Abstract
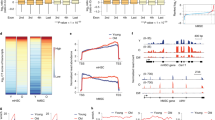
Mammalian aging is characterized by the progressive loss of tissue function and increased risk for disease. Accumulation of senescent cells in aging tissues partly contributes to this decline, and targeted depletion of senescent cells in vivo ameliorates many age-related phenotypes. The fundamental molecular mechanisms responsible for the decline of cellular health and fitness during senescence and aging are largely unknown. In this study, we investigated whether chromatin-mediated loss of transcriptional fidelity, known to contribute to fitness and survival in yeast and worms, also occurs during human cellular senescence and mouse aging. Our findings reveal aberrant transcription initiation inside genes during senescence and aging that co-occurs with changes in the chromatin landscape. Interventions that alter these spurious transcripts have profound consequences on cellular health, primarily affecting intracellular signal transduction pathways. We propose that age-related spurious transcription promotes a noisy transcriptome and degradation of coherent transcriptional networks.






Similar content being viewed by others
Data availability
The GEO accession number for all genome-wide datasets generated in this article is GSE156829. The following publicly available datasets downloaded from the GEO and used in this study are:
1. Shah et al., 2013 – GSE36616
2. Cruickshanks et al., 2013 – GSE48580
3. Rai et al., 2014 – GSE56307
4. Sen et al., 2019 – GSE106146
5. Chan et al., 2022 – GSE175533
All image data have been deposited to Mendeley Data (https://doi.org/10.17632/mtns8fjk9w.1). See also Source Data (for qPCR analysis, western blot and microscopy images) accompanying this manuscript.
Code availability
All code used in this manuscript has been deposited to GitHub: https://github.com/gdonahue/Sen_NA_2022.
References
Kornberg, R. D. Eukaryotic transcriptional control. Trends Cell Biol. 9, M46–M49 (1999).
Smolle, M., Workman, J. L. & Venkatesh, S. reSETting chromatin during transcription elongation. Epigenetics 8, 10–15 (2013).
Sen, P. et al. H3K36 methylation promotes longevity by enhancing transcriptional fidelity. Genes Dev. 29, 1362–1376 (2015).
Pu, M. et al. Trimethylation of Lys36 on H3 restricts gene expression change during aging and impacts life span. Genes Dev. 29, 718–731 (2015).
Neri, F. et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543, 72–77 (2017).
Brocks, D. et al. DNMT and HDAC inhibitors induce cryptic transcription start sites encoded in long terminal repeats. Nat. Genet. 49, 1052–1060 (2017).
McCauley, B. S. et al. Altered chromatin states drive cryptic transcription in aging mammalian stem cells. Nat. Aging 1, 684–697 (2021).
Munoz-Espin, D. & Serrano, M. Cellular senescence: from physiology to pathology. Nat. Rev. Mol. Cell Biol. 15, 482–496 (2014).
He, S. & Sharpless, N. E. Senescence in health and disease. Cell 169, 1000–1011 (2017).
Sharpless, N. E. & Sherr, C. J. Forging a signature of in vivo senescence. Nat. Rev. Cancer 15, 397–408 (2015).
Collado, M. & Serrano, M. The power and the promise of oncogene-induced senescence markers. Nat. Rev. Cancer 6, 472–476 (2006).
Coppe, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
Baker, D. J. et al. Naturally occurring p16Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
Krimpenfort, P. & Berns, A. Rejuvenation by therapeutic elimination of senescent cells. Cell 169, 3–5 (2017).
Sen, P., Shah, P. P., Nativio, R. & Berger, S. L. Epigenetic mechanisms of longevity and aging. Cell 166, 822–839 (2016).
Yang, N. & Sen, P. The senescent cell epigenome. Aging (Albany NY) 10, 3590–3609 (2018).
Alcorta, D. A. et al. Involvement of the cyclin-dependent kinase inhibitor p16 (INK4a) in replicative senescence of normal human fibroblasts. Proc. Natl Acad. Sci. USA 93, 13742–13747 (1996).
Shah, P. P. et al. Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Genes Dev. 27, 1787–1799 (2013).
Freund, A., Laberge, R. M., Demaria, M. & Campisi, J. Lamin B1 loss is a senescence-associated biomarker. Mol. Biol. Cell 23, 2066–2075 (2012).
