Abstract
Random forests are a popular type of machine learning model, which are relatively robust to overfitting, unlike some other machine learning models, and adequately capture non-linear relationships between an outcome of interest and multiple independent variables. There are relatively few adjustable hyperparameters in the standard random forest models, among them the minimum size of the terminal nodes on each tree. The usual stop** rule, as proposed by Breiman, stops tree expansion by limiting the size of the parent nodes, so that a node cannot be split if it has less than a specified number of observations. Recently an alternative stop** criterion has been proposed, stop** tree expansion so that all terminal nodes have at least a minimum number of observations. The present paper proposes three generalisations of this idea, limiting the growth in regression random forests, based on the variance, range, or inter-centile range. The new approaches are applied to diabetes data obtained from the National Health and Nutrition Examination Survey and four other datasets (Tasmanian Abalone data, Boston Housing crime rate data, Los Angeles ozone concentration data, MIT servo data). Empirical analysis presented herein demonstrate that the new stop** rules yield competitive mean square prediction error to standard random forest models. In general, use of the intercentile range statistic to control tree expansion yields much less variation in mean square prediction error, and mean square prediction error is also closer to the optimal. The Fortran code developed is provided in the Supplementary Material.
Similar content being viewed by others
Introduction
Breiman developed the idea of bootstrap aggregation (bagging) models1, commonly used with bootstrap averages of tree models, as a way of flexibly modeling data. Bootstrap averaging is a way of reducing the prediction variance of single tree models. However, correlations between trees implied that there would be limits to the reduction in prediction errors achieved by increasing the number of trees. The random forest (RF) model was developed by Breiman2 as a way of reducing correlation between bootstrapped trees, by limiting the number of variables used for splitting at each tree node. RF models often achieve much better prediction error than bagging models. RF models have proved a straightforward machine learning method, much used because of their ability to provide accurate predictions for large and complex datasets and availability in many software packages. The semi-parametric model is determined by three user specified parameters, one of the more critical being the stop** criterion for node splitting, the minimum node size of each potential parent node. The node size regulates the model complexity of each tree in the forest and has implications on the statistical performance of the algorithm. In a recent paper Arsham et al.3 proposed using as stop** criteria the size of the offspring nodes and showed in a series of simulation studies circumstances in which performance over a standard RF model could be improved in this way.
The original RF algorithm by Breiman2 used the minimum size of the parent node to limit tree growth. This implementation of the RF algorithm has been utilized in several packages including the randomForest4 and ranger5 packages; ranger5 appears to be among the most efficient implementation of the standard RF algorithm. The problem of how to select the node size in RF models has been much studied in the literature6,7. There are a number of available packages that allow for alternatives to the standard parental node size limit for node splitting. In particular the randomForestSRC8 and the partykit9,10 R packages both allow for splits to be limited by the size of the children nodes.
In this short paper we outline a number of variant types of RF algorithms, generalizations of the RF model developed by Breiman2, and which use a number of different criteria for stop** tree expansion, in addition to the canonical ones of Breiman2 and Arsham et al.3. We illustrate fits of model to the National Health and Nutrition Examination Survey (NHANES) data and four other datasets, the Tasmanian Abalone data, the Boston Housing crime rate data, the Los Angeles ozone concentration data, and the MIT servo data; these last four datasets are all as used in the paper of Breiman2. Further description of the data is given in Table 1.
Results
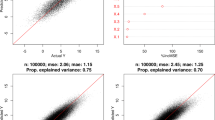
As can be seen from Table 2 and Fig. 1, for the NHANES, Tasmanian Abalone and Los Angeles Ozone datasets the default (parent node size) tree-expansion limitation yields the lowest mean square prediction error (MSPE), although in all cases the MSPE is very close for most other tree-expansion limitation statistics. In particular the MSPE using leaf-node limitation is within 2% of that for parent node limitation. However, for the Boston Housing data leaf-node limitation yields an MSPE that is substantially better, by about 4%, than parent-node limitation, and indeed any other method of tree-limitation. The MSPE using 25–75% intercentile range limitation is substantially better than any for the MIT servo data, the only other method that works nearly as well uses 10–90% intercentile range. All other methods of tree-expansion limitation, in particular both leaf-node and parent-node methods, have MSPE that is at least 15% larger (Table 2). In general use of the two intercentile range statistics (intercentile 10–90% range, intercentile 25–75% range) to control tree expansion yield much less variation in MSPE; in particular, using the 25–75% range, the MSPE does not exceed 5% of the MSPE for the best tree-expansion method for each dataset (Fig. 1).
