Abstract
Fecal microbiota transplantation (FMT) is a therapeutic intervention for inflammatory diseases of the gastrointestinal tract, but its clinical mode of action and subsequent microbiome dynamics remain poorly understood. Here we analyzed metagenomes from 316 FMTs, sampled pre and post intervention, for the treatment of ten different disease indications. We quantified strain-level dynamics of 1,089 microbial species, complemented by 47,548 newly constructed metagenome-assembled genomes. Donor strain colonization and recipient strain resilience were mostly independent of clinical outcomes, but accurately predictable using LASSO-regularized regression models that accounted for host, microbiome and procedural variables. Recipient factors and donor–recipient complementarity, encompassing entire microbial communities to individual strains, were the main determinants of strain population dynamics, providing insights into the underlying processes that shape the post-FMT gut microbiome. Applying an ecology-based framework to our findings indicated parameters that may inform the development of more effective, targeted microbiome therapies in the future, and suggested how patient stratification can be used to enhance donor microbiota colonization or the displacement of recipient microbes in clinical practice.
Similar content being viewed by others
Main
Fecal microbiota transplantation involves the transfer of gut microbes, viruses and luminal content to modulate a recipient’s microbiome, for therapeutic purposes. While the efficacy of FMT has been demonstrated for various diseases1,2,3, such as recurrent Clostridioides difficile infection (rCDI)4,5 or ulcerative colitis (UC6,7), it may also facilitate microbiome recovery following disturbance8 and can enhance microbiome-mediated responses to other therapies9,10. Nevertheless, despite demonstrable efficacy in a growing range of clinical applications, the mode of action of FMT remains poorly understood3 and neither clinical success nor adverse outcomes are currently predictable with accuracy.
Because FMT primarily targets the microbiome, the engraftment of ‘beneficial’ and/or displacement of ‘detrimental’ microbes are expected to cause clinical effects3, in conjunction with more specific processes of host–microbiome interplay, such as the modulation of immune responses11, restored short-chain fatty acid (SCFA) metabolism12 or reinstated phage pressure13,14. It has been argued that both microbiome engraftment and clinical success are mainly determined by donor factors, and that rationally selected ‘super-donors’ may improve therapeutic efficacy15,16. This donor-centric view has since been questioned, at least for some indications17, highlighting the importance of recipient18,19,20 or procedural21 factors instead.
Changes in microbial compositions following FMT have been studied with regard to phages22 or fungi23,24, yet the bulk of current knowledge is focused on bacteria and archaea where colonization by donor microbes and the persistence of indigenous recipient microbes emerge at the strain level of microbial populations25. Strain-level studies suggest that colonization levels following FMT vary across indications: whereas donor and recipient strains coexist long term in metabolic syndrome (MetS) patients25, donor takeover is the most common outcome in rCDI26,27,28, with intermediate outcomes in UC29 or obesity30,31. However, the factors sha** these differential strain-level outcomes remain poorly understood. In small pilot study cohorts, colonization success of donor strains leading to short-term persistence was associated with species phylogeny, broad microbial phenotypes and relative fecal abundances in rCDI26,27, but with more adaptive metabolic phenotypes in UC32.
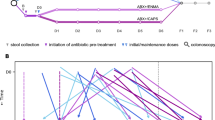
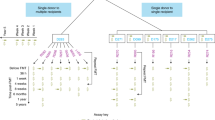
Here we conducted a meta-analysis of novel and published metagenomes from fecal samples collected before and after FMT to compare the fate of donor and recipient strain populations across multiple disease indications. We hypothesized that drivers of FMT response are best studied from an ecological perspective:33,34,5). d, Ternary diagram of the strain population space for conspecific recipient strain persistence, donor strain colonization, donor–recipient coexistence and influx of novel strains.