Dou, Z. et al. Autophagy mediates degradation of nuclear lamina. Nature 527, 105–109 (2015).
Shimi, T. et al. The role of nuclear lamin B1 in cell proliferation and senescence. Genes Dev. 25, 2579–2593 (2011).
O’Sullivan, R. J., Kubicek, S., Schreiber, S. L. & Karlseder, J. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat. Struct. Mol. Biol. 17, 1218–1225 (2010).
Bahar, R. et al. Increased cell-to-cell variation in gene expression in ageing mouse heart. Nature 441, 1011–1014 (2006).
Burgess, D. J. Epigenetics: therapy-induced transcription is cryptically widespread. Nat. Rev. Cancer 17, 456 (2017).
Mahat, D. B. et al. Base-pair-resolution genome-wide map** of active RNA polymerases using precision nuclear run-on (PRO-seq). Nat. Protoc. 11, 1455–1476 (2016).
Campisi, J. Replicative senescence: an old lives’ tale? Cell 84, 497–500 (1996).
Sen, P. et al. Histone acetyltransferase p300 induces de novo super-enhancers to drive cellular senescence. Mol. Cell 73, 684–698 (2019).
Ernst, J. et al. Map** and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
**, Y., Eser, U., Struhl, K. & Churchman, L. S. The ground state and evolution of promoter region directionality. Cell 170, 889–898 (2017).
Li, B. et al. Infrequently transcribed long genes depend on the Set2/Rpd3S pathway for accurate transcription. Genes Dev. 21, 1422–1430 (2007).
Heyn, P., Kalinka, A. T., Tomancak, P. & Neugebauer, K. M. Introns and gene expression: cellular constraints, transcriptional regulation, and evolutionary consequences. Bioessays 37, 148–154 (2015).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Guenther, M. G., Levine, S. S., Boyer, L. A., Jaenisch, R. & Young, R. A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130, 77–88 (2007).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Landt, S. G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Andersson, R. & Sandelin, A. Determinants of enhancer and promoter activities of regulatory elements. Nat. Rev. Genet. 21, 71–87 (2020).
Giaimo, B. D., Ferrante, F., Herchenrother, A., Hake, S. B. & Borggrefe, T. The histone variant H2A.Z in gene regulation. Epigenetics Chromatin 12, 37 (2019).
Wagner, E. J. & Carpenter, P. B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 13, 115–126 (2012).
**, C. et al. H3.3/H2A.Z double variant-containing nucleosomes mark ‘nucleosome-free regions’ of active promoters and other regulatory regions. Nat. Genet. 41, 941–945 (2009).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Shaulian, E. & Karin, M. AP-1 as a regulator of cell life and death. Nat. Cell Biol. 4, E131–E136 (2002).
Rai, T. S. et al. HIRA orchestrates a dynamic chromatin landscape in senescence and is required for suppression of neoplasia. Genes Dev. 28, 2712–2725 (2014).
Chan, M. et al. Novel insights from a multiomics dissection of the Hayflick limit. eLife 11, e70283 (2022).
Ellyard, J. I. & Vinuesa, C. G. A BATF-ling connection between B cells and follicular helper T cells. Nat. Immunol. 12, 519–520 (2011).
Pakos-Zebrucka, K. et al. The integrated stress response. EMBO Rep. 17, 1374–1395 (2016).
Chu, T. et al. Chromatin run-on and sequencing maps the transcriptional regulatory landscape of glioblastoma multiforme. Nat. Genet. 50, 1553–1564 (2018).
Mitchell, S. J. et al. Effects of sex, strain, and energy intake on hallmarks of aging in mice. Cell Metab. 23, 1093–1112 (2016).
Wang, S. et al. Mechanistic heterogeneity in site recognition by the structurally homologous DNA-binding domains of the ETS family transcription factors Ets-1 and PU.1. J. Biol. Chem. 289, 21605–21616 (2014).
Kowalczyk, M. S. et al. Intragenic enhancers act as alternative promoters. Mol Cell 45, 447–458 (2012).
Martinez-Zamudio, R. I. et al. AP-1 imprints a reversible transcriptional programme of senescent cells. Nat. Cell Biol. 22, 842–855 (2020).