Discussion
We have presented a number of alternative tree-expansion stop** rules for RF models. It appears that for some datasets, in particular the NHANES, Tasmanian Abalone and Los Angeles Ozone data the new types of stop** rules that we fit have very similar MSPE as the standard stop** rules normally used by RF models (Table 2, Fig. 1). However, for two other datasets, the Boston Housing and MIT Servo data, it is clear that two particular variant stop** rules fit substantially better than the standard RF model (Table 2, Fig. 1). In general, use of the intercentile 25–75% range statistic to control tree expansion yields much less variation in MSPE, and MSPE also closer to the optimal. The MSPE for this measure does not exceed 5% of the MSPE for the best tree-expansion method for each dataset (Fig. 1).
One of the parameters in the RF algorithm is the minimum size of the node below which the node would remain unsplit. This is very commonly available in implementations of the RF algorithm, in particular in the randomForest package4. The problem of how to select the node size in RF models is much studied in the literature. In particular Probst et al.7 review the topic of hyperparameter tuning in RF models, with a subsection dedicated to the choice of terminal node size. This has also been discussed from a more theoretical point of view in a related article by Probst et al.6. As Probst et al. document, the optimal node size is often quite small, and in many packages the default is set to 1 for classification trees and 5 for regression trees7. There are a number of packages available that allow for alternatives to the standard parental node size limit for node splitting. In particular the randomForestSRC8 and the partykit9,10 R packages both allow for splits to be limited by the size of the offspring node. As far as we are aware no statistical package uses the range, variance or centile range based limits demonstrated here. It should be noted that the use of limits of parental and offspring node size are not equivalent. While it is obviously the case that if the offspring nodesize is at least \(n\) then the parental node size must be at least \(2n\), the reverse is clearly not the case. For example, it may be that among the candidate splits of a particular node of size \(2n\) would in general be offspring nodes of sizes \(1,2,...,n - 1,n,n + 1,...2n - 1\). Were one to insist on terminal nodes being of size \(n\) then only the split into two nodes each of size \(n\) would be considered, whereas without restriction on the size of the terminal nodes potential candidates would in general include nodes of size \(1,2,...,n - 1,n + 1,...2n - 1\) also, although the splitting variables might not in general allow all these to occur.
Numerous variants of the RF model have been created, many with implementations in R software. For example, quantile regression RF was introduced by Meinshausen11 and combines quantile regression with random forests and its implementation provided in the package quantregForest. Garge et al.12 implemented a model-based partitioning of the feature space, and developed associated R software mobForest (although this has now been removed from the CRAN archive). Seibold et al.13 also used recursive partioning RF models which were fitted to amyotrophic lateral sclerosis data. Seibold et al. have also developed software for fitting such models, in the R model4you package14. Segal and ** rules with specific application to regression trees. However, the basic idea would obviously easily carry over to classification trees, using for example the Gini or cross-entropy loss functions.
Methods
Data
The NHANES data that we use comprises data for the 2015–2016 and 2017–2018 screening samples, the former used to train the RF and the latter as test set. There are n = 4292 individuals in the 2015–2016 data, and n = 4051 individuals in the 2017–2018 data. A total of 19 descriptive variables (features) are used in the model, with laboratory glycohemoglobin percentage as the outcome variable, a continuous measure. The population weights given in these two datasets are used to weight mean square error (MSE). The version of the NHANES data is exactly as used in the paper of Arsham et al.3. We also employ four other datasets, the Tasmanian Abalone data, the Boston Housing crime rate data, the Los Angeles ozone concentration data, and the MIT servo data; these last four datasets are all as used in the paper of Breiman2. A description of all these datasets is given in Table 1. The five datasets are all given in Supplement S1.
