Abstract
Ferroptosis is an iron-dependent form of cell death induced by lipid oxidation with an essential role in diseases, including cancer. Since prognostic value of ferroptosis-dependent related genes (FDRGs) in colorectal cancer (CRC) remains unclear, we explored the significance of FDRGs in CRC through comprehensive single-cell analysis. We downloaded the GSE161277 dataset for single-cell analyses and calculated the ferroptosis-dependent gene score (FerrScore) for each cell type. According to each cell type-specific median FerrScore, we categorized the cells into low- and high-ferroptosis groups. By analyzing differentially-expressed genes across the two groups, we identified FDRGs. We further screened these prognosis-related genes used to develop a prognostic signature and calculated its correlation with immune infiltration. We also compared immune checkpoint gene efficacy among different risk groups, and qRT-PCR was performed in colorectal normal and cancer cell lines to explore whether the signature genes could be used as clinical prognostic indicators. In total, 523 FDRGs were identified. A prognostic signature including five signature genes was constructed, and patients were divided into two risk groups. The high-risk group had poor survival rates and displayed high levels of immune infiltration. Our newly developed ferroptosis-based prognostic signature possessed a high predictive ability for CRC.
Similar content being viewed by others
Introduction
Of the most common cancer types, colorectal cancer (CRC) accounts for 9.4% of all cancer deaths worldwide1. Currently, surgery and chemotherapy have significantly improved the survival rates of CRC patients2. However, owing to a lack of efficient clinical treatment and prognostic biomarkers, the overall prognosis of CRC is poor. Moreover, because of tumor heterogeneity, the clinical and histopathological features of tumors cannot currently be used to accurately predict the course of CRC. Therefore, it is critical to identify new prognostic factors and treatment targets for CRC.
Over the past few years, tumor heterogeneity has been shown to be a significant challenge in the treatment and prognosis of cancer. Recently, single-cell RNA sequencing (scRNA-seq) has attracted considerable attention. It allows for the genome-wide analysis of individual cells and makes it possible to understand cellular heterogeneity3,4. Li et al.5 compared the intra-tumor cell heterogeneity between carcinoma and normal tissues in CRC using scRNA-seq. Poonpanichakul et al.6 used a droplet-based scRNA-seq method to profile intra-tumor cell heterogeneity in CRC ascites. However, few studies have been conducted on the cellular heterogeneity of CRC during its evolution from adenoma to carcinoma. In this study, we analyzed the cellular heterogeneity of adenomas and carcinomas by single-cell analysis, and used this information to develop an effective treatment strategy for CRC.
Ferroptosis, an iron-dependent cell death process, is characterized by lipid peroxidation. It is morphologically and mechanistically distinct from the other types of cell death7. Increasing evidence suggests that ferroptosis plays a role in various cancers8. Lu et al.9 found that downregulation of KLF2 inhibits ferroptosis by reducing the transcriptional repression of GPX4 and promoting the invasive activity of renal cell carcinoma. Moreover, because iron metabolism and homeostasis are associated with tumor immunity, they also play a significant role in immunity10. Wang et al.11 demonstrated that the activation of CD8+ T cells could increase ferroptosis and the efficacy of immunotherapy. However, the mechanisms underlying ferroptosis in CRC, and the role that ferroptosis-dependent related genes (FDRGs) of CRC remain unclear. Therefore, it is necessary to understand the pathophysiology and underlying mechanisms of FDRGs in CRC.
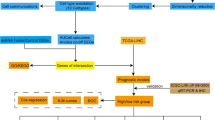
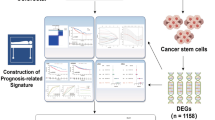
In this study, we calculated ferroptosis-dependent gene scores (FerrScores) for different CRC cell types and used them to define FDRGs. We also constructed a prognostic signature for CRC. Our findings may provide a novel therapeutic strategy for CRC.
Methods
Data collection
To compare the cell heterogeneity in colorectal adenoma and carcinoma tissues, we downloaded the GSE161277 CRC sequence library from the Gene Expression Omnibus database (GEO) (http://www.ncbi.nlm.nih.gov/geo/) and selected four colorectal adenoma and four colorectal carcinoma samples (GSM4904234, GSM4904235, GSM4904236, GSM4904238, GSM4904239, GSM4904242, GSM4904243, and GSM4904245) for single-cell analysis12. We also downloaded the RNA-Seq data and relevant clinical information on CRC from the official website of The Cancer Genome Atlas (TCGA) (https://portal.gdc.cancer.gov/). Furthermore, we obtained the validation dataset GSE17538 from the GEO database. A list of ferroptosis-dependent genes was compiled using the FerrDb website (http://www.zhounan.org/ferrdb/current/).
