred. Therefore, the observed bias towards maternal mutations cannot be explained by this mechanism. On the contrary, if this mechanism really exists, it confirms that mutation is restricted to females. The mutational pattern observed at Ei8 needs further study. Sequencing of alleles will be an important step in explaining the results, providing more insight in the process of microsatellite mutation and evolution in general, and in the olive ridley sea turtle in particular.
References
Amos W, Sawcer SJ, Feakes RW, Rubinsztein DC (1996). Microsatellites show mutational bias and heterozygote instability. Nat Genet, 13: 390–391.
Bennet JM (1986). A method for sampling blood from hatchling loggerhead turtles. Herp Rev, 17: 43.
Bollmer JL, Irwin ME, Rieder JP, Parker PG (1999). Multiple paternity in loggerhead turtle clutches. Copeia, 2: 475–478.
Carmichael AN, Fridolfsson AK, Halverson J, Ellegren H (2000). Male-biased mutation rates revealed from Z and W chromosome-linked ATP synthase α-subunit (ATP5a1) sequences in birds. J Mol Evol, 50: 443–447.
Cooper G, Burroughs NJ, Rand DA, Rubinsztein DC, Amos W (1999). Markov chain Monte Carlo analysis of human Y-chromosome microsatellites provides evidence of biased mutation. Proc Natl Acad Sci USA, 96: 11916–11921.
Crow JF (1993). How much do we know about spontaneous human mutation rates? Environ Mol Mutagen, 21: 122–129.
Curtis C, Williams CJ, Spotila JR (2000). Mating system of Caribbean leatherback turtles as indicated by analysis of microsatellite DNA from hatchlings and adult females. In: Abreu-Grobois FA, Briseño-Dueñas R, Márquez R, Sarti L (eds) Proceedings of the Eighteenth Annual Symposium on Sea Turtle Biology and Conservation, NOAA Technical Memorandum NMFS-SEFSC- 436: p 155.
Dutton PH, Bixby E, Davis SK (2000). Tendency towards single paternity in leatherbacks detected with microsatellites. In: Abreu-Grobois FA, Briseño-Dueñas R, Márquez R, Sarti L (eds) Proceedings of the Eighteenth Annual Symposium on Sea Turtle Biology and Conservation, NOAA Technical Memorandum NMFS-SEFSC- 436: p 39.
Ewing HE (1943). Continued fertility in female box turtles following mating. Copeia, 2: 112–114.
Fitzsimmons NN, Moritz C, Moore SS (1995). Conservation and dynamics of microsatellite loci over 300 million years of marine turtle evolution. Mol Biol Evol, 12: 432–440.
Fitzsimmons NN (1998). Single paternity of clutches and sperm storage in the promiscuous green turtle (Chelonia mydas). Mol Ecol, 7: 575–584.
Fu YH, Kuhl DPA, Pizzuti A, Pieretti M, Sutcliffe JS, Richards S et al. (1991). Variation of the CGG repeat at the fragile X site results in genetic instability: resolution of the Sherman paradox. Cell, 67: 1047–1058.
Galbraith DA, Boag PT, White BN (1989). Sample size decisions in fingerprinting large clutches. Fingerprint News, 2: 8–10.
Gist DH, Jones JM (1989). Sperm storage within the oviduct of turtles. J Morph, 199: 379–384.
Grimaldi MC, Crouau-Roy B (1997). Microsatellite allelic homoplasy due to variable flanking sequences. J Mol Evol, 44: 336–340.
Hanotte OT, Burke T, Armour JAL, Jeffreys AJ (1991). Hypervariable minisatellite DNA sequences in the Indian pea fowl Pavo cristatus. Genomics, 9: 587–597.
Harry JL, Briscoe DA (1988). Multiple paternity in the loggerhead turtle (Caretta caretta). J Hered, 79: 96–99.
Hoekert WEJ, Schouten AD, Van Tienen LHG, Weijerman M (1996). The Sea Turtles of Galibi, Suriname. An update on status and nesting ecology. Technical report No 1, Biotopic Foundation, Amsterdam, The Netherlands: 35pp.
Jeffreys AJ, Macleod A, Tamaki K, Neil DL, Monckton DG (1991). Minisatellite repeat coding as a digital approach to DNA ty**. Nature, 354: 204–209.
Jennions MD, Petrie M (2000). Why do females mate multiply? A review of the genetic benefits. Biol Rev, 75: 21–64.
Kalb H, Valverde R, Owens D (1992). What is the reproductive patch of the olive ridley sea turtle? In: Richardson JA, Richardson TH (eds) Proceedings of the Twelfth Annual Workshop on Sea Turtle Biology and Conservation, NOAA Technical Memorandum NMFS-SEFSC- 361: pp57–60.