Statistical methods
There are minimal adjustable parameters in the standard RF algorithm2, specifically the number of trees (i.e. the number of bootstrap samples, ntree), and the number of variables sampled per node (mtry) used to determine the growth of the tree, and the maximum number of nodes per tree (maxnodes). The version of the algorithm that we have implemented incorporates a number of additional parameters that determine whether tree generation is halted, specifically:
-
(a)
The proportion of the total variance (in the total dataset) of the outcome variable in a given node used to determine whether to stop the further development of the tree from that node downwards;
-
(b)
The proportion of the total range (= maximum − minimum) (in the total dataset) of the outcome variable in a given node used to determine whether to stop the further development of the tree from that node downwards;
-
(c)
The proportion of the intercentile range [X%, 100 − X%] (in the total dataset) of the outcome variable in a given node used to determine whether to stop the further development of the tree from that node downwards. We used X = 10% and X = 25%.
-
(d)
The minimum number of observations per parent node.
-
(e)
The minimum number of observations per terminal (leaf) node.
The tree generation at a particular node is halted if any of conditions (a)–(e) is triggered. In most implementations of the standard RF model2, for example the R randomForest package4, only criteria (d) is available; in some software, in particular in the randomForestSRC8 and partykit9 R packages criteria (d) and (e) are available as options. The paper of Arsham et al.3 outlined the use of criterion (e) in the context of regression trees. Table 2 outlines the minimum mean square prediction error (MSPE) obtained using the 2017–2018 NHANES data as test set, with model training via the 2015–2016 data. For all other datasets MSPE was defined via tenfold cross validation. In all cases MSPE was the minimum value using ntree = 1000 trees with maxnodes = 1000. We employed a number of sampled variables per node mtry generally about half the total number of independent variables, so mtry = 10, 4, 7, 5, 2, for the NHANES, Tasmanian Abalone, Boston Housing, Los Angeles Ozone and MIT Servo datasets, respectively.
In all cases the categorical variables are treated simply as numeric (non-categorical) variables. We also performed additional model fits in which we used Breiman’s method of coding categorical variables2, but as these generally yielded inferior model fits, as measured by the minMSPE, we do not report these further.
The Fortran 95-2003 code implementing the regression random forest algorithm described above is given in Supplement S1, along with a number of parameter steering files for the five datasets fitted.
Ethics declaration
This study has been approved annually by the National Cancer for Health Statistics Research Ethics Review Board (ERB), and all methods were performed in accordance with the relevant guidelines and regulations of that ERB. All participants signed a form documenting their informed consent, and participants gave informed consent to storing specimens of their blood for future research.
Data availability
The National Health and Nutrition Examination Survey data is freely available for download from https://wwwn.cdc.gov/nchs/nhanes/continuousnhanes/default.aspx?BeginYear=2015 (2015–2016 data) and https://wwwn.cdc.gov/nchs/nhanes/continuousnhanes/default.aspx?BeginYear=2017 (2017–2018 data).
References
Breiman, L. Bagging predictors. Mach. Learn. 24, 123–140. https://doi.org/10.1007/bf00058655 (1996).
Breiman, L. Random forests. Mach. Learn. 45, 5–32. https://doi.org/10.1023/a:1010933404324 (2001).
Arsham, A., Rosenberg, P. & Little, M. Effects of stop** criterion on the growth of trees in regression random forests. New Engl. J. Stat. Data Sci. https://doi.org/10.51387/22-NEJSDS5 (2022).
randomForest: Breiman and Cutler's Random Forests for Classification and Regression. Version 4.6-14 (CRAN—The Comprehensive R Archive Network, 2018).
ranger. Version 0.12.1 (CRAN—The Comprehensive R Archive Network, 2020).
Probst, P., Boulesteix, A.-L. & Bischl, B. Tunability: Importance of hyperparameters of machine learning algorithms. J. Mach. Learn. Res. 20, 1–32 (2019).
Probst, P., Wright, M. N. & Boulesteix, A.-L. Hyperparameters and tuning strategies for random forest. WIREs Data Mining Knowl. Discov. 9, e1301. https://doi.org/10.1002/widm.1301 (2019).
randomForestSRC. Version 2.9.3 (CRAN—The Comprehensive R Archive Network, 2020).
partykit. Version 1.2-15 (CRAN—The Comprehensive R Archive Network, 2021).
Hothorn, T. & Zeileis, A. partykit: A modular toolkit for recursive partytioning in R. J. Mach. Learn. Res. 16, 3905–3909 (2015).
Meinshausen, N. Quantile regression forests. J. Mach. Learn. Res. 7, 983–999 (2006).