Processing of sc-RNAseq data
We used the “Seurat” R package (version 4.1.1) and integrated downstream analysis of single-cell transcriptome profiles. The data were quality controlled. Cells with fewer than 300 features and genes expressed in fewer than three cells were excluded. In addition, the proportion of mitochondria was limited to less than 20%. We then normalized the data using the LogNormalization method. We also screened the 2000 highly variable genes with the “FindVariableFeatures” function. Uniform manifold approximation and projection (UMAP) was used for data visualization in 2 dimensions The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request. Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 71(3), 209–249 (2021). Puccini, A. & Lenz, H. J. Colorectal cancer in 2017: Practice-changing updates in the adjuvant and metastatic setting. Nat. Rev. Clin. Oncol. 15(2), 77–78 (2018). Islam, S. et al. Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res. 21(7), 1160–1167 (2011). Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Meth. 6(5), 377–382 (2009). Li, H. et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49(5), 708–718 (2017). Poonpanichakul, T. et al. Capturing tumour heterogeneity in pre- and post-chemotherapy colorectal cancer ascites-derived cells using single-cell RNA-sequencing. Biosci. Rep. 41(12), BSR20212093 (2021). Dixon, S. J. et al. Ferroptosis: An iron-dependent form of nonapoptotic cell death. Cell 149(5), 1060–1072 (2012). Chen, P. et al. Combinative treatment of beta-elemene and cetuximab is sensitive to KRAS mutant colorectal cancer cells by inducing ferroptosis and inhibiting epithelial-mesenchymal transformation. Theranostics 10(11), 5107–5119 (2020). Lu, Y. et al. KLF2 inhibits cancer cell migration and invasion by regulating ferroptosis through GPX4 in clear cell renal cell carcinoma. Cancer Lett. 522, 1–13 (2021). Wang, Y. et al. Iron metabolism in cancer. Int. J. Mol. Sci. 20(1), 95 (2018). Wang, W. et al. CD8(+) T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569(7755), 270–274 (2019). Zheng, X. et al. Single-cell transcriptomic profiling unravels the adenoma-initiation role of protein tyrosine kinases during colorectal tumorigenesis. Sign. Transduct. Target Ther. 7(1), 60 (2022). McInnes, L., Healy, J. & Melville, J. Umap: Uniform manifold approximation and projection for dimension reduction. ar**v preprint ar**v:180203426 (2018). Aran, D., Hu, Z. & Butte, A. J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18(1), 220 (2017). Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013). Tibshirani, R. Regression shrinkage and selection via the lasso. J. Roy. Stat. Soc.: Ser. B (Methodol.) 58(1), 267–288 (1996). Yu, G. et al. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287 (2012). Galon, J. et al. The immune score as a new possible approach for the classification of cancer. J. Transl. Med. 10, 1 (2012). Shimada, K. et al. Frequent genetic alterations in immune checkpoint-related genes in intravascular large B-cell lymphoma. Blood 137(11), 1491–1502 (2021). Jiang, F. et al. An immune checkpoint-related gene signature for predicting survival of pediatric acute myeloid leukemia. J. Oncol. 2021, 5550116 (2021). Song, D. et al. A model of seven immune checkpoint-related genes predicting overall survival for head and neck squamous cell carcinoma. Eur. Arch. Otorhinolaryngol. 278(9), 3467–3477 (2021). Tian, M. et al. A novel immune checkpoint-related seven-gene signature for predicting prognosis and immunotherapy response in melanoma. Int. Immunopharmacol. 87, 106821 (2020). Mei, Y. et al. Single-cell analyses reveal suppressive tumor microenvironment of human colorectal cancer. Clin. Transl. Med. 11(6), e422 (2021). Ren, J. et al. Development and validation of a metabolic gene signature for predicting overall survival in patients with colon cancer. Clin. Exp. Med. 20(4), 535–544 (2020). Luo, D. et al. Development and validation of a novel epigenetic signature for predicting prognosis in colon cancer. J. Cell. Physiol. 235(11), 8714–8723 (2020). Zhou, Y., Guo, Y. & Wang, Y. Identification and validation of a seven-gene prognostic marker in colon cancer based on single-cell transcriptome analysis. IET Syst. Biol. 16(2), 72–83 (2022). Jiang, C. et al. In silico development and clinical validation of novel 8 gene signature based on lipid metabolism related genes in colon adenocarcinoma. Pharmacol. Res. 169, 105644 (2021). Zuo, D. et al. Construction and validation of a metabolic risk model predicting prognosis of colon cancer. Sci. Rep. 11(1), 6837 (2021). Fang, Z. et al. Identification of a prognostic gene signature of colon cancer using integrated bioinformatics analysis. World J. Surg. Oncol. 