Donor strain colonization is independent of clinical outcome
Summarized across all tracked species, the colonization and persistence of donor and recipient strains, respectively, varied greatly among allogenic FMT patients (Fig. 2a,b). We observed neither complete recipient strain turnover (loss of all strains) nor complete donor rejection (failure to colonize) in any analyzed FMT instance, although persistence of recipient strains or colonization by donor strains was very low in some patients. Outcomes varied depending on the presence of the species before FMT: takeover by donor strains (accounting for 18.0 ± 16.0% species post FMT) and persistence of recipient strains (11.3 ± 9.1%) occurred more frequently among species present in either donor or recipient, but not in both. In contrast, in cases where species were present in both donor and recipient before FMT, coexistence of donor and recipient strains (19.0 ± 11.8%) was the most frequent outcome compared with donor colonization (4.5 ± 4.0%) and recipient persistence (5.6 ± 5.2%). Among post-FMT strain populations, 41.5 ± 21.0% were attributable to novel strains or entirely novel species not present in either donor or recipient pre FMT (or previously below detection limits). Such major turnover towards novel strains was probably associated with the intervention itself, because novel or previously undetected strains accounted for 50 ± 10.1% in autologous FMTs.
a, Microbiome-level outcomes of 228 scorable allogenic FMT time series, summarized across all strain populations observed in donor and recipient (rec.). Fractions are normalized to the number of species observed in the recipient post FMT. b, Contextual data on indication, procedure and clinical outcome for each FMT time series in a.
Takeover by donor and novel strains was characteristic of patients with rCDI or UC whereas MetS FMTs mostly resulted in conspecific strain coexistence, with varied outcomes in the other tested indications. Clinical response was not associated with strain-level dynamics for any indication; in other words, patient remission was not significantly linked to donor strain colonization or recipient strain displacement—for individual species and across all tracked species (Supplementary Fig. 1). In particular, our data did not support earlier hypotheses that reinstatement of SCFA production is a hallmark of remission in UC and rCDI, because an increased carriage of gut metabolic modules (GMMs; Methods) for acetogenesis, propionigenesis and butyrogenesis following FMT did not correlate with clinical outcome.
Recipient, not donor, factors drive post-FMT strain dynamics
To identify factors associated with colonization outcome, we trained a series of predictive machine learning models using cross-validated LASSO-regularized linear regression (Methods). Among possible predictors we distinguished ex ante variables (that is, knowable before the FMT intervention; Fig. 3a) from post hoc variables (measurable after FMT; Fig. 3b). Moreover, we categorized predictors based on variable scope (procedural, donor related and recipient related) and resolution (host, community and species level), totaling >400 variables as regularization inputs (Supplementary Table 6). We then built cross-validated models for individual predictor categories (for example, using procedural variables only), as well as combined models to assess the overall predictability of outcomes.
a, Ex ante predictability of microbial community-wide outcomes for individual FMTs (summarized across all trackable strain populations in a triad of donor, recipient pre-FMT and recipient post-FMT samples; Fig. 2) using cross-validated LASSO linear models with regularized subsets of different variable categories or a combination of all variables (‘full’ model) knowable before the intervention (Methods and Supplementary Table 6). Within each category, only the most relevant predictors are included. Predictive performance for each outcome index is shown as R2 on the left, and variable importance and directionality for the most predictive factors as cross-validated LASSO coefficients on the right. b, Association of FMT outcomes with LASSO-regularized sets of post hoc variables (measured after the intervention).
Using regularized combinations of ex ante variables, the fractions of species exhibiting post-FMT coexistence of donor and recipient strains and post-FMT recipient strain persistence were predictable with moderate accuracy (LASSO R2 = 0.58 and 0.49, respectively), with lower variation explained for colonization by donor (R2 = 0.34) and pre-FMT recipient strain resilience (R2 = 0.35; Fig. 3a). Interestingly, the fraction of donor strains that successfully took over was not well predicted (R2 = 0.1309).
To identify the major determinants of strain outcomes, we compared the accuracy of models that used restricted subsets of variables with those of full models (which chose from all variables). Models that were restricted to community diversity indices (including species richness) or species abundances in the recipient before FMT achieved similar accuracies, reflecting the importance of these two factors in predicting the fate of donor and recipient strains after FMT. Moreover, across all models, variables capturing recipient factors or donor–recipient microbiome complementarity (for example, community dissimilarity) were more predictive than donor factors. The most important predictors of strain-level outcome included recipient species richness and abundances of selected species in the recipient before FMT, in particular Bacteroides uniformis, Bacteroides vulgatus and one Oscillibacter species, which were positively associated with overall recipient strain persistence and coexistence). In contrast, models based on procedural, metabolic or donor species variables were less accurate (Fig. 3a, left). Notably, donor carriage of GMMs related to SCFA synthesis was not associated with increased strain colonization, contrary to previous findings12. However, high carriage of butyrogenesis genes in the recipient before FMT was moderately associated with overall strain persistence—that is, recipient communities with higher butyrogenesis potential were generally more resilient, further highlighting the role of the recipient microbiome in post-FMT strain dynamics.