Zhang, C. et al. ATF3 drives senescence by reconstructing accessible chromatin profiles. Aging Cell 20, e13315 (2021).
Moulton, K. S., Semple, K., Wu, H. & Glass, C. K. Cell-specific expression of the macrophage scavenger receptor gene is dependent on PU.1 and a composite AP-1/ets motif. Mol. Cell. Biol. 14, 4408–4418 (1994).
Wu, H., Moulton, K., Horvai, A., Parik, S. & Glass, C. K. Combinatorial interactions between AP-1 and ets domain proteins contribute to the developmental regulation of the macrophage scavenger receptor gene. Mol. Cell. Biol. 14, 2129–2139 (1994).
Madrigal, P. & Alasoo, K. AP-1 takes centre stage in enhancer chromatin dynamics. Trends Cell Biol. 28, 509–511 (2018).
Martin-Herranz, D. E. et al. Screening for genes that accelerate the epigenetic aging clock in humans reveals a role for the H3K36 methyltransferase NSD1. Genome Biol. 20, 146 (2019).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Cruickshanks, H. A. et al. Senescent cells harbour features of the cancer epigenome. Nat. Cell Biol. 15, 1495–1506 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Aoi, Y. et al. NELF regulates a promoter-proximal step distinct from RNA Pol II pause-release. Mol. Cell 78, 261–274 (2020).
Blumberg, A. et al. Characterizing RNA stability genome-wide through combined analysis of PRO-seq and RNA-seq data. BMC Biol. 19, 30 (2021).
Acknowledgements
We would like to thank K. Alexander and D. Mahat for PRO-seq/PRO-cap/ChRO-cap guidance and members of the Berger laboratory for critical reading of the manuscript. We would also like to thank D. Jha for insightful discussions. This work was supported by National Institutes of Health/National Institute on Aging grant P01AG031862 to S.L.B., American Heart Association grant 15POST21230000, AFAR Irene Diamond Transition Award DIAMOND 17113 and National Institute on Aging Intramural Research Program grant ZIA-AG-000679 to P.S.
Author information
Authors and Affiliations
Contributions
P.S. and S.L.B. conceptualized the work. P.S. generated most genome-wide datasets used in this study (except those already published, as indicated in Supplementary Table 2). C.L. performed functional experiments, such as qPCR detection of PRO-cap peaks and replicative senescence assays with cells overexpressing BATF. G.E. performed PRO-cap on cells overexpressing BATF. N.Y. helped in wet lab experiments. G.D., E.K. and Y.L. performed bioinformatics analyses of data. N.R. processed WGBS data, which were generated in the laboratory of P.D.A. D.C.S. provided concentrated lentiviral preparations used in this study. P.P.S. performed ATAC-seq and participated in discussions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Jesus Gil and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information.
Supplementary Figure Legends 1–9, Supplementary Tables 1–4 (Supplementary Table 1 supplied as Supplementary Data), Supplementary Source Data, Supplementary References and Supplementary Figs. 1–9
Supplementary Table 1.
List of Proliferating-not-Senescent, Senescent-not-Proliferating, Young-not-Old and Old-not-Young peaks.
Source data
Source Data Fig. 1
qPCR raw data for Fig. 1i
Source Data Fig. 5
β-gal assay and EdU assay raw data and calculations for Fig. 5a,i
Source Data Fig. 5
Unprocessed western blot and EdU images for Fig. 5e,i
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sen, P., Donahue, G., Li, C. et al. Spurious intragenic transcription is a feature of mammalian cellular senescence and tissue aging. Nat Aging 3, 402–417 (2023). https://doi.org/10.1038/s43587-023-00384-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-023-00384-3
- Springer Nature America, Inc.
This article is cited by
-
Map** medically relevant RNA isoform diversity in the aged human frontal cortex with deep long-read RNA-seq
Nature Biotechnology (2024)
-
Age-associated transcriptional stress due to accelerated elongation and increased stalling of RNAPII
Nature Genetics (2023)
-
Cryptic initiation drives transcriptional junk in ageing
Nature Reviews Genetics (2023)
-
Spurious transcription may be a hallmark of aging
Nature Aging (2023)