19(1), 13 (2021). Cheng, X. et al. A novel enterocyte-related 4-gene signature for predicting prognosis in colon adenocarcinoma. Front. Immunol. 13, 1052182 (2022). Huang, W. et al. Prognostic costimulatory molecule-related signature risk model correlates with immunotherapy response in colon cancer. Sci. Rep. 13(1), 789 (2023). Qiu, C. et al. Identification of molecular subtypes and a prognostic signature based on inflammation-related genes in colon adenocarcinoma. Front. Immunol. 12, 769685 (2021). Zheng, L., Yang, Y. & Cui, X. Establishing and validating an aging-related prognostic four-gene signature in colon adenocarcinoma. Biomed. Res. Int. 2021, 4682589 (2021). Ouyang, S. et al. Inhibition of STAT3-ferroptosis negative regulatory axis suppresses tumor growth and alleviates chemoresistance in gastric cancer. Redox. Biol. 52, 102317 (2022). Guo, J. et al. Ferroptosis: A novel anti-tumor action for cisplatin. Cancer Res Treat. 50(2), 445–460 (2018). Nersisyan, S. et al. ECM-receptor regulatory network and its prognostic role in colorectal cancer. Front. Genet. 12, 782699 (2021). Crotti, S. et al. Extracellular matrix and colorectal cancer: How surrounding microenvironment affects cancer cell behavior?. J. Cell Physiol. 232(5), 967–975 (2017). Maltseva, D. V. & Rodin, S. A. Laminins in metastatic cancer. Mol. Biol (Mosk). 52(3), 411–434 (2018). **e, Q. et al. Tspan5 promotes epithelial-mesenchymal transition and tumour metastasis of hepatocellular carcinoma by activating Notch signalling. Mol. Oncol. 15(11), 3184–3202 (2021). Zhu, H., Zhao, N. & Jiang, M. Isovitexin attenuates tumor growth in human colon cancer cells through the modulation of apoptosis and epithelial-mesenchymal transition via PI3K/Akt/mTOR signaling pathway. Biochem. Cell Biol. 99(6), 741–749 (2021). Gorka, J. et al. MCPIP1 inhibits Wnt/beta-catenin signaling pathway activity and modulates epithelial-mesenchymal transition during clear cell renal cell carcinoma progression by targeting miRNAs. Oncogene 40(50), 6720–6735 (2021). Liao, P. et al. CD8(+)T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell 40(4), 365-378.e6 (2022). Li, X. Y. et al. A novel model based on necroptosis-related genes for predicting prognosis of patients with prostate adenocarcinoma. Front. Bioeng. Biotechnol. 9, 814813 (2021). Oshi, M. et al. A novel 4-gene score to predict survival, distant metastasis and response to neoadjuvant therapy in breast cancer. Cancers (Basel) 12(5), 1148 (2020). Shou, Y. et al. TIMP1 indicates poor prognosis of renal cell carcinoma and accelerates tumorigenesis via EMT signaling pathway. Front. Genet. 13, 648134 (2022). Ma, B. et al. TIMP1 promotes cell proliferation and invasion capability of right-sided colon cancers via the FAK/Akt signaling pathway. Cancer Sci. 113(12), 4244–4257 (2022). Farrar, J. E. et al. Ribosomal protein gene deletions in diamond-blackfan anemia. Blood 118(26), 6943–6951 (2011). Ren, Y. et al. A prognostic model for colon adenocarcinoma patients based on ten amino acid metabolism related genes. Front. Public Health. 10, 916364 (2022). Daugaard, M., Jaattela, M. & Rohde, M. Hsp70-2 is required for tumor cell growth and survival. Cell Cycle 4(7), 877–880 (2005). Chen, Q. et al. LASP1 promotes proliferation, metastasis, invasion in head and neck squamous cell carcinoma and through direct interaction with HSPA1A. J. Cell. Mol. Med. 24(2), 1626–1639 (2020). Yang, L. et al. MANF ameliorates DSS-induced mouse colitis via restricting Ly6C(hi)CX3CR1(int) macrophage transformation and suppressing CHOP-BATF2 signaling pathway. Acta Pharmacol. Sin. 44(6), 1175–1190 (2023). Yang, L. et al. Circular RNA hsa_circ_0004277 contributes to malignant phenotype of colorectal cancer by sponging miR-512–5p to upregulate the expression of PTMA. J. Cell. Physiol. https://doi.org/10.1002/jcp.29484 (2020). The authors gratefully acknowledge each editor and reviewer for their efforts on this study. X.X. and X.Z. designed the study and wrote the manuscript. Q.L. and Y.Q. performed the analysis. Y.L. collected the dataset. W.T. reviewed and revised the manuscript. The authors declare no competing interests. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. Xu, X., Zhang, X., Lin, Q. et al. Integrated single-cell and bulk RNA sequencing analysis identifies a prognostic signature related to ferroptosis dependence in colorectal cancer.
Sci Rep 13, 12653 (2023). https://doi.org/10.1038/s41598-023-39412-y Received: Accepted: Published: DOI: https://doi.org/10.1038/s41598-023-39412-yData availability
References
Acknowledgements
Author information
Authors and Affiliations
Contributions
Corresponding author
Ethics declarations
Competing interests
Additional information
Publisher's note
Rights and permissions
About this article
Cite this article