In the study population used here, rCDI state was associated with a higher fraction of successfully colonizing donor strains in the post-FMT microbiome. However, we note that while >90% of patients with rCDI in our dataset received antibiotics before intervention, most patients for other indications did not (or underwent extended washout periods), hence rCDI and the effect of antibiotics cannot be disentangled. Moreover, in full models choosing from all variables, higher species richness in the recipient and individual species abundances were more robust predictors for the persistence of recipient strains than rCDI state. This suggests that the high levels of donor strain colonization observed in patients with rCDI may be due in part to a more precarious microbial community (possibly instigated or exacerbated by antibiotic use), rather than being a disease-specific effect.
Models trained on post hoc variables were found to be highly accurate, in particular when describing donor colonization (Fig. 3b). As expected, the strength of community-wide compositional shifts in the recipient (Bray–Curtis dissimilarity and metabolic dissimilarity pre to post FMT) were associated with lower persistence of recipient strains. Interestingly, no individual species’ abundance post FMT was strongly associated with colonization outcome. However, successful colonization of particular species (Fig. 3b, right) was highly predictive of overall colonization of donor strains, in particular B. uniformis, B. vulgatus, several Oscillospiraceae sp. and Lachnospiraceae sp., including Anaerostipes hadrus. These might be considered indicator species, the successful engraftment of which is associated with an overall higher influx of donor strains.
Post-FMT strain outcomes are species specific and predictable
Whereas the above analyses describe summarized outcomes across all tracked species, we next investigated the strain population dynamics within each species post FMT. For sufficient statistical power, we focused on the 307 species detected in >50 allogenic FMTs across our study dataset (Fig. 4 and Supplementary Figs. 1 and 2). Recipient persistence, donor colonization, coexistence and influx of novel strains were observed for all species, with no notable phylogenetic signal. We did not observe any species with consistent patterns of colonization (‘super-colonizers’) or persistence (‘super-persisters’) across all FMTs. However, we observed two broadly distinct types of post-FMT strain dynamics in conspecific FMT triads (that is, for species present in both donor and recipient before the intervention; Fig. 4a and Supplementary Fig. 2). Most species showed a strong propensity towards donor–recipient strain coexistence that was independent of initial strain abundances. Notably, these included prevalent commensals like Bacteroides sp., Blautia sp., Dorea sp., Ruminoccocus sp. and Faecalibacterium sp. In contrast, for Veillonella parvula, several Streptococcus spp., Eggerthella lenta, Akkermansia muciniphila and Prevotella copri, strain populations strongly tended towards dominance of either donor, recipient or novel strains, with infrequent coexistence, indicating that these species may be inherently less prone to conspecific strain carriage within the same host.
a, Strain-level outcomes for selected species are shown for conspecific FMT triads—that is, time series where the focal species was present in both donor and recipient pre FMT. Outcomes are scored as recipient strain persistence (dominance by recipient strains, yellow), donor takeover (blue), donor–recipient coexistence (orange) or influx of novel or previously undetected strains (purple), as indicated in the schematic on the left. Each dot corresponds to one scored FMT. b, Stacked bars representing outcomes for each species across scorable FMTs, scaled to the number of FMTs where the species was observed in the recipient following the intervention. Dashed lines indicate averages for recipient strain persistence within taxonomic groups (x axis). Outcome frequencies across all species are summarized on the left. c, Frequency of colonization by donor or novel (previously undetected) strains per species, as subsets of the data in b. Averages per taxonomic group are represented by dotted lines. d, Prediction accuracies of LASSO models for different binarized FMT outcomes (indicated on the left; Methods) as AUROC, averaged across cross-validation folds per species.
Strain-level FMT outcomes varied within each major taxonomic group, with no relevant differences between clades (Fig. 4b,c). Strains of facultatively aerobic species colonized less successfully (analysis of variance (ANOVA), R2 = 0.02, P = 0.002), whereas carriage of butyrogenesis (R2 = 0.026, P = 2 × 10−4) or propionigenesis (R2 = 0.008, P = 0.05) pathway genes or a generally saccharolytic (R2 = 0.046, P = 1.1 × 10−6) or proteolytic (R2 = 0.047, P = 8.5 × 10−7) metabolic setup was associated with higher colonization success.