Garge, N. R., Bobashev, G. & Eggleston, B. Random forest methodology for model-based recursive partitioning: The mobForest package for R. BMC Bioinform. 14, 125. https://doi.org/10.1186/1471-2105-14-125 (2013).
Seibold, H., Zeileis, A. & Hothorn, T. Model-based recursive partitioning for subgroup analyses. Int. J. Biostat. 12, 45–63. https://doi.org/10.1515/ijb-2015-0032 (2016).
model4you. Version 0.9-7 (CRAN—The Comprehensive R Archive Network, 2020).
Segal, M. R. & **ao, Y. Multivariate random forests. Wiley Interdiscipl. Rev. Data Mining Knowl. Discov. 1, 80–87 (2011).
MultivariateRandomForest. Version 1.1.5 (CRAN—The Comprehensive R Archive Network, 2017).
Wager, S. & Athey, S. Estimation and inference of heterogeneous treatment effects using random forests. J. Am. Stat. Assoc. 113, 1228–1242. https://doi.org/10.1080/01621459.2017.1319839 (2018).
Foster, J. C., Taylor, J. M. & Ruberg, S. J. Subgroup identification from randomized clinical trial data. Stat. Med. 30, 2867–2880. https://doi.org/10.1002/sim.4322 (2011).
Li, J. et al. A multicenter random forest model for effective prognosis prediction in collaborative clinical research network. Artif. Intell. Med. 103, 101814. https://doi.org/10.1016/j.artmed.2020.101814 (2020).
Speiser, J. L. et al. BiMM forest: A random forest method for modeling clustered and longitudinal binary outcomes. Chemometr. Intell. Lab. Syst. 185, 122–134. https://doi.org/10.1016/j.chemolab.2019.01.002 (2019).
Quadrianto, N. & Ghahramani, Z. A very simple safe-Bayesian random forest. IEEE Trans. Pattern Anal. Mach. Intell. 37, 1297–1303. https://doi.org/10.1109/TPAMI.2014.2362751 (2015).
Ishwaran, H., Kogalur, U. B., Blackstone, E. H. & Lauer, M. S. Random survival forests. Ann. Appl. Stat. 2, 841–860 (2008).
Díaz-Uriarte, R. & Alvarez de Andrés, S. Gene selection and classification of microarray data using random forest. BMC Bioinform. 7, 3. https://doi.org/10.1186/1471-2105-7-3 (2006).
Diaz-Uriarte, R. GeneSrF and varSelRF: A web-based tool and R package for gene selection and classification using random forest. BMC Bioinform. 8, 328. https://doi.org/10.1186/1471-2105-8-328 (2007).
van Lissa, C. J. metaforest: Exploring Heterogeneity in Meta-analysis Using Random Forests. R Package Version 0.1.3. https://CRAN.R-project.org/package=metaforest (2020). Accessed August 2022.
Georganos, S. et al. Geographical random forests: A spatial extension of the random forest algorithm to address spatial heterogeneity in remote sensing and population modelling. Geocarto Int. 36, 121–136. https://doi.org/10.1080/10106049.2019.1595177 (2021).
Zhang, G. & Lu, Y. Bias-corrected random forests in regression. J. Appl. Stat. 39, 151–160. https://doi.org/10.1080/02664763.2011.578621 (2012).
Song, J. Bias corrections for random forest in regression using residual rotation. J. Korean Stat. Soc. 44, 321–326. https://doi.org/10.1016/j.jkss.2015.01.003 (2015).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning. Data Mining, Inference, and Prediction 2nd edn, 1–745+i-xxii (Springer, 2017).
Acknowledgements
This work was supported by the Intramural Research Program of the National Institutes of Health, the National Cancer Institute, Division of Cancer Epidemiology and Genetics.
Funding
Open Access funding provided by the National Institutes of Health (NIH).
Author information
Authors and Affiliations
Contributions
M.P.L.: Conceptualization, Methodology, Investigation, Software, Formal analysis, Validation, Writing original draft, Data curation. P.S.R.: Writing—review and editing. A.A.: Investigation, Data curation, Writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Little, M.P., Rosenberg, P.S. & Arsham, A. Alternative stop** rules to limit tree expansion for random forest models. Sci Rep 12, 15113 (2022). https://doi.org/10.1038/s41598-022-19281-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-19281-7
- Springer Nature Limited