Kichler K, Holder MT, Davis SK, Marquez-M R, Owens DW (1999). Detection of multiple paternity in the Kemp's ridley sea turtle with limited sampling. Mol Ecol, 8: 819–830.
Moran P, Garcia-Vazquez E (1998). Multiple paternity in Atlantic salmon: a way to maintain genetic variability in relicted populations. J Hered, 89: 551–553.
Parker PG, White TA, Peare T (1996). Paternity studies in animal populations. In: Smith TB, Wayne RK (eds) Molecular Genetic Approaches in Conservation, Oxford University Press: New York. pp 413–423.
Primmer CR, Ellegren H (1996). Directional evolution in germline microsatellite mutations. Nat Genet, 13: 391–393.
Raymond M, Rousset F (1995). GENEPOP (V.1.2): a population genetics software for exact tests and ecumenicism. J Hered, 86: 248–249.
Reichart HA, Fretey J (1993). WIDECAST Sea Turtle Recovery Action Plan for Suriname (Eckert KL, editor). CEP Technical Report No. 24. UNEP Caribbean Environment Programme, Kingston, Jamaica xiv + 65 pp.
Rieder JP, Parker PG, Spotila JR, Irwin ME (1998). The mating system of the leatherback turtle: a molecular approach. In: Byles R, Fernandez Y (eds) Proceedings of the Sixteenth Annual Symposium on Sea Turtle Biology and Conservation, NOAA Technical Memorandum NMFS-SEFSC-412. pp 120–121.
Schlötterer C, Ritter R, Harr B, Brem G (1998). High mutation rate of a long microsatellite allele in Drosophila melanogaster provides evidence for allele-specific mutation rates. Mol Biol Evol, 15: 1269–1274.
Schuett GW, Gillingham JC (1986). Sperm storage and multiple paternity in the copperhead, Agkistrodon contortrix. Copeia, 3: 807–811.
Schulz JP (1975). Sea turtles nesting in Suriname. Zoologische Verhandelingen, Rijksmuseum van Natuurlijke Historie: Leiden, No.143 143 pp.
Sugg DW, Chesser RK (1994). Effective population sizes with multiple paternity. Genetics, 137: 1147–1155.
Tautz D, Schlötterer C (1994). Simple sequences. Curr Opin Genet Devel, 4: 832–837.
Vergnaud G, Mariat D, Apiou F, Aurias A, Lathrop M, Lauthier V (1991). The use of synthetic tandem repeats to isolate new VNTR loci: cloning of a human hypermutable sequence. Genomics, 11: 135–144.
Weber JL (1990). Informativeness of human (dC-dA)n* (dG-dT)n polymorphisms. Genomics, 7: 524–530.
Weber JL, Wong C (1993). Mutation of human short tandem repeats. Hum Mol Genet, 2: 1123–1128.
Westneat DF, Fredrick PC, Haven Wiley R (1987). The use of genetic markers to estimate the frequency of successful alternative reproductive tactics. Behav Ecol Sociobiol, 21: 35–45.
Zeh JA (1997). Polyandry and enhanced reproductive success in the harlequin-beetle-riding pseudoscorpion. Behav Ecol Sociobiol, 40: 111–118.
Acknowledgements
We would like to thank H Reichart (LBB), M Held, C Bram, and H Sijlbing (STINASU) and R Pané, for making it possible to work in the Galibi Nature Reserve; K Kichler, N FitzSimmons and B Bowen for providing DNA samples; N FitzSimmons for providing primers; L van Tienen and M Weijerman for their help with the fieldwork and sampling; and M Girondot, N FitzSimmons, K Vijverberg, L Raijmann, and I Williams for useful comments on the manuscript. Financial support came from the Beijerinck-Pop** Foundation, the Society for the Advancement of Research in the Tropics (Treub Society), and the World Wildlife Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hoekert, W., Neuféglise, H., Schouten, A. et al. Multiple paternity and female-biased mutation at a microsatellite locus in the olive ridley sea turtle (Lepidochelys olivacea). Heredity 89, 107–113 (2002). https://doi.org/10.1038/sj.hdy.6800103
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800103
- Springer Nature Switzerland AG
Keywords
This article is cited by
-
Do female amphibians and reptiles have greater reproductive output if they have more mates?
Behavioral Ecology and Sociobiology (2022)
-
Is multiple paternity in elasmobranchs a plesiomorphic characteristic?
Environmental Biology of Fishes (2020)
-
Microsatellite instability testing in colorectal cancer using the QiaXcel advanced platform
BMC Cancer (2018)
-
Multiple paternity in a Pacific marine turtle population: maternal attributes, offspring outcomes and demographic inferences
Marine Biology (2018)
-
Polyandry and multiple paternities in the threatened Agassiz’s desert tortoise, Gopherus agassizii
Conservation Genetics (2011)