To disentangle the factors contributing to post-FMT strain outcomes for each species, we built species-specific cross-validated logistic LASSO regression models using ex ante and post hoc sets of predictor variables, analogous to those discussed above (Fig. 4d). For each species we categorized strain-level outcomes, defining recipient resilience as events where recipient strains persisted (as dominant populations or coexisting with donor strains; yellow), donor colonization (donor strains successfully colonized as dominant or coexisting populations; light blue), donor takeover (donor strains become dominant; dark blue) and recipient turnover (dominance by donor strains and/or new or previously undetectable strains; purple). When training models using all available ex ante variables, recipient resilience (LASSO area under the curve (AUC) = 0.62 ± 0.13), donor colonization (0.58 ± 0.10) and donor takeover (0.65 ± 0.14) were predictable with moderate accuracy, with some variation within and between taxonomic clades (Fig. 4d). In contrast, recipient strain turnover (AUC = 0.94 ± 0.05) was predictable with high accuracy across almost all species, indicating that the displacement of resident strain populations in the recipient (not only by donor strain takeover, but by any means) may in general be a more deterministic process.
Recipient microbiome drives species-specific strain dynamics
We built LASSO models that were restricted to different subcategories of predictor variables and compared their performance with full models trained on the entire complements of ex ante or post hoc variables (Fig. 5a). Models trained exclusively on recipient pre-FMT species abundances, on abundance and strain population characteristics of the focal species and, to a lesser degree, on microbiome community diversity variables achieved highest accuracies, comparable to those of full models. Notably, predictive power of individual recipient species was due almost entirely to exclusion effects, meaning that the enrichment of certain species in the recipient was associated with less donor takeover or recipient strain turnover of others, while facilitation effects did not have a contributing role. Models restricted to procedural factors (including disease indication), pre-FMT metabolic state or donor species abundances achieved much lower accuracies than full models, indicating that these variable groups were less predictive of strain-level outcomes. Overall, we observed similar trends for models trained on post hoc variables (Fig. 5a, right).
a, Logistic LASSO models were trained to predict FMT binarized outcomes (recipient resilience, yellow; recipient turnover, purple; donor takeover, blue) for n = 307 species across FMT time series, using different subsets of ex ante variables (knowable before the intervention). Each dot represents data for one species. Data are shown for full models (choosing from all available variables) and models trained on variable subsets categorized by type (procedural, community-level diversity and so on). Predictive performance of species models is shown as average AUROC across LASSO cross-validation folds in marginal box plots, ranging from 0.5 to 1.0; center line, median; box limits, upper and lower quartiles; whiskers, maxima/minima within 1.5× interquartile range from upper/lower quartiles. b, Variable importance across full models to predict takeover by donor strains. Each edge indicates the importance of a predictor variable (top row) when predicting donor takeover for a given species (bottom row). Dot size for predictors indicates summed variable importance across all species; dot size for species (bottom) indicates total number of relevant predictors. Edge color and width indicate direction and strength of the association, respectively. c, Variable importance for individual predictor categories, as subsets of the data in b.
For most species, we found that strain turnover could be accurately predicted using only two community-level microbiome diversity measures—species richness in the pre-FMT recipient and donor–recipient community dissimilarity, the main factors selected in models restricted to community diversity variables (Fig. 5b). Low richness and a strong compositional shift in the recipient microbiome relative to healthy donors are hallmarks of disease-associated microbiome states, and our data indicate that the strength of this diffuse imbalance, correlated to disease (such as rCDI or UC in our dataset) or other disturbances (for example, antibiotics pretreatment or bowel cleansing), is directly linked with FMT outcome in most species. In contrast, donor richness or functional redundancy, previously proposed to be relevant49, were only subordinately predictive, if at all. Metabolic variables were likewise unreliable predictors. Community-wide butyrogenesis potential was negatively associated with turnover in the recipient (that is, strain populations were more resilient in recipients carrying high loads of butyrate production genes), but higher butyrogenesis levels in the donor did not correspondingly promote colonization. However, in full models for recipient strain turnover, these variables were superseded by indicator species in the recipient microbiome (see below) and focal species characteristics (in particular, recipient strain population diversity; Fig. 5b).
The strongest predictor of takeover by donor strains was a high donor/recipient abundance ratio of a species (as suggested previously for rCDI27), indicating that the amount of incoming viable donor microbes (also referred to as propagule pressure) may provide a neutral baseline estimate for donor strain colonization success, in particular for species not present in the recipient pre FMT (Fig. 5b,c). In general, while the donor/recipient ratio was most predictive, the underlying signal was driven by species abundance (or absence) in the recipient microbiota, much less so in the donor microbiota. Intraspecific strain population properties—donor/recipient strain population dissimilarity and recipient (and, to a much lesser extent, donor) strain population diversity—were also highly predictive but effects were more nuanced: donor strain takeover was more likely in species with complementary strain populations between donor and recipient, while diverse recipient populations (not dominated by individual strains) were more resilient than uneven ones. Moreover, incoming species that were phylogenetically complementary to the recipient community (that is, adding novelty—for example, by filling an unoccupied niche) were more likely to colonize or turn over the resident population.
Resident ‘gatekeeper’ species inhibit donor strain engraftment
Given that FMTs involve the pitting of the recipient’s residual microbial community against incoming microbiota from the donor, we specifically explored the impact of individual species on the engraftment of others by training models restricted to donor or recipient pre-FMT species abundances (Fig. 5a) and exploration of individual species’ relevance as predictors in full models (Fig. 5b,c). We extracted networks of engraftment inhibition and facilitation, associating the abundance of putative effector species in the donor and recipient with donor takeover events in focal species. The vast majority of interactions was inhibitive (Fig. 5a–c): for most species, higher abundance in both donor and recipient correlated negatively with engraftment of other species. These exclusion effects were stronger for the resident community of the recipient (AUC = 0.63 ± 0.14) than the donor (AUC = 0.53 ± 0.06).
Colonization inhibition was phylogenetically concentrated—that is, inhibitive interactions were more common between related species within the same clade than between clades (Fig. 5B). Bacteroidales in the recipient microbiota, in particular B. uniformis, B. vulgatus, Alistipes shahii and Parabacteroides distasonis, were among the strongest colonization inhibitors, but also included two of the most strongly inhibited species, Bacteroides xylanisolvens and Bacteroides ovatus. In other words, the enrichment of gatekeeper species such as Bacteroidales in the recipient microbiota inhibited colonization for a broad panel of species, and vice versa, in line with previous findings that subgroups of Bacteroidales are generally highly persistent also in healthy individuals50. Lactococcus lactis, Streptococcus salivarius and Dialister invisus in the recipient were the foremost colonization facilitators. In contrast to colonization inhibition, facilitation typically affected phylogenetically distant species—for example, the facilitation of Paraprevotella clara and Erysipelatoclostridium ramosum colonization by recipient Pauljensenia sp. (an Actinobacterium) were among the strongest interactions observed across all species.
We observed few prominent predictive species in the donor microbiota, most notably B. vulgatus and Evtepia gabavorous. Facilitation and inhibition effects of donor species were generally limited and overall less predictive of colonization success, indicating that the donor microbiota has limited impact on colonization outcome beyond intraspecific strain dynamics.
Adaptive and neutral processes shape the post-FMT microbiome
The accurate prediction of strain-level outcomes after FMT is informative beyond mere descriptive associations when construed through the lens of gut ecology: FMTs are community-level perturbation experiments, interpretable in a framework of invasion ecology and community assembly to identify processes and mechanisms that shape the microbiome33,34,51,52, community multistability leading to enterotypes53,54, priority55 or ‘Anna Karenina’ effects56. We found limited evidence for colonization facilitation across species boundaries, both in donor and recipient. Likewise, our data did not support a strong role for community-wide metabolic states: neither general metabolic setup nor specific metabolic modules such as SCFA production in donor or recipient greatly impacted FMT outcomes.
The strongest effects toward donor strain colonization emerged at species and strain level. Incoming species were more likely to colonize if they were phylogenetically or metabolically complementary to the residual community, implying that they were able to take over unoccupied niches. Colonization success was associated with complementarity specifically to the local community. High conspecific diversity in the donor and low diversity in the recipient were also linked with engraftment success: recipient populations dominated by single strains were less resilient, and donor strains from more diverse panels were more likely to colonize, probably due to strain-level-limiting similarity effects. Indeed, conspecific donor strain populations colonized more successfully if they were dissimilar to recipient strains, indicating strong inhibitive intraspecific priority effects.
However, we note once more that the colonization of individual species was predictable with only moderate accuracy, irrespective of the variable sets used—unlike residual strain population turnover, which was highly predictable. This implies that colonization success may be stochastic to